Abstract
Since no universal codominant markers are currently available, dominant genetic markers, such as amplified fragment length polymorphism (AFLP), are valuable tools for assessing genetic diversity in tropical trees. However, the measurement of genetic diversity (H) with dominant markers depends on the frequency of null homozygotes (Q) and the fixation index (F) of populations. While Q can be estimated for AFLP loci, F is less accessible. Through a modelling approach, we show that the monolocus estimation of genetic diversity is strongly dependent on the value of F, but that the multilocus diversity estimate is surprisingly robust to variations in F. The robustness of the estimate is due to a mechanistic effect of compensation between negative and positive biases of H by different AFLP loci exhibiting contrasting frequency profiles of Q. The robustness was tested across contrasting theoretical frequency profiles of Q and verified for 10 neotropical species. Practical recommendations for the implementation of this analytical method are given for genetic surveys in tropical trees, where such markers are widely applied.
Similar content being viewed by others
Introduction
To date, genetic diversity in tropical trees has only been investigated for a limited number of species (Loveless, 1992, 2002; Caron et al, 2004; Lowe et al, 2005; Ward et al, 2005). The scarcity of studies most likely originates from the lack of molecular tools and genetic markers applicable to a wide range of species. Development of universal markers for gene diversity studies remains, at least for the time being, confined to the chloroplast genomes (Demesure et al, 1995; Grivet et al, 2001). To date, no universal marker system is available for codominant nuclear loci. Such markers need to be developed separately for each species, and can be extremely costly in resources and time (Squirrell et al, 2003). Transferability of microsatellites between species has been reported in several temperate woody taxa (eg Eucalyptus, Byrne et al, 1996; Conifers, Echt et al, 1999; Vitis, Di Gaspero et al, 2000; Fagaceae, Barreneche et al, 2004), and more recently for some tropical species (Meliaceae, White and Powell, 1997; Caryocariaceae, Collevatti et al, 1999). However, all these examples showed that SSR transferability remains limited to phylogenetically related species.
The technical limitations of interspecific transfer of codominant markers has lead to the use of random amplification techniques for assessing genetic diversity, such as RAPDs (random amplified polymorphic DNA, Williams et al, 1990) or AFLPs (amplified fragment length polymorphism, Vos et al, 1995). The RAPD technique has now become less popular due to problems with reproducibility; nevertheless, both random priming techniques can produce an unlimited number of markers, and make them attractive for species where no other codominant markers or DNA sequence data are available.
Recently, two meta-analysis showed that diversity surveys made with dominant markers provide comparable data to surveys undertaken with codominant markers (Nybom and Bartish, 2000; Nybom, 2004). Both class of markers suggest that long-lived, outcrossing, late succession taxa retain most of their genetic variability within populations. One of the major advantages of using random primers to amplify genomic products is that large number of DNA sites can be prospected for polymorphism (Cavers et al, 2005). As shown by simulation studies, the large number and wide distribution of AFLP markers throughout the genome compensates for the poor genetic information content at each locus (Mariette et al, 2002a, 2002b). However, the major drawback to using dominant markers for assessing diversity in species is that the estimation of genetic diversity depends on F, the fixation index of the population, and so the use of dominant markers can be limited when F is unknown, as is usually the case for tropical trees.
Gene diversity studies based on dominant markers compiled in the meta-analysis of Nybom (2004), either assumed that populations were in Hardy–Weinberg equilibrium (F=0) or used associated estimates of F from codominant markers. Much attention was then placed on procedures for estimating allelic frequency and related sampling strategy, but assuming that the F value of the population was known (Lynch and Milligan, 1994). Refined estimation procedures using Bayesian methods have been proposed to reduce the bias in estimating allelic frequencies (Zhivotovsky, 1999). The different estimation procedures produce very similar results for outcrossing species in most cases (Krauss, 2000). However, these papers do little to address the problem of analysing dominant data directly when no F value is available.
The objective of this paper is to evaluate the sensitivity of the diversity estimation procedures to variation in F. As a corollary, we intend to provide empirical recommendations for surveys of genetic diversity in species where no information on F values is available, as is commonly the case for tropical tree species.
Genetic diversity for a diallelic dominant locus
Consider a dominant marker. M and m are, respectively, the marker and the null allele at the considered locus. The genotypes MM and Mm cannot be distinguished, which explains that the marker is considered as dominant in relation to the null allele. Let p be the frequency of the marker allele and q be the frequency of the null allele (q=1−p). Let P be the frequency of the MM genotypes, PMm be the frequency of the Mm genotypes and Q be the frequency of the mm genotypes.
The frequencies of the three genotypic classes in a population are as follows:

where F is the fixation index of Wright, or Hardy–Weinberg deviation.
Based on equation (1), the frequency of the null allele (m) can be calculated as follows:

Hence Nei's genetic diversity can be calculated in the case of a biallelic as follows (see also Caron et al, 2004):

In most tropical species F is unknown. Hence, we considered two extreme cases of F encompassing the range of possible values (F=0 and 1). F=0 corresponds to the case where the population is in Hardy–Weinberg equilibrium, and is generally applicable for outcrossing species. Nei's genetic diversity is then (HHW):

F=1 corresponds to a population where there is no heterozygote (see equation (1)), for example, a fully selfed species. In this case, the frequency of bands is considered as the frequency of the M allele, and all phenotypes observed are homozygous genotypes. As seen in equation (1) if F=1, then Q=q and P=p. As a result, the diversity calculated in this case has also been called the phenotypic diversity (HPH) (Mariette et al, 2002a, 2002b)

It is worthwhile noting that H (equation (3)) varies differently as a function of F depending on the Q values (Figure 1): when Q is ≤0.25, H is a monotonous decreasing function of F, meaning that HHW and HPH are, respectively, the maximum and minimum value attainable of H. When 0.25<Q<0.5, H is a monotonous function of F with a local maximum that does not correspond to HHW or HPH. For example when Q=0.4, HHW=0.465 and HPH=0.48 and the maximum of H (0.5) is obtained when F=0.6. When Q is ≥0.5, H is a monotonous increasing function of F, meaning that HHW and HPH are, respectively, the minimum and maximum value attainable of H. Hence, the use of HHW or HPH to calculate H can be misleading. The bias in measuring H with HHW or HPH can be calculated by

The estimation of H might be biased in different ways (Figure 2). The importance of the biases is clearly dependent on the value of Q. Regardless of the F value, HPH will underestimate H when Q is ≤0.38, and overestimate H when Q is ≥0.5; HHW will overestimate H when Q is ≤0.25, and underestimate H when Q is >0.38. However, at intermediate values of Q (0.38–0.5) and at large values of Q (>0.90), the biases are reduced whatever the method of estimation is used (HHW or HPH).
Genetic diversity for multiple dominant loci
At a single locus, we have shown that HHW and HPH produce biased estimates of H, and that the level of bias depends on the frequency of null allele homozygotes in the population (Q). As genetic diversity is calculated as a mean over all loci, the bias of a multilocus estimate of diversity will strongly depend on the distribution of the Q values of the different AFLP fragments. As an example, consider a preferentially outcrossing species with F value of 0.20, diversity measured by HHW is overestimated for AFLP fragments, which are present at a frequency of 0.60, but underestimated for AFLP fragments with a frequency of 0.2. Hence, the overall diversity over a large number of loci depends on the frequency profile of the AFLP fragments, as biases over all loci may either cumulate or compensate depending on their Q value. We considered here different cases of Q frequency profiles that are likely to encompass the different experimental situations: U-shaped frequency profile of Q (most AFLP fragments showing either low or high frequencies), inverse U-shaped frequency profile (most fragments are at intermediate frequencies), J-shaped frequency profile (excess of fragments at high frequencies), inverse J-shaped frequency profile (excess of AFLP fragments at low frequencies). These frequency profiles are generated by sampling 200 AFLP loci from a beta distribution with parameters a and b (Figure 3). For each profile, 10 repetitions are obtained by sampling with replacement in the beta distribution. And for each repetition, H is calculated according to equation (3) by varying F between −0.2 and 1 (Figure 3).
Multilocus measurement of diversity as a function of the fixation index F. Distribution of the frequency of null homozygote (Q) were generated according to a beta distribution with parameters a and b (a=0.7 and b=0.7 for the U-shaped frequency profile; a=5 and b=5 for the inverse U-shaped profile; a=1 and b=0.6 for the J-shaped profile; a=0.3 and b=0.7 for the inverse J-shaped profile). In all, 10 repetitions were used for each frequency profile and the vertical bars represent the range of variation of H values among the 10 repetitions.
Overall, the multilocus measures of diversity are much less sensitive to the variation in F values than the monolocus measures (comparison of Figures 3 and 1). For the U- and inverse J-shaped frequency profiles, the mean values of diversity do not vary as a function of F. There is a slight increase of H as a function of F in the case of the inverse U-shaped profile, and a slight decrease for the J-shaped profile. In the two latter cases, the diversity measures remain unchanged over a wide range of F values. For example, for F varying between −0.2 and 0.2 (corresponding to a preferentially out-crossed species), or for F varying between 0.8 to 1 (corresponding to a preferentially selfed species), changes of H remain extremely low. Variance due to the stochastic sampling of loci is highest for the J-shaped profiles, corresponding to the case when most of the AFLP fragments exhibit low Q values. In such cases, the estimation of allele frequencies can also be strongly biased and their sampling variance inflated (Lynch and Milligan, 1994).
We also explored other theoretical cases for each of the four categories of frequency profiles (U, inverse U, J and inverse J). The results shown in Figure 3 are consistent across most of the cases investigated (data not shown), with a few exceptions. For the J- and U-shaped profiles, when the profiles are characterized by an extremely large proportion of fragments with high frequencies (eg a=b=0.2 for the U shaped, a=2 and b=0.2 for the J shaped), then H decreases with increasing values of F. Similarly for the inverse J profiles, H increases as a function of F, when the proportion of fragments present in high frequency increases (eg a=0.2 and b=2).
Case studies: genetic diversity of neotropical trees
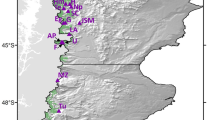
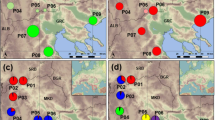
Following the same strategy as in theoretical cases, we estimated genetic diversity for 10 tree species distributed throughout Central and South America with contrasting distributional ranges (from local to continental). For seven out of the 10 species, surveys of AFLP diversity were available for several populations (Table 1). In this study, we bulked all material coming from different populations into a single population (species level) for which diversity was estimated. This is because our investigations are mainly concerned with the impact of the frequency profile on the estimation of diversity and not the comparison of diversity among species. As a result, the fixation index (F) to be considered is FIT, cumulating both the within- and among-population deviations of Hardy–Weinberg equilibrium. Diversity (H) was estimated using the Bayesian method of Zhivotovsky (1999), taking as a prior distribution of Q the observed value of Q in the species. As shown by Zhivotovsky (1999), this method is the least biased estimation procedure, especially when Q is small. Computations of H and the corresponding sampling variances were carried out according to Vekemans et al (2002). Diversity was estimated by considering successive values of F ranging between −0.3 and 1. The frequency profiles of the 10 species (Figure 4) fitted to one of the four theoretical cases considered earlier (ie U, inverse U, J, inverse J; Figure 3), with slight deviations from these general patterns for Anacardium, Cedrela and Voschysia.
AFLP frequency profiles (Freq) of null homozygotes (Q) in 10 neotropical species. Values within brackets correspond to the a and b parameters of the corresponding beta distribution. To compare the observed frequency profiles to the theoretical ones (Table 1 and Figure 3), a and b parameters of the beta distribution were calculated according to Zhivotovsky (1999).
The diversity values for the 10 different species confirmed the results obtained with the theoretical AFLP profiles. HHW or HPH were the two extreme values of H, when F varied between 0 and 1 (data not shown). We did not represent the whole range of variation but only the two extreme values (HHW or HPH; Figure 5). As a result, the real value of H is situated within this range. Among the 10 species investigated, three did not exhibit any difference between HHW and HPH. Anacardium, Cedrela, Virola and Swietenia were among those that showed the largest differences between HHW and HPH. The two former species are also among those that showed irregular frequency profiles (see Figure 4). However, most significantly these four species were those with the lowest number of scored AFLP fragments (Table 1). Finally, all species showed similar values for H when F varied between −0.2 and 0.2, which is the most likely range for variation in fixation index for species exhibiting preferential outcrossing (data not shown).
Diversity estimates of 10 neotropical species as a function of the fixation index. For each species, HPH (−) and HHW ( × ) were estimated, together with their confidence interval (±1.96σ), represented as vertical bars. E.u., Eugenia uniflora; P.m., Pseudobombax munguba; V.f., Voschysia ferruginea; V.m., Virola michelii; E.g., Eperua grandiflora; C.s., Chrysophyllum sanguinolentum; A.o., Anacardium occidentale; C.o., Cedrela odorata; L.c., Lonchocarpus costaricensis; and S.m., Swietenia macrophylla.
Discussion
The measurement of genetic diversity with dominant markers depends on the frequency of null homozygotes (Q) and the fixation index (F) of the populations. While Q can be assessed directly with random priming molecular marker systems, F is less accessible, unless codominant markers are available. The level of the fixation index in a population depends on the mating system and genetic structure (Wahlund effect; Hartl and Clark, 1989). Estimates of the fixation index in tropical trees originate from two sources: mating system analysis and studies of population structure. Indirect estimates of F (F=S/(2−S)) can be obtained from estimates of selfing rates (S). In a review on 26 tropical trees species, Murawski (1995) showed that selfing rates varied between 0 and 80%, but that a majority of tree species were outcrossed. These results were confirmed in two more recent reviews (Loveless, 2002; Ward et al, 2005), partly overlapping with the previous study. In the review of Loveless (2002), only one species among 30 exhibited a selfing rate higher than 15%, and in the review of Ward et al (2005), mixed mating systems were only found in representatives of a single family. Thus, overall we can expect that the fixation coefficient would be, in most cases, less than 10%. However, these values might be inflated in the presence of a Wahlund effect. Significant variation in F has also been noted between adult and juvenile cohorts (Murawski, 1995) and were interpreted as the result of selection favouring heterozygotes. Hence, values of F tend to decrease with population age (Murawski, 1995), which may actually counterbalance any inflating influence from a Wahlund effect.
We show in this study that the monolocus estimation of gene diversity has the potential to vary strongly with variations in F, but that the multilocus estimate is rather robust to deviations in Hardy–Weinberg equilibrium. The robustness of the estimate is due to a mechanistic effect of compensation between negative and positive biases of H estimates for different AFLP loci exhibiting contrasting frequencies of the null homozygote. Surprisingly, the robustness is maintained across a large spectrum of frequency profiles of Q. Existing population data for Q suggest that frequency profiles are in most cases U shaped (Miyashita et al, 1999; Borowsky, 2001). In our survey of 10 neotropical species, we found in addition J- and inverse J-shaped distribution profiles. The robustness of diversity estimation is strongest when frequencies profiles of Q are balanced (U-shaped frequency profiles) or in the case of an inverse J-shaped distribution.
These results lead to important applied consequences for monitoring genetic diversity in species where little information is available on the genetic structure of natural populations. Surveys of genetic diversity in natural populations of tropical trees should therefore follow a stepwise procedure. First, the monitoring should be based on a large number of loci. The usefulness of AFLP for multilocus assessments of diversity lies in the fact that negative and positive biases of H at different loci average out. However, this mechanistic compensation is only effective when many markers are used. From previous simulations (Mariette et al, 2002a, 2002b) and experimental studies (Caron et al, 2004), a few hundred AFLP markers should ideally be recorded in order to cope with the intragenome heterogeneity of diversity. A similar trend was observed in our survey of 10 tropical species (Figure 5): when less than 250 fragments are recorded, the difference between HPH and HHW increases. When no information on the fixation index is available, we recommend estimating H by HPH and HHW. In all theoretical and experimental cases investigated in this study (Figures 3 and 5), these two estimates represent the extreme values of the range of variation of H, and in most cases, this range was extremely small. However, the procedure should be used more cautiously in the case of J- or inverse U-shaped frequency profiles of Q (Figure 3). When there is indirect information available on the mating system, and when the species is considered to be outcrossing or mixed mating, then HHW would be the diversity measure to choice. However, if there is evidence that the species is selfed, then estimation of H by HPH is recommended.
References
Barreneche T, Casasoli M, Russell K, Akkak A, Meddour M, Plomion C et al (2004). Comparative mapping between Quercus and Castanea using simple-sequence repeats (SSRs). Theor Appl Genet 108: 558–566.
Borowsky RL (2001). Estimating nucleotide diversity from random amplified polymorphic DNA and amplified fragment length polymorphism data. Mol Phylogenet Evol 18: 143–148.
Byrne M, Marquez-Garcia MI, Uren T, Smith DS, Moran GF (1996). Conservation and genetic diversity of microsatellite loci in the genus Eucalyptus. Aust J Bot 44: 331–341.
Caron H, Bandou E, Kremer A (2004). Multilocus assessment of levels of genetic diversity in tropical trees in Paracou stands. In: Gourlet Fleury S, Guehl JM, Laroussinie O (eds) Ecology and Management of a Neotropical Rainforest. Elsevier: Amsterdam. pp 160–171.
Cavers S, Navarro C, Lowe AJ (2004). A combination of molecular markers (cpDNA, PCR-RFLP, AFLP) identifies evolutionarily significant units in Cedrela odorata L. (Meliaceae) in Costa Rica. Conserv Genet 4: 571–580.
Cavers S, Degen B, Caron H, Hardy O, Lemes M, Gribel R et al (2005). Optimal sampling strategy for estimation of spatial genetic structure in tree populations. Heredity 95: 281–289.
Cavers S, Navarro C, Hopkins P, Ennos RA, Lowe AJ (in review). Regional and population-scale influences on genetic diversity partitioning within Costa Rican populations of the pioneer tree Vochysia ferruginea Mart. Silvae Genet.
Collevatti RG, Brondnai RV, Grattapaglia D (1999). Development and characterization of microsatellite markers for genetic analysis of a Brazilian endangered tree species. Heredity 83: 748–756.
Demesure B, Sodzi N, Petit RJ (1995). A set of universal primers for amplification of polymorphic non-coding regions of mitochondrial and chloroplast DNA in plants. Mol Ecol 4: 129–131.
Di Gaspero G, Peterlunger E, Testolin R, Edwards KJ, Cipriani G (2000). Conservation of microsatellite loci within the genus Vitis. Theor Appl Genet 101: 301–308.
Echt CS, Vendramin GG, Nelson CD, Marquardt P (1999). Microsatellite DNA as shared genetic markers among conifer species. Can J Forest Res 29: 365–371.
Grivet D, Heinze B, Vendramin GG, Petit RJ (2001). Genome walking with consensus: application to the large single copy region of chloroplast DNA. Mol Ecol Not 1: 345–349.
Hartl DL, Clark DL (1989). Principles of Population Genetics. Sinauer Associates: Sunderland, MA.
Krauss SL (2000). Accurate gene diversity estimates from amplified fragment length polymorphism (AFLP) markers. Mol Ecol 9: 1241–1245.
Loveless MD (1992). Isozyme variation in tropical trees: patterns of genetic organization. New Forests 6: 67–94.
Loveless MD (2002). Genetic diversity and differentiation in tropical trees. In: Degen B, Loveless MD, Kremer A (eds) Modelling and Experimental Research on Genetic Processes in Tropical and Temperate Forest. Embrapa: Belem, PA, Brazil. pp 3–30.
Lowe AJ, Jourde B, Breyne P, Colpaert N, Navarro C, Cavers S (2003). Fine scale genetic structure and gene flow within Costa Rican populations of Mahogany (Swietenia macrophylla). Heredity 90: 268–275.
Lowe AJ, Boshier D, Ward M, Bacles CFE, Navarro C (2005). Genetic resource loss following habitat fragmentation and degradation; reconciling predicted theory with empirical evidence. Heredity 95: 255–273.
Lynch M, Milligan BG (1994). Analysis of population genetic structure with RAPD markers. Mol Ecol 3: 91–99.
Margis R, Felix DB, Caldas JF, Salgueiro F, de Araujo DSD, Breyne P et al (2002). Biodiversity on three neighboring populations of Eugenia uniflora (pitanga) from Brazilian Atlantic rain forest accessed by AFLP markers. Biodivers and Conserv 11: 149–163.
Mariette S, Cottrell J, Csaikl U, Goicoechea P, König A, Lowe AJ et al (2002a). Comparison of levels of genetic diversity detected by AFLP and microsatellite markers within and among mixed Q. petraea (Matt.) Liebl. and Q. robur L. stands. Silvae Genet 51: 72–80.
Mariette S, Le Corre V, Austerlitz F, Kremer A (2002b). Sampling within the genome for measuring within-population diversity: trade-offs between markers. Mol Ecol 11: 1145–1156.
Miyashita NT, Kawabe A, Innan H (1999). DNA variation in the wild plant Arabidoposis thaliana revealed by amplified fragment length polymorphism analysis. Genetics 152: 1723–1731.
Murawski DA (1995). Reproductive biology and genetics of tropical trees from a canopy perspective. In: Loma M, Nadkwarm N (eds) Forest Canopies. Academic Press: New York. pp 457–491.
Navarro C, Cavers S, Colpaert N, Hernandez G, Breyne P, Lowe AJ (In Review). Chloroplast and total genomic diversity in the endemic Costa Rican tree Lonchocarpus costaricensis (JD Smith) Pittier (Papilionaceae). Silvae Genet.
Nybom H (2004). Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol Ecol 13: 1143–1155.
Nybom H, Bartish I (2000). Effects of life history traits and sampling strategies on genetic diversity estimates obtained with RAPD markers in plants. Perspect Plant Ecol Evol Syst 3/2: 93–114.
Squirrell J, Hollingsworth PM, Woodhead M, Russell J, Lowe AJ, Gibby M et al (2003). How much effort is required to isolate nuclear microsatellites from plants? Mol Ecol 12: 1339–1348.
Vekemans X, Beauwens T, Lemaire M, Roldan Ruiz I (2002). Data from amplified fragment length polymorphism (AFLP) markers show indication of size homoplasy and of a relationship between degree of homoplasy and fragment size. Mol Ecol 11: 139–151.
Vos P, Hogers R, Bleeker M, Reijans M, Vandelee T, Hornes M et al (1995). AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23: 4407–4414.
Ward M, Dick CW, Gribel R, Lemes M, Caron H, Lowe AJ (2005). To self, or not to self… A review of outcrossing and pollen-mediated gene flow in neotropical trees. Heredity 95: 246–254.
White G, Powell W (1997). Cross species amplification of SSR loci in the Meliaceae family. Mol Ecol 6: 1195–1197.
Williams JGK, Kubelik AR, Livak KJ, Rafalski A, Tingey SV (1990). DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18: 6531–6535.
Zhivotovsky LA (1999). Estimating population structure in diploids with multilocus dominant DNA markers. Mol Ecol 8: 907–913.
Acknowledgements
This study is part of the EC-funded project GENEO-TROPECO, Contract Number ICA4-CT-2001-10101; http://thoth.nbu.ac.uk/geneo. Samples were collected in Brazil, Costa Rica and French Guiana by researchers at INPA, UFRJ, CATIE and INRA, respectively.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kremer, A., Caron, H., Cavers, S. et al. Monitoring genetic diversity in tropical trees with multilocus dominant markers. Heredity 95, 274–280 (2005). https://doi.org/10.1038/sj.hdy.6800738
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800738
Keywords
This article is cited by
-
Genetic variability of Araucaria angustifolia in the Argentinean Parana Forest and implications for management and conservation
Trees (2018)
-
DNA polymorphisms and genetic relationship among populations of Acacia leucophloea using RAPD markers
Journal of Forestry Research (2018)
-
Genetic structure of Lima bean (Phaseolus lunatus L.) landraces grown in the Mayan area
Genetic Resources and Crop Evolution (2018)
-
AFLP diversity and spatial structure of Calycophyllum candidissimum (Rubiaceae), a dominant tree species of Nicaragua’s critically endangered seasonally dry forest
Heredity (2017)
-
Forest genetic monitoring: an overview of concepts and definitions
Environmental Monitoring and Assessment (2016)



 . βPH is the bias resulting from the assumption that frequencies of phenotypes are equal to allele frequencies (thin curves)
. βPH is the bias resulting from the assumption that frequencies of phenotypes are equal to allele frequencies (thin curves)  .
.

