Abstract
The genetic population structure of Polyommatus coridon (Poda 1761) over large regions of France, Italy and Germany was studied by allozyme electrophoresis. The genetic diversity within populations was high for all parameters analysed (number of alleles 2.72; observed and expected heterozygosity 19.6% and 20.3%, respectively; percentage of polymorphic loci: total: 76.4% and, with polymorphism if the frequency of the commonest allele is below 95%: 53.1%), whereas genetic differentiation between populations was comparatively low (FST = 0.021 ± 0.002). The mean number of alleles declined significantly from southern to northern populations (r = −0.53, P = 0.0005). Similar effects were found also for other parameters of genetic diversity. This is interpreted as a loss of genetic diversity during postglacial expansion. However, samples from France and Italy had similar patterns of genetic diversity indicating no significant loss in this region. Populations from southern Germany were genetically uniform, well differentiated from French populations and showed a significant loss of genetic diversity. Probably, this is due to a bottleneck during passing through the Burgundian Gap, which is a migration corridor from north-eastern France to southern Germany. In contrast to southern German populations, western German populations were not well differentiated from French populations. Nevertheless, they were genetically impoverished, probably as a result from local bottlenecks and post-expansion phenomena.
Similar content being viewed by others
Introduction
The distribution patterns of animals and plants has been subjected to dramatic changes throughout time. Recent changes in the distributions of species are frequently discussed (eg, Parmesan et al, 1999). However, changes in species’ distributions in Europe in the past appear to have been far larger than recent ones (eg, Hewitt, 1996, 1999; Taberlet et al, 1998; Comes and Kadereit, 1999; and references therein). The climate during the last ice-age, the Würm, being totally different from today (eg, Frenzel et al, 1992), forced nearly all plant and animal species to change their European distribution ranges (Huntley and Birks, 1983; Gliemeroth, 1995; Hewitt, 1996, 1999; Taberlet et al, 1998; Comes and Kadereit, 1999). Species thought to have been widely distributed throughout Central Europe during the ice-ages, only survive in alpine or tundral relict areas (Coope, 1970, 1994; Varga, 1977). Thermophilic species were restricted to the more temperate regions of the Mediterranean during glaciations (Hewitt, 1996).
As a consequence of global warming at the end of the Würm, thermophilic species expanded and colonised Central Europe (Hewitt, 1996, 1999). Expansions of tree species can be studied by analysing pollen (eg, Huntley and Birks, 1983; Gliemeroth, 1995). For animals, sufficient remains only exist for some groups, such as beetles or terrestrial gastropods (eg, Coope, 1970, 1994; Hertelendy et al, 1992); for the great majority of invertebrates, hardly any evidence of this kind is available.
The expansion routes can be followed in some cases, particularly plants, through characteristic distribution patterns (see Haeupler and Schönfelder, 1988). However, such clear patterns only rarely are found in mobile invertebrates. Most of these species are now more or less widespread in their distribution ranges. Their presence or absence in a location is mostly determined by their habitat requirements. In such cases, genetic analyses can be a useful tool for coming to a more detailed understanding of postglacial expansions, as has been shown for several animal and plant species (Taberlet et al, 1998; Comes and Kadereit, 1999; Hewitt, 1999; and references therein). However, only very few of these studies have concerned invertebrates (eg, Cooper et al, 1995). Therefore, conclusions about the postglacial expansions of invertebrates are still mainly based on their current distribution patterns (eg, Sternberg, 1998).
Polyommatus coridon (Poda, 1761) is a model organism for mobile invertebrates with strong habitat dependency. These thermophilic animals are mostly restricted to barren grasslands of chalk slopes, and their myrmicophilic larvae feed mainly on Hippocrepis comosa (eg, Ebert and Rennwald, 1991).
We have analysed the population structure of P. coridon from 39 localities over major regions of France, Italy and Germany. In a previous study, we showed that P. coridon colonised our study area from an ice-age refugium in Italy (Schmitt and Seitz, 2001a). Now, we present the analysis of the genetic effects of the postglacial expansion process and their consequences for the identification of possible expansion corridors. Furthermore, the importance of regional geomorphologic structures for the population genetic structure resulting from expansion processes is discussed.
Materials and methods
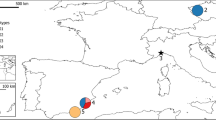
Butterflies were collected at 39 sample stations (see Figure 1), and stored in liquid nitrogen immediately after being captured. Allozyme electrophoresis on acetate plates was performed for 20 loci using standard protocols (Hebert and Beaton, 1993). The alleles were labelled according their relative mobility. Details on enzymes and running conditions are described in Schmitt and Seitz (2001b).
Sampling localities of Polyommatus coridon. Sample numbers correspond to those in Table 1. The northern limit of distribution (Tolman and Lewington, 1998, modified) is indicated by a broken line, the border between the Adriato- and the Ponto-Mediterranean genetic lineages conforming phenotypic differentiation (Schmitt and Seitz, 2001a) is indicated by a dotted line and the distribution range of the Adriato-Mediterranean lineage by a shaded area.
The discrimination between some alleles of Ldh was not always possible. Therefore, the results for this enzyme were excluded from all calculations of genetic distances and identities and all further calculations based on these values.
The allele frequencies, F-statistics (Weir and Cockerham, 1984), Nei's standard genetic distances (Nei, 1978) and RxC χ2-tests (Sokal and Rohlf, 1995) were calculated with G-Stat (Siegismund, 1993). Hierarchical variance analyses and F-statistics were done with ARLEQUIN 2.000 (Schneider et al, 2000). Autocorrelation was computed for the most common allele of each locus with Le Progliciel R 4.0d3 (Casgrain and Legendre, 2001). For the computation of autocorrelation, the geographical distances were subdivided into 20 equidistant classes. Tests on Hardy–Weinberg equilibrium (Louis and Dempster, 1987) and genetic linkage disequilibrium (Weir, 1991) were performed with GENEPOP (Raymond and Rousset, 1995). PHYLIP (Felsenstein, 1993) was used for the cluster analyses; we used the neighbour-joining (Saitou and Nei, 1987) and the UPGMA method. Differences between means were tested with two-tailed U-tests, and correlation analyses were calculated using STATISTICA (Stat Soft Inc. 1993). Means are given with their standard deviations.
Results
Genetic variability
All 20 loci analysed were polymorphic and showed banding patterns consistent with known quaternary structures (Richardson et al, 1986). The highest number of distinguishable alleles was 17 for Pgi; at Aat-1, Acon, Gpd and Fum, only four alleles were distinguished. The mean number of alleles per locus over all populations was 7.3 (±3.3 s.d.). The allele frequencies of all loci are available on request from the authors.
The average number of alleles per locus per population was 2.72 (±0.29 s.d.), ranging from 2.2 to 3.5. The percentage of polymorphic loci (Ptot) had a mean of 76.4% (±7.9 s.d.), ranging from 60% to 95%. The mean percentage of polymorphic loci with the most common allele not exceeding 95% (P95) was 53.1% (±6.4 s.d.), ranging from 45% to 70%. This high percentage of polymorphic loci coincided with high observed heterozygosities (Ho), ranging from 15.9% to 23.6%, with a mean of 19.6% (±1.8 s.d.). The mean expected heterozygosity (He) was slightly higher (20.3% ±1.6 s.d.). All these data are presented in more detail in Table 1.
Hardy–Weinberg equilibrium, linkage disequilibrium and genetic structure
All loci (except 6-Pgdh) showed banding patterns consistent with autosomal inheritance (Richardson et al, 1986). Since all females analysed appeared homozygous at the 6-Pgdh locus, we conclude that this locus is located on the z-chromosome. Thus, females were hemizygous for this locus. It was therefore excluded from the calculation of Hardy–Weinberg equilibrium and FIS.
Over all samples, neither significant deviation from Hardy–Weinberg expectations nor linkage disequilibrium was observed. Therefore, further analyses applied standard population genetical methods.
The genetic distances (Nei, 1978) among samples ranged from 0.014 to 0.041 with an average of 0.020 (±0.004 s.d.). The highest average genetic distance from all other samples was calculated for the population from Carol (Pyrenees) with 0.029 (±0.004). FST (0.021 ± 0.002 s.d.) and FIS (0.022 ± 0.008 s.d.) for all samples analysed were each significantly different from zero.
Regional genetic structures
The mean number of alleles was negatively correlated with the latitude of the sampling localities (r = −0.53, t = 3.82, P = 0.0005, see Figure 2). Three samples showed surprisingly low allele numbers (sample 37: Carol, Pyrenees; sample 38: Monte Baldo, southern Alps; sample 6: Hilbringen, western Germany). Without these three sites, the correlation was stronger (r = −0.67, t = 5.30, P < 0.0001). A significant decline from the south to the north was detected also for the other parameters of genetic diversity. Only the relation between latitude and number of polymorphic loci at the 95% level was not significant (P = 0.093). The genetic distances showed no significant correlation with the geographical distances between sample stations.
Correlation between the latitude of the sample stations and the mean number of alleles per locus for 39 populations of Polyommatus coridon from France, Italy and Germany (r = −0.53; P = 0.0005). Within each of the three distinguished geographical groups (triangles: France and Italy; open circles: southern Germany; filled circles: western Germany) no significant correlation between latitude and allele numbers was found. The samples marked by arrows showed remarkably low mean numbers of alleles.
As can be seen from Figure 2, samples from the southern group (consisting of the samples from France and Italy) predominantly had 2.8 or more alleles per locus. The remaining samples from western and southern Germany usually had less than 2.8 alleles per locus. The mean of the southern group was significantly higher than that of both northern groups (U tests: P = 0.0007 and 0.0145, respectively). Similar results were also obtained for the other studied parameters of genetic diversity. Within each of these three geographical groups, no significant correlation between the parameters of genetic diversity and the latitude was found (eg, number of alleles per locus: France and Italy: r = −0.01, t = −0.04, P = 0.97; southern Germany: r = 0.12, t = 0.32, P = 0.76, western Germany: r = 0.12, t = 0.47, P = 0.64).
Rare alleles (with an overall frequency less than 5.0%, but present in at least three populations) were not equally distributed over all sample stations. Thus, four rare alleles were only found in southern France and Italy (Figure 3a), and one more allele was found in France (including north-eastern France) and Italy, but was absent in Germany. Nine alleles were only found in France/Italy and western Germany (Figure 3b), and another nine alleles only in France/Italy and southern Germany (Figure 3c). Eight alleles were not found in southern France and Italy, however, none of these alleles was found in more than eight populations. Two alleles were restricted to Germany, however, none of these was found in more than four populations.
(a–d) Distribution of several rare alleles (overall percentage less than 5%) in all analysed populations of Polyommatus coridon. (a) Idh-2 allele 2; (b) Pgm allele 1; (c) Me allele 4; (d) number of private alleles (alleles found in only one or two populations). In the black circles, the percentage of the respective allele is given. Populations with open circles lack the allele. For the private alleles, the number of private alleles are given; white circle: no private allele; grey circle: one or two private alleles; black circle: three or more private alleles. The northern limit of distribution is marked by a broken line and the border between the Adriato- and the Ponto-Mediterranean genetic lineages conforming phenotypic differentiation is indicated by a dotted line.
In the samples from the Pyrenees (sample 37: Carol), the southern Alps (sample 38: Monte Baldo) and Central Italy (sample 39: Gran Sasso), we did not find several of the rare alleles. Four, five and four rare alleles, respectively, that were found in the majority of all populations were missing in these three populations. The number of private alleles (rare alleles found in only one or two samples) was significantly higher in samples from south-eastern France (mean: 4.0 ± 1.8) than in all the other samples (mean: 1.2 ± 1.3; U-test: P = 0.0046; Figure 3d).
Based on Nei's (1978) genetic distances, we performed neighbour-joining and UPGMA cluster analyses, each of which showed similar results. The neighbour-joining analysis was the most instructive (Figure 4): all samples from southern Germany (all populations from Baden-Württemberg, Bavaria and Thuringia (excluding Hessel)) clustered together. Within this southern German group, a subgroup can be identified including the samples from the Schwäbische Alb (samples 20 and 21), the Fränkische Jura (samples 25 and 26) and south-western Thuringia (sample 23) which did not differ significantly in their allelic distribution applying an R × C χ2-test.
Neighbour-joining diagram of Polyommatus coridon populations based on genetic distances (Nei, 1978). Sample numbers correspond to Table 1 and Figure 1. With one exception, all nodes are significant at the 5% confidence level before Bonferoni correction. Those not significant after Bonferoni correction are marked with a filled circle. Abbreviations: First part: county code; Second part: BW: Baden-Württemberg, BY: Bavaria, C: Central, NE: North-East, Pyr: Pyrenees, RP: Rhineland-Palatinate, S: South, SL: Saarland, TH: Thuringia; Third part (only for western Germany): CE: central Eifel, Na: Nahe-Bergland region, SE: southern Eifel, SM: Saar-Moselle region; last part: name of sampling locality.
In contrast, samples from western Germany clustered apparently randomly together with populations from France. Ten of these 17 samples formed a group that included only one additional sample from western Lorraine (sample 28: Chauvontcourt). The allelic distribution within this cluster differed significantly between populations (R × C χ2-test).
Hierarchical variance analyses (see Table 2) revealed a high between group variance between France/Italy and southern Germany as well as between western Germany and southern Germany. The between-group variance between France/Italy and western Germany was moderate. The France/Italy group showed no significant variance between its two geographical subgroups southern France/Italy and north-eastern France. Within southern Germany, a relatively high between-group variance was detected between the two major regional subgroups, whereas no significant further substructuring of these two regional subgroups was found. In western Germany, there is no evidence for any such differentiation into regional subgroups.
Significant FST values were calculated for the majority of the geographical clusters analysed (Table 2). In general, FST values of more geographically restricted population clusters were smaller than those from geographically less restricted clusters. Only the four samples from the continuous limestone area of southern Germany (samples 20, 21, 25, 26) showed no significant differentiation.
Autocorrelation analyses for the most common allele of each locus showed significant positive autocorrelation in the first distance group for Idh-1, Aat-2, Gpd, Fum, Me and Acon, and monotonic decline of Moran I values in the following distance groups till 170 to 400 km. The Moran I values obtained for the first distance group did not exceed 0.25. For the other 13 loci, no such autocorrelation was found.
Discussion
Genetic diversity within populations and differentiation between populations
The P. coridon populations analysed showed intrapopulational genetic diversity at a high level. The mean number of alleles per locus (2.72 ± 0.29 s.d.) was much higher than in nearly all available data on butterflies and moths. However, even higher allele numbers have been found for Aglais urticae with 2.84 (Vandewoestijne et al, 1999) and for Heliothis virescens with 3.00 (Sluss et al, 1978). The values for observed and expected heterozygosity as well as the percentages of polymorphic loci were also higher than the average in Lepidoptera (Graur, 1985; Packer et al, 1998). Such high levels of intrapopulational genetic diversity are considered typical for populations with large effective population sizes (Vanderwoestijne et al, 1999). For many species, large effective population sizes result from rather moderate local densities in combination with high vagility and open population structure, eg, Pieris napi (Porter and Geiger, 1995) and Aglais urticae (Vanderwoestijne et al, 1999). P. coridon has a closed population structure. Therefore, its large effective population sizes, more likely result from unusually high local population densities and not from mobility which is only moderate in this species (Bink, 1992; Cowley et al, 2001).
The genetic differentiation between the analysed P. coridon populations is comparatively low (mean genetic distance (Nei, 1978) 0.020 (±0.004 s.d.); FST-value 0.021 (± 0.002 s.d.)). Highly differentiated species (eg, subspecific differentiation) have much higher values such as those seen in Parnassius apollo (Descimon, 1995), Parnassius mnemosyne (Napolitano and Descimon, 1994), Euphydryas editha (Britten et al, 1995) and Euphydryas gilettii (Debinski, 1994). Therefore, the genetic differentiation of the populations of P. coridon analysed appears too weak for a splitting into different subspecies. For more detailed information see Schmitt and Seitz (2001b).
Postglacial expansion
During the last ice-age, thermophilic animal and plant species did not survive in Western and Central Europe (cf. Hewitt, 1996). In the postglacial, many species colonised this region from southern refugia (cf. Taberlet et al, 1998; Comes and Kadereit, 1999; Hewitt, 1999). This was also postulated for P. coridon (eg, Varga, 1977). We assume that the significant negative correlation between latitude of the sampled populations and the mean numbers of alleles as well as other parameters of genetic diversity is a consequence of the expansion from the Adriato-Mediterranean refugium to Central Europe. This phenomenon of genetic erosion during expansion processes has also been frequently observed in other species (cf. Stone and Sunnucks, 1993; Hewitt, 1996; Comes and Kadereit, 1999; with references therein).
The monotonic decline in Moran's I up to distances of 400 km also supports the hypothesis of directional migration with limited subsequent gene flow. Similar results were obtained for the expansive gallwasp Andricus quercuscalicis of which the expansion history is well documented during the last 400 years (Stone and Sunnucks, 1993). However, the genetic differentiation of P. coridon was too weak for the detection of this effect at distances greater than 400 km.
Gradual and continuous expansion processes result in linear loss of genetic variability. This was shown clearly in Andricus quercuscalicis, where the dispersal distance alone explained 79% and 85%, respectively, of the variance of the expected heterozygosity and the mean number of alleles per locus (Stone and Sunnucks, 1993). For P. coridon, latitude, as an estimator of expansion distance, only explained 28% of the geographic variance in the mean number of alleles. For the other parameters analysed, the correlation was even less. Thus, a steady erosion of genetic diversity during the postglacial expansion process is unlikely: different types of dispersal (cf. Ibrahim et al, 1996), acting during different phases of the expansion process, have to be postulated.
The allele numbers and the other parameters reflecting genetic diversity of the populations in or near the assumed glacial refugium (Italy and maybe south-eastern France) did not differ significantly from other more northern French populations. The variance of allele frequencies between these two groups was not significant. Apparently, no major genetic modification or erosion took place during the expansion toward large regions of eastern France. Possibly, a stepping stone expansion without major losses of genetic diversity (as described in Ibrahim et al, 1996) took place in the large eastern French limestone regions that offer favourable conditions for this species.
Similar effects were observed for Aedes albopictus, Pieris napi and Aglais urticae at an even larger geographical level (Kambhampati et al, 1990; Porter and Geiger, 1995; Vanderwoestijne et al, 1999; respectively). One major migration corridor of P. coridon in eastern France is assumed to be along the warm Rhône-Saône river systems (Figure 5) as is also supposed for several dragonfly species (Sternberg, 1998).
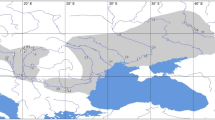
Assumed postglacial migration routes of Polyommatus coridon. The assumed refugium is shown as a hatched area and supposed migration corridors by solid arrows; the Burgundian Gap is marked by a circle. The northern limit of distribution is marked by a broken line and the border between the Adriato- and the Ponto-Mediterranean genetic lineages conforming phenotypic differentiation is indicated by a dotted line. Mountain regions are shaded.
The sample from the Pyrenees (station 37: Carol) showed by far the highest genetic differentiation compared to all other populations. This population was genetically depauperate in comparison with the others from France. This might be due to the marginal geographical position in the Pyrenees and a relatively great distance from the assumed refugium so that a loss of diversity happened during the expansion through southern France. The low level of genetic diversity might, however, be partly explained by the low population density at this sample station. The lack of genetic variation in this sample also confirms the hypothesis that French and German populations of P. coridon are not descended from a glacial refugium in Spain. This is remarkable because France was mainly colonised from Iberian and Balkan refugia and not from Italy, because the Alps acted as efficient dispersal barrier (Taberlet et al, 1998; Hewitt, 1999).
The population from the south-eastern Alps (sample 38: Monte Baldo) had a very low level of genetic heterogeneity compared to other populations from Italy or France. This might be a consequence of the relatively long expansion distance along the southern slopes of the Alps. An Adriato-Mediterranean origin for the south-eastern Alps’ populations has frequently been found in previous works as, for example, in the grasshopper Chorthippus parallelus (Cooper et al, 1995).
The sample from Central Italy (station 39: Gran Sasso) had a medium genetic diversity compared to samples from southern France. However, four widely distributed rare alleles were missing. This supports the hypothesis that the refugium of P. coridon included also northern parts of Italy and, eventually, south-eastern France. This region is known to be one of the most northern arboreal refugia (Huntley and Birks, 1983). However, it has generally been seen as been separated from the Adriato-Mediterranean region of Italy (eg, Konnert and Bergmann, 1995).
The existence of an Adriato-Mediterranean refugium extending as far north as south-eastern France is further supported by the large number of private alleles found in south-eastern France, where the great majority of alleles found in the postglacial expansion area are present. These patterns could be explained if expansion into Central Europe took place from the northern edge of the refugium and if this leading edge of the Adriato-Mediterranean refugium was located in south-eastern France (cf. Hewitt, 1993). A similar situation was shown for the grasshopper Chorthippus parallelus in the Balkans (Cooper et al, 1995).
In Germany, we found less genetic variability than in France and Italy. However, remarkable differences exist between the southern and western German populations. The southern German samples are well differentiated from the French and Italian samples (over 40% of the total variance was between these two groups) and the southern German samples clustered in a separate branch of the neighbor-joining tree. Probably, this is a consequence of the colonisation of this region through the Burgundian Gap between the Jura in the south and the Vosges in the north (see circle in Figure 5). This passage might have caused a genetic bottleneck, unifying southern German populations. However, eight rare alleles passed through this gap from France but are absent from western Germany, hereby showing this colonisation route. The Burgundian Gap is generally suggested an important migration corridor from north-eastern France to southern Germany, for example for dragonflies (Sternberg, 1998).
Within southern Germany, we observed no further significant loss of genetic diversity. Perpaps dispersal conditions in this area were so favourable for P. coridon that, as in eastern France, continuous stepping stone expansion occurred without genetic erosion. However, a division into a more north-western (samples 18, 19, 22, 24) and a more south-eastern group (samples 20, 21, 23, 25, 26) was detected by hierarchical variance analyses (more than 40% of the total variance was between these two groups). This might reflect two main colonisation routes within southern Germany (Figure 5).
For the eastern group, we found a very low level of differentiation between populations: FST values are sometimes not significant, the variance between populations is very low and no hierarchical structure exists. All these populations are linked via a large limestone mountain region extending from the eastern Black Forest to the Basin of Thuringia. This mountain area offers a high density of well suited habitats for this species and likely represents a migration corridor. We suggest that the north-western group used the warm Rhine river valley as an expansion corridor as is well known, for example, from the distribution patterns of many plants (Haeupler and Schönfelder, 1988).
In contrast to the samples from southern Germany, the western German samples (west of the river Rhine) are not well differentiated from the French and Italian samples (only little more than 20% of the total variance was between these two groups). The western German samples clustered in the neighbour-joining diagram apparently at random, together with samples from France and Italy, and eight rare alleles that are absent from southern Germany exist in western Germany and France/Italy. We assume that, during their expansion west of the Vosges (Figure 5), western German populations were not subjected to such a unifying bottleneck as occurred in crossing the Burgundian Gap.
Western and southern German samples showed similar FST values, although the sampled area in western Germany was smaller and the density of sample stations higher than in southern Germany; this indicates a higher level of differentiation between western German populations than between southern German ones. Hierarchical variance analyses revealed no significant hierarchical structure within western Germany. This might result from the relatively unfavourable environmental conditions in western Germany for P. coridon, maybe causing (i) leptokurtic long distance expansion patterns with local losses of genetic diversity (cf. Ibrahim et al, 1996) and /or (ii), genetic erosion in established populations after the expansion process. Comparable results were shown for the gallwasp Andricus quercuscalicis that had much lower genetic differentiation between populations in its favourable native range than in its less favourable expansion area (Stone and Sunnucks, 1993). Station 6 (Hilbringen) apparently is the clearest example for in situ genetic erosion: this geographically isolated sample station has a rather small population size compared to the other populations studied, and it is genetically the most depauperate sample. This phenomenon of genetic erosion of small populations is frequently reported (eg, Buza et al, 2000; Hudson et al, 2000; Jäggi et al, 2000; Madson et al, 2000).
References
Bink, FA (1992). Ecologische Atlas van de Dagvlinders van Noordwest-Europa. Schuyt & Co: Uitgevers en Importeurs, Haarlem.
Britten, HB, Brussard, PF, Murphy, DD, Ehrlich, PR (1995). A test for isolation-by-distance in Central Rocky Mountain and Great Basin populations of Edith's Checkerspot Butterfly (Euphydryas editha). J Hered, 86: 204–210.
Buza, L, Young, A, Thrall, P (2000). Genetic erosion, inbreeding and reduced fitness in fragmented populations of the endangered tetraploid pea Swainsona recta. Biol Conserv, 93: 177–186.
Casgrain, P, Legendre, P (2001). The R package for multivariate and spatial analysis Version 40 (development release 3). Département de Sciences Biologiques, Université de Montréal: Montréal.
Comes, HP, Kadereit, JW (1999). The effect of Quarternary climatic changes on plant distribution and evolution. Trends Plant Sci, 3: 432–438.
Coope, GR (1970). Interpretation of Quaternary insect fossils. Ann Rev Entomol, 15: 97–120.
Coope, GR (1994). The response of insect faunas to glacial-interglacial climatic fluctuations. Phil Trans R Soc Lond, B344: 19–26.
Cooper, SJ, Ibrahim, KM, Hewitt, GM (1995). Postglacial expansion and genome subdivision in the European grasshopper Chorthippus parallelus. Mol Ecol, 4: 49–60.
Cowley, MJR, Thomas, CD, Roy, DB, Wilson, RJ, Léon-Cortés, JL, Guitiérrez, D et al (2001). Density-distribution relationships in British butterflies. I. The effect of mobility and spatial scale. J Animal Ecol, 70: 410–425.
Debinski, DM (1994). Genetic diversity assessment in a metapopulation of the butterfly Euphydryas gilettii. Heredity, 70: 25–30.
Descimon, H (1995). La conservation des Parnassius en France: aspects zoogéographiques, écologiques, démographiques et génétiques. Editions OPIE, Vol. 1: pp 1–54.
Ebert, G, Rennwald, E (eds) (1991). Die Schmetterlinge Baden-Württembergs, Vol. 2. Eugen Ulmer: Stuttgart.
Felsenstein, J (1993). PHYLIP (Phylogeny Inference Package) Ver. 3.5.c. Department of Genetics, University of Washington: Seattle, Washington.
Frenzel, B, Pécsi, M, Velichko, AA (eds) (1992). Atlas of Paleoclimates and Paleoenvironments of the Northern Hemisphere Late Pleistocene-Holocene. Gustav Fischer: Stuttgart.
Gliemeroth, AK (1995). Paläoökologische Untersuchungen über die letzten 22.000 Jahre in Europa. Akademie der Wissenschaften und Literatur, Paläoklimaforschung, Vol. 18. Gustav Fischer Verlag: Stuttgart.
Graur, D (1985). Gene diversity in Hymenoptera. Evolution, 39: 190–199.
Haeupler, H, Schönfelder, P (eds) (1988). Atlas der Farn- und Blütenpflanzen der Bundesrepublik Deutschland. Eugen Ulmer: Stuttgart.
Hebert, PDN, Beaton, MJ (1993). Methodologies for allozyme analysis using cellulose acetat electrophoresis. Helena Laboratories: Beaumont, Tx.
Hertelendy, E, Sümegi, P, Szöör, G (1992). Geochronological and paleoclimatic characterisation of Quaternary sediments in the Great Hungarian Plain. Radiocarbon, 34: 833–839.
Hewitt, GM (1993). Postglacial distribution and species substructure: lessons from pollen, insects and hybrid zones. Lin Soc Symp Ser, 14: 97–123.
Hewitt, GM (1996). Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc, 58: 247–276.
Hewitt, GM (1999). Post-glacial re-colonization of European biota. Biol J Linn Soc, 68: 87–112.
Hudson, QJ, Wilkins, RJ, Waas, JR, Hogg, ID (2000). Low genetic variability in small populations of New Zealand kokako Callaeas cinerea wilsoni. Biol Conserv, 96: 105–112.
Huntley, B, Birks, HJB (1983). An atlas of past and present pollen maps of Europe: 0–13000 years ago. Cambridge University Press: Cambridge.
Ibrahim, KM, Nichols, RA, Hewitt, GM (1996). Spatial patterns of genetic variation generated by different forms of dispersal during range expansion. Heredity, 77: 282–291.
Jäggi, C, Wirth, T, Baur, B (2000). Genetic variability in subpopulations of the asp viper (Vipera aspis) in the Swiss Jura mountains: implications for a conservation strategy. Biol Conserv, 94: 69–77.
Kambhampati, S, Black, WC, Rai, KS, Sprenger, D (1990). Temporal variation in genetic structure of a colonising species – Aedes albopictus in the United States. J Hered, 64: 281–287.
Konnert, M, Bergmann, F (1995). The geographical distribution of genetic variation of silver fir (Abies alba, Pinaceae) in relation to its migration history. Plant System Evol, 196: 19–30.
Louis, EJ, Dempster, ER (1987). An exact test for Hardy–Weinberg and multiple alleles. Biometrics, 43: 805–811.
Madsen, T, Olsson, M, Wittzell, H, Stille, B, Gullberg, A, Shine, R et al (2000). Population size and genetic diversity in sand lizards (Lacerta agilis) and adders (Vipera berus). Biol Conserv, 94: 257–262.
Napolitano, M, Descimon, H (1994). Genetic structure of French populations of the mountain butterfly, Parnassius mnemosyne L. (Lepidoptera: Papilionidae). Biol J Linn Soc, 53: 325–344.
Nei, M (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 89: 583–590.
Packer, L, Taylor, JS, Savignano, DA, Bleser, CA, Lane, CP, Sommers, LA et al (1998). Population biology of an endangered butterfly, Lycaeides melissa samuelis (Lepidoptera: Lycaenidae): genetic variation, gene flow, and taxonomic status. Can J Zool, 76: 320–329.
Parmesan, C, Ryrholm, N, Stefanescu, C, Hill, JK, Thomas, CD, Descimon, H et al (1999). Poleward shifts in geographical ranges of butterfly species associated with regional warming. Nature, 399: 579–583.
Porter, AH, Geiger, H (1995). Limitations to the inference of gene flow at regional geographic scales – an example from the Pieris napi group (Lepidoptera: Pieridae) in Europe. Biol J Linn Soc, 54: 329–348.
Raymond, M, Rousset, F (1995). Genepop (V. 1.2): a population genetics software for exact tests and ecumenicism. J Hered, 86: 248–249.
Richardson, BJ, Baverstock, PR, Adams, M (1986). Allozyme Electrophoresis A Handbook for Animal Systematics and Population Studies. Academic Press: San Diego.
Saitou, N, Nei, M (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol, 4: 406–425.
Schmitt, T, Seitz, A (2001a). Allozyme variation in Polyommatus coridon (Lepidoptera: Lycaenidae): identification of ice-age refugia and reconstruction of post-glacial expansion. J Biogeography, 28: 1129–1136.
Schmitt, T, Seitz, A (2001b). Intraspecific structuring of Polyommatus coridon (Lycaenidae). Nota lepid, 24: 53–63.
Schneider, S, Roessli, D, Excoffier, L (2000). Arlequin ver 2000 – A software for population genetics data analysis. Anthropology: University of Genève.
Siegismund, HR (1993). G-Stat, ver. 3, Genetical statistical programs for the analysis of population data. The Arboretum, Royal Veterinary and Agricultural University: Horsholm, Denmark.
Sluss, TP, Sluss, ES, Graham, HM, Dubois, M (1978). Allozyme differences between Heliothis virescens and H. zea. Ann Entomol Soc Am, 71: 191–195.
Sokal, RR, Rohlf, FJ (1995). Biometry, 3rd edn. W.H. Freeman: New York.
Sternberg, K (1998). The postglacial colonization of Central Europe by dragonflies, with special reference to southwestern Germany (Insecta, Odonata). J Biogeography, 25: 319–337.
Stone, GN, Sunnucks, P (1993). Genetic consequences of an invasion through a patchy environment – the cynipid gallwasp, Andricus quercuscalicis (Hymenoptera: Cynipidae). Mol Ecol, 2: 251–268.
Taberlet, P, Fumagalli, L, Wust-Saucy, A-G, Cosson, J-F (1998). Comparative phylogeography and postglacial colonization routes in Europe. Mol Ecol, 7: 453–464.
Tolman, T, Lewington, R (1998). Die Tagfalter Europas und Nordwestafrikas. Franckh-Kosmos: Stuttgart.
Vandewoestijne, S, Nève, G, Baguette, M (1999). Spatial and temporal population genetic structure of the butterfly Aglaisurticae L. (Lepidoptera, Nymphalidae). Mol Ecol, 8: 1539–1543.
Varga, Z (1977). Das Prinzip der areal-analytischen Methode in der Zoogeographie und die Faunenelement-Einteilung der europäischen Tagschmetterlinge (Lepidoptera: Diurna). Acta Biol Debrecina, 14: 223–285.
Weir, BS (1991). Genetic data analysis. Sinauer: Sunderland, MA.
Weir, BS, Cockerham, CC (1984). Estimating F-statistics for the analysis of population structure. Evolution, 38: 1358–1370.
Acknowledgements
We thank our colleagues Dr Harald Schreiber and Dr Norbert Zahm from the University of the Saarland for the sample from Gran Sasso and many other colleagues for very friendly help in finding many of the sampling localities and for several excursions done together, to Dr Karl Fath (New York, USA) for correcting our English and to two anonymous referees whose annotations considerably improved the manuscript. We also thank the local authorities in Saarbrücken, Trier, Koblenz, Neustadt (Weinstraße), Freiburg im Breisgau, Erfurt, Würzburg, Ansbach, Bayreuth, Regensburg, München and Verona for the permits to collect butterflies and to work in several protected areas. We gratefully acknowledge the Centre for Environmental Studies (ZFU) of the Johannes Gutenberg-University of Mainz and the Deutsche Forschungsgemeinschaft (DFG, grant number SE 506/6–1) for financial support and scholarship of the Graduiertenkolleg of the ZFU for T Schmitt.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Schmitt, T., Gießl, A. & Seitz, A. Postglacial colonisation of western Central Europe by Polyommatus coridon (Poda 1761) (Lepidoptera: Lycaenidae): evidence from population genetics. Heredity 88, 26–34 (2002). https://doi.org/10.1038/sj.hdy.6800003
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800003
Keywords
This article is cited by
-
Suitable climate space and genetic diversity of the mountain-affiliated moth Cosmosoma maishei (Erebidae: Arctiinae: Arctiini: Euchromiina) in cloud forests of Chiapas, Mexico
Journal of Insect Conservation (2023)
-
Genetic variability of the riffle beetle Elmis maugetii Latreille, 1802 (Coleoptera: Elmidae) in Europe and North Africa
Biologia (2022)
-
Regional population structure of the endangered Bridle Shiner (Notropis bifrenatus)
Conservation Genetics (2018)
-
Extra-Mediterranean glacial refugia in a Mediterranean faunal element: the phylogeography of the chalk-hill blue Polyommatus coridon (Lepidoptera, Lycaenidae)
Scientific Reports (2017)
-
Genetic structure of Parnassius mnemosyne (Lepidoptera: Papilionidae) populations in the Carpathian Basin
Organisms Diversity & Evolution (2016)