Abstract
Previous attempts to test for small-scale stock structuring within Atlantic herring (Clupea harengus L.) with molecular markers have been hampered by uninformative levels of genetic variation. Here we report the first application of microsatellite DNA markers to investigate population subdivision in Atlantic herring from Norwegian waters and the Barents Sea, and also examine microsatellite differentiation between C. harengus and Pacific herring (C. pallasi). Results from four microsatellite loci indicate high, and informative, variation compared to molecular markers used previously: number of alleles per locus=18–41; mean expected heterozygosity within samples=0.90–0.93. Significant genetic differences were detected between almost all samples representing postulated Icelandic summer-spawner, Norwegian spring-spawner and Norwegian fjord stocks, using Fisher’s exact test, FST and RST values. Levels of allele frequency differentiation between Atlantic and Pacific herring overlapped the range seen among Atlantic herring samples, indicating that microsatellites are poor indicators of the degree of species differentiation. Comparison with allozyme and mitochondrial DNA restriction fragment length polymorphism (RFLP) datasets from the same samples suggests that microsatellites may detect structuring at a finer scale, but are less informative at larger scales of divergence.
Similar content being viewed by others
Introduction
The specific life-history features of many marine pelagic and demersal fish, such as their high mobility and dispersal capacity, prodigious fecundity and large population sizes, combined with a lack of clear environmental boundaries, have led to doubts concerning the power of established genetic approaches to discriminate stocks or elucidate population structure (Smith et al., 1990; Hedgecock, 1994). Indeed, allozyme and mitochondrial DNA (mtDNA) studies have revealed generally low levels of genetic subdivision when compared with freshwater fish (Hedgecock, 1994). Attention has turned to the development of increasingly polymorphic genetic markers in an attempt to improve discriminatory power. Variable number of tandem repeat (VNTR) loci within nuclear DNA have been identified as a potential source for markers of genetic variation within and among fish populations, initially using minisatellite DNA loci (e.g. Bentzen et al., 1991), but increasingly the more easily assayed microsatellite DNA loci (García de Leon et al., 1997; Estoup et al., 1998; O’Connell et al., 1998; Ruzzante et al., 1998).
Microsatellite loci characteristically exhibit high levels of length mutation, resulting in extensive allelic variation and levels of heterozygosity in fish ranging from 24% to 90% (O’Connell & Wright, 1997). Such levels of variation make them especially suitable for stock identification in species previously exhibiting low levels of detectable variation using allozymes or mtDNA (Bentzen et al., 1991); for example, Ruzzante et al. (1998) found significant heterogeneity at small geographical scales among Atlantic cod populations using microsatellites, in contrast to previous surveys using allozymes (Mork et al., 1985) or mtDNA (Carr & Marshall, 1991). Similarly, O’Connell et al. (1998) used microsatellite markers to demonstrate significant genetic heterogeneity among Pacific herring populations from the Gulf of Alaska and the Bering Sea previously thought to be genetically homogeneous (Grant & Utter, 1984).
The Atlantic herring (Clupea harengus L.) exhibits a mosaic of spatially or temporally separate spawning aggregations throughout the North Atlantic Ocean. Most spawning aggregations have historically been thought to represent separate stocks, based on morphological characters as well as spawning time and location (Svetovidov, 1963; Haegele & Schweigert, 1985). To date however, genetic markers (principally allozymes) have generally shown uniformity across the NW and NE Atlantic (Jørstad et al., 1991), as well as within the offshore waters of the NE Atlantic (Ryman et al., 1984; King et al., 1987), which suggests a lack of locally distinct genetic stocks in contradiction to the morphological and life-history data. Similarly, mtDNA restriction fragment length polymorphism (RFLP) studies have shown genetic homogeneity or low levels of differentiation over large geographical distances (Kornfield & Bogdanowicz, 1987; Dahle & Eriksen, 1990). In contrast to the pattern seen at large geographical scales, discrete localized allozymic differentiation has been detected in Norwegian fjord populations, most notably those within the Balsfjord and Trondheimsfjord (Jørstad et al., 1994; Turan et al., 1998).
The aim here was to test microsatellite primers developed from Pacific herring (Clupea pallasi L.) for their utility in studying genetic variation and population structuring within C. harengus among supposedly discrete spawning aggregations within the NE Atlantic. We also compare levels of genetic divergence indicated by allozyme and mtDNA RFLP data on the same Atlantic herring samples to examine: (i) the utility of the three types of markers for detecting different levels of divergence within a pelagic marine fish; and (ii) the ability of microsatellites to differentiate between C. harengus and C. pallasi, currently considered separate species on the basis of genetic data (Grant, 1986; Domanico et al., 1996) but previously considered morphologically/ecologically similar enough to be subspecies (Svetovidov, 1963).
Materials and methods
Laboratory procedures
Atlantic herring samples were collected by pelagic trawl from sites (indicated in Fig. 1) in the Norwegian Sea and Barents Sea, comprising putative Icelandic summer-spawners (IC) and Norwegian spring-spawners (NS1 and NS2), and from a Norwegian fjord (Balsfjord, BF). One sample of Pacific herring was collected off Vancouver Island (PC). Further details on samples (sampling depth, gear used, mean lengths, age, etc.) can be found in Turan (1997).
Map of the NE Atlantic Ocean showing herring sample locations: Icelandic summer-spawners (IC), Norwegian spring-spawners (Barents Sea) (NS1), Norwegian spring-spawners (central Norwegian Sea) (NS2), Balsfjord herring (BF). The Pacific herring sample (PC) was collected in the NE Pacific, near Vancouver Island.
DNA extraction, from ethanol-preserved muscle tissue, followed the protocol of Taggart et al. (1992). PCR reactions were performed using primers specific to four Pacific herring microsatellite loci (Cha17, Cha20, Cha63, Cha113) as used by O’Connell et al. (1998) under the following conditions: 120 s at 94°C; then 30 cycles of 60 s at 92°C, 60 s at the specific annealing temperature, and 30 s at 72°C. A fifth locus (Cha123) used by O’Connell et al. (1998) did not give consistent amplifications, although subsequent optimization has shown that this locus can also be used reliably for Atlantic herring (P. Bloor, pers. comm.). Reaction mixes contained 20 ng template DNA, 1 mM MgCl, 0.2 mM each nucleotide, 0.5 μM of each primer (forward primer 5′ end-labelled with a Cy5 fluorescent dye group), 0.2 U Taq polymerase (Bioline, U.K.) with the manufacturer’s supplied 1× buffer, in a final reaction volume of 10 μL. Amplified products were checked for yield on an agarose minigel before loading, along with size standards internal to every lane, onto a 6% denaturing acrylamide gel. Gels were run on an ALFexpressTM (Pharmacia Biotech) automated sequencer. Data collation and sizing of alleles were determined using Fragment Manager v1.2 software (Pharmacia Biotech).
Data analysis
The most suitable means of quantifying microsatellite allele frequency differentiation between samples is still a matter of argument, depending upon the mutational model adopted (Garza et al., 1995; O’Connell et al., 1997), although O’Connell & Wright (1997) have recommended a conservative approach of conventional F-statistics for commercial fish species. In view of the continuing debate and for reasons of comparison, we chose to estimate, and thus compare, genetic differentiation among samples by three different methods. First, we tested for simple frequency differentiation between pairs of samples with Fisher’s exact test, using GENEPOP v.3 (Raymond & Rousset, 1995). Secondly, we estimated the degree of the total genetic variation apportioned among samples with unbiased RST (which assumes a Stepwise Mutation Model) using the RST-CALC program (Goodman, 1997), which adjusts for differences in sample size and allele size variances among loci, and finally FST (which assumes an Infinite Allele Model), estimated by theta (θ; Weir & Cockerham, 1984), using F-STAT (Goudet, 1996). Significant departure from genetic homogeneity was tested for in all three cases using permutation tests, contained within the respective software packages. Genotype distributions within samples were tested for conformity to Hardy–Weinberg expectations with exact tests with significance determined by a Markov chain method (GENEPOP v.3). Where multiple tests were involved, significance levels were adjusted according to the sequential Bonferroni procedure (Rice, 1989).
Patterns of genetic distance between samples were displayed using multidimensional scaling techniques (SPSS), a method which avoids clustering of similar samples into groups as in tree building methods, but instead computes co-ordinates for each sample such that the distances between points fit as closely as possible (estimated by r2) to the measured distances between the respective samples. Nei’s (1972) distance (DS) was used, as IAM-based distance measures are predicted to be more accurate for microsatellites in closely related taxa (Slatkin, 1995), although an SMM-based distance (Δμ2; Slatkin, 1995) was also displayed for comparison.
Results
All four Pacific herring microsatellite loci were highly polymorphic in Atlantic herring, with high allelic diversity and heterozygosity in all samples (Table 1). Total number of alleles detected across all Atlantic herring samples pooled ranged from 18 at Cha63 to 41 at Cha20, and expected mean heterozygosity within samples ranged from 90% to 93%. Allele size ranges, allelic diversity and heterozygosity were comparable between the Atlantic herring and Pacific herring samples (Table 1), except for noticeably fewer alleles at Cha20 in the Pacific herring sample (15 vs. 26–27 in Atlantic samples). Allele and genotype frequencies are available from the authors on request.
Genotypic frequencies conformed to Hardy–Weinberg expectations in all but three single-locus tests (Table 1). Multilocus combinations of single-locus tests within samples, using Fisher’s (1954) combined probabilities method, however, indicated that three out of four Atlantic herring samples (IC, NS1 and NS2) exhibited overall departures from out-crossing expectations, and all were caused by deficits of heterozygotes. The sensitivity of the combined probability procedure to single extremely low values should be borne in mind—significance within NS1 and NS2 is primarily caused by results at single loci (Cha20 and Cha113, respectively).
Four-locus combined probabilities of pairwise exact tests between samples indicated that all samples differed significantly in their allele frequencies from all others (all P << 0.001), except NS2 and BF (P=0.039, nonsignificant after Bonferroni correction). Estimates of the among-sample component of genetic variation, whether expressed as FST or RST, indicated significant structuring within the dataset: across all samples (FST=0.024, P < 0.001; RST=0.151, P < 0.001); across all Atlantic herring (FST=0.020, P < 0.001; RST=0.129, P < 0.001); and within Atlantic samples, excluding NS1 (FST=0.009, P < 0.001; RST=0.051, P < 0.001). Values of pairwise RST among samples were consistently much higher (as much as an order of magnitude) than equivalent FST values (Table 2), although both measures indicated the same pattern of relative genetic similarity among samples: the Pacific herring sample was most divergent (means vs. Atlantic: FST=0.037; RST= 0.257); the Barents Sea spring-spawner sample (NS1) was most divergent among the Atlantic herring (FST= 0.025–0.037; RST=0.146–0.273); and the Icelandic summer-spawner (IC), Norwegian Sea spring-spawner (NS2) and the Balsfjord (BF) samples form a closer grouping (FST=0.001–0.018; RST=0.022–0.076). Permutation testing indicated all pairwise estimates of FST and RST were significantly greater than zero except for NS2 × BF (FST and RST) and NS2 × IC (RST only). Paradoxically, the level of differentiation indicated between the Pacific herring and the Atlantic herring samples overlaps that observed within Atlantic herring.
Other than the obvious difference in magnitude between estimates of FST and RST there was no apparent difference between the two statistics in the pattern of genetic differentiation exhibited. Although the data presented here are not extensive enough for a conclusive test of relationship between the two measures, several observations from the data are of potential interest. First, a linear regression of pairwise microsatellite FST on RST was signifcant at P < 0.001, i.e. FST and RST appear to show a linear correlation over the range of values observed here. Secondly, if plotted against pairwise values of FST estimated from allozyme data (see Discussion), both measures appear to approach an asymptote in the region of comparisons between PC/NS1 and the others (data not shown).
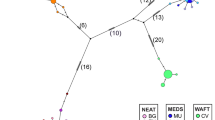
The patterns of genetic differences among the samples described above from FST and RST are reproduced by the multidimensional scaling (MDS) plot (Fig. 2) of pairwise genetic distances (Nei’s DS): the IC/NS2/BF grouping is clearly separate from NS1 within the Atlantic herring, with NS2 being genetically most similar to all three other samples than they are to each other (i.e. being intermediate between the three). The plot of pairwise Δμ2 (Fig. 2) indicated essentially the same pattern, but with distances within the IC/NS2/BF grouping more pronounced relative to distances between this group and NS1.
Multidimensional scaling plots of (a) pairwise DS (Nei, 1972), and (b) Δμ2 (Slatkin, 1995) values between samples of Atlantic herring and Pacific herring (PC). Sample locations: Icelandic summer-spawners (IC), Norwegian spring-spawners (Barents Sea) (NS1), Norwegian spring-spawners (central Norwegian Sea) (NS2), Balsfjord herring (BF). The Pacific herring sample (PC) was collected in the NE Pacific, near Vancouver Island. (MDS plot r2=0.999 and 0.989, respectively).
Discussion
Cross-species utility of herring primers
This is the first report of a microsatellite analysis of population structure in Atlantic herring. Four out of five primer sets designed originally from Pacific herring (C. pallasi) DNA sequences (see O’Connell et al., 1998) work well in C. harengus (and the fifth locus has been successfully optimized subsequent to the present study; P. Bloor, pers. comm.), which indicates a high degree of conservation of primer sites between these two species. Conservation of priming sites has been reported previously among closely related species of fish (Brooker et al., 1994), indicating a promising trans-specific utility of microsatellite primers. One note of caution, however, may be highlighted by the present data: significant deficits of heterozygotes within samples, both at single loci and overall, may indicate a significant presence of nonamplifying null alleles.
All four loci tested were highly polymorphic in C. harengus, with 18–41 alleles per locus over all samples, and mean expected heterozygosities (He) within samples ranging from 0.90 to 0.93 (mean=0.91). Allele size ranges and levels of polymorphism at all four loci within the Atlantic herring samples are very similar to those observed in the Pacific herring sample, and also to those reported previously for Pacific herring from the Gulf of Alaska and the Bering Sea by O’Connell et al. (1998). The few other studies of marine fish to date have shown similar levels of microsatellite polymorphism in Atlantic cod (He=0.85; Brooker et al., 1994) and European sea bass (He=0.79; García de Leon et al., 1997).
Genetic structure of the Atlantic herring population
The most important finding was the degree of genetic structuring indicated within the Atlantic herring population of the Norwegian Sea–Barents Sea area. All tests showed these samples to be genetically differentiated to a degree which suggests restricted gene flow. This finding supports earlier suggestions, based on morphological characters and spawning time/location, of a complex stock structure in Atlantic herring (Svetovidov, 1963; Haegele & Schweigert, 1985). The apparent conflict with earlier allozyme and mtDNA data, which found large-scale genetic homogeneity (Ryman et al., 1984; King et al., 1987; Kornfield & Bagdanowicz, 1987; Dahle & Eriksen, 1990), may arise from the greater information content (polymorphism) of microsatellites. It has been proposed that higher rates of mutation (and therefore polymorphism) of DNA markers results in enhanced power for testing population differentiation (Rousset & Raymond, 1995; Goudet et al., 1996). Estoup et al. (1998) demonstrated within brown trout populations that microsatellites indicated more pronounced differentiation at small geographical scales (within rivers) than allozymes, although both sets of markers gave similar estimates of FST at larger scales (within drainages). Similarly, Shaw et al. (1999) have shown that microsatellites uncover genetic structuring in squid populations at a scale that previous allozyme and mtDNA studies did not detect.
The significant genetic differences observed here between Icelandic summer-spawners and Norwegian spring-spawners support the discrete character of these stocks described on morphological and physiological grounds (Fridriksson, 1958; Turan, 1997). Interestingly, the sample of Norwegian spring-spawners from the Barents Sea (NS1) is the most genetically distinct among the NE Atlantic herring surveyed, including the other Norwegian spring-spawners (NS2). This is also consistent with morphological differences, observed between northern and southern fish (Debarros & Holst, 1995; Stenevik et al., 1996), and indicates that Norwegian spring-spawners should not be treated as a single stock. Balsfjord herring have previously been shown by the use of allozymes and mtDNA to be genetically differentiated from coastal stocks (Jørstad et al., 1994). In contrast to these reports the present study found no evidence for such differentiation between the Balsfjord sample and the southern Norwegian spring-spawner sample (NS2). Studies using allozyme and mtDNA markers on the same samples also did not show significant genetic differences between the BF and NS2 samples (Turan et al., 1998; see Table 3 and discussion below). One possibility is that migratory Norwegian spring-spawners may have been sampled instead of, or together with, resident Balsfjord fish: Norwegian spring-spawners migrate into fjords to feed in the upper water layers, before leaving to spawn on offshore banks (Jørstad et al., 1994). The time (September) and depth (15–20 m) at which our sample was collected raise the possibility that it contained a mix of the two populations with relatively few Balsfjord resident fish. In fact the absence of significant deficits of heterozygotes within this sample, as would be expected from mixing of differentiated populations, suggests that it may contain only migratory fish. It is therefore of obvious importance in future studies to consider localized migratory behaviour and depth distribution when collecting herring from these waters.
Although our results indicate significant genetic differences among populations of Atlantic herring, the proportion of total genetic variation occurring among populations (FST) is relatively small: approximately 1–3%, although much higher (2–27%) if RST is accepted as being a more representative measure. Such levels of differentiation are common for marine fish, estimated from microsatellites or allozymes, and are generally thought to result from levels of migration high enough to prevent genetic isolation of populations, but insufficient to prevent short-term drift in allele frequencies and/or localized adaptation. What factors could cause a pattern of restricted gene flow without reproductive isolation of populations of pelagic marine fish? Four possibilities are considered here: (i) sampling error introducing ‘noise’ into such low estimates (see Waples, 1998); (ii) isolation-by-distance effects; (iii) hydrographic effects; and (iv) behavioural effects. The relationship between sample sizes, large number of alleles per locus and statistical rigour is of some concern to studies employing microsatellites. Ruzzante (1998) has demonstrated empirically that the numbers of individuals and loci screened can substantially affect variance in estimates of a number of genetic distance and structuring measures, but that variance in FST and RST increased substantially only for sample sizes below 40–50 individuals. We consider that the sample sizes employed in the current study (all but one between 45 and 50) give acceptable precision for the statistics employed and for the levels of variability observed. In fact binning of allelic classes to reduce total number of alleles per locus to 10–13 and pooling rare classes results in an increase in pairwise FST values (as would be expected, RST is reduced slightly), so the raw data presented here represent a conservative estimate of among-sample differentiation. Patterns of genetic differentiation among samples may reflect the geographical separation of populations sampled, in that restrictions to gene flow arising from the limited dispersal of individuals results in between-population allele frequency variation increasing with geographical distance, as suggested for Atlantic cod populations (Mork et al., 1985). In the present study Mantel tests (GENEPOP, 2000 replications) indicated no significant relationship between geographical distance and pairwise values of RST (P=0.248) or FST (P=0.386), although there is perhaps a qualitative suggestion of a relationship between genetic distance and geographical location (see Figs. 1 and Fig. 2). Topographic and hydrographic factors, as suggested to contribute to population differentiation in cod (Ruzzante et al., 1998), may be more important in structuring genetic variation among herring populations; for example, differences in water temperature between the Barents Sea and more southerly regions, and the particular characteristics of fjords, may result in differently adapted or physically isolated populations. Finally, behavioural effects, such as adults returning to their natal sites to spawn as suggested for Pacific herring (Hourston, 1982), will promote and maintain genetic differences among populations. McQuinn (1997) has recently argued that patterns of dispersal and spawning behaviour in marine teleosts such as C. harengus that inhabit unstable northern waters are unlikely to arise entirely from hydrographic factors. He argues that the phenotypic differentiation of sympatric spawning groups arises and is maintained not necessarily by natal spawning behaviour, but by social transmission of spatial learning between adults and prespawning individuals, resulting in repeated returns of spawning groups to specific nursery areas. Such an ‘adopted-migrant’ hypothesis forms the underlying mechanism for a postulated metapopulation structuring in herring, where a balance between the high return rates of adults (Wheeler & Winters, 1984) and occasional straying among seasonally sympatric groups results in a mosaic of locally persistent populations with occasional gene flow. The levels of differentiation reported in the current study are not inconsistent with such an ‘adopted-migrant’ hypothesis, although further work involving temporal sampling will be essential to determine persistence of spatial structuring.
Comparison of microsatellite, allozyme and mtDNA RFLP data
Table 3 presents pairwise genetic differentiation (FST) among the same set of samples estimated from allozyme and mtDNA datasets (data from Turan, 1997 and Turan et al., 1998; brief details on number of loci, and gene region/number of restriction enzymes used given in Table 3). Comparison of pairwise FST values from the three different markers (FST is used as RST has no equivalent statistic for allozyme data) suggests that broad patterns and levels of differentiation indicated are comparable among the three datasets, although the mtDNA data are generally less informative. The allozyme data support present patterns revealed by microsatellites in suggesting several levels of divergence among the Atlantic herring: FST values are low among IC1, NS2 and BF (but larger, and significant, differences are indicated by microsatellites); the NS1 sample stands out as the most divergent among the Atlantic samples, with FST values from allozymes 2–3 times higher than those from microsatellites. The mtDNA results indicate no significant structuring among the Atlantic samples, but do show consistently higher FST between NS1 and the rest, and confirm that the Pacific herring sample is the most divergent. Estimates of genetic differentiation between Atlantic and Pacific herring show the most discordant comparisons between the three marker types. Microsatellites indicate that the level of differentiation between Pacific herring and the Atlantic samples (FST=0.027–0.043, mean=0.037) overlaps with that observed within NE Atlantic herring in comparisons involving the Barents Sea spring spawner sample (NS1, FST=0.025–0.037, mean=0.032). By contrast, allozyme and mtDNA data both indicate Pacific herring are far more differentiated from any Atlantic samples (allozymes mean FST=0.743, mtDNA mean FST = 0.124) than seen within Atlantic herring (allozymes mean FST=0.074, mtDNA mean FST=0.003).
What factors might contribute to the observed disparities in divergence indicated by microsatellites, allozymes and mtDNA RFLP? First, the lack of resolution of the mtDNA analysis below interspecific comparisons (where allozymes and microsatellites detected widespread differentiation) most probably results from statistical problems with the extremely high haplotypic diversity observed, with 96 different haplotypes (71 of them unique) in 196 fish screened (Turan et al., 1998). This problem has been noted for mtDNA RFLP data in several pelagic fish species (Hauser & Ward, 1998); replacement of RFLP by direct sequencing of mtDNA has been demonstrated to provide far more informative data at the intraspecific level in some fish (Carr & Marshall, 1991; Grant et al., 1998). The detection of values of FST significantly different from zero among the IC1/NS2/BF group by microsatellites, where allozymes indicated no significant divergence, probably results from the increased power of the microsatellite analysis resulting from higher variability (but with fewer alleles per locus, and more even distribution of frequencies, than the mtDNA). Several other studies (e.g. Estoup et al., 1998; Shaw et al., 1999) have indicated similar patterns, and microsatellite studies have now shown that whiting (Rico et al., 1997), cod (Ruzzante et al., 1998) and Pacific herring (O’Connell et al., 1998) all exhibit small but significant genetic differentiation within populations previously thought to be uniform over regional geographical ranges based on allozyme and mtDNA data. Above this subtle level of population differentiation, divergence (FST) as indicated by allozymes appears to increase more rapidly than by equivalent estimates from microsatellites. The lower microsatellite value probably results from greater homoplasy in this dataset, resulting from the much higher rates of mutation observed for microsatellite loci and allele size range constraints, which is expected to reduce FST values (Goldstein et al., 1995; Slatkin, 1995), and the stepwise mutational process assumed for the type of perfect repeat arrays used in this study (Valdes et al., 1993). A similar result has been found in the squid Loligo forbesi, where the isolated Azores population is extremely divergent from other populations (based on mtDNA sequence divergence): FST estimated from microsatellites is half that estimated from allozymes (0.245 vs. 0.536; Shaw et al., 1999). In contrast, Estoup et al. (1998) found FST values from allozymes and microsatellites in brown trout largely comparable over a wide range of divergence (up to mean FST of 0.279 across drainages). The apparent contradiction here might be explained by differences in population sizes: large effective sizes in herring and squid would promote microsatellite homoplasy by increasing the effect of mutation relative to drift (large number of new, possibly convergent, mutants per generation with low probabilities of loss by random drift) greatly reducing the effects of genetic drift on rate of loss of new mutations; small effective population sizes in trout would promote the opposite. Anecdotal support for these effects may be apparent in the widely differing numbers of alleles detected per locus in herring and brown trout (39 and 3.6, respectively), although differences in repeat type and structure may also be resulting in differences in allelic diversity.
From a similar perspective the level of differentiation implied for the Pacific herring sample, overlapping that observed within Atlantic herring samples, indicates that microsatellites may not be useful for estimating separation and divergence between these species. Studies with mammals and turtles have likewise reported a poor phylogenetic capability of microsatellites, because of small genetic distances observed between species relative to those within species (Garza et al., 1995; Jarne & Lagoda, 1996). Estimates of genetic divergence from microsatellites have been predicted to depart from linearity with increasing time since divergence of taxa, and nonstepwise (IAM-based) estimators are predicted to lose linearity earlier than stepwise (SMM-based) estimators, because of homoplasy resulting from range constraints and mutation rates (Goldstein et al., 1995). Although the data presented here are certainly not extensive enough for a test of such ideas in a pelagic fish, they show some interesting patterns that could be further tested. If allozymes are assumed to give a more representative estimate of true divergence between the samples examined here (particularly between the more differentiated samples), then it appears that both FST and RST estimated from microsatellites similarly lose linearity in the region from moderately divergent populations (NS1 vs. the rest) to closely related species (Pacific vs. Atlantic herring).
In summary, the data obtained from the analysis of four polymorphic microsatellite loci in NE Atlantic herring demonstrate the existence of genetically distinct populations, and indicate the utility of microsatellite markers in detecting genetic differentiation in a highly mobile, pelagic teleost. Moreover, microsatellite DNA markers provide a new perspective on previous estimates of the low levels of genetic differentiation detected in this species using allozyme and mtDNA data. Although genetic differences between populations are small, levels of gene flow (effective migration) indicated would suggest that depletion or extinction of any one of these populations may not be balanced by recruitment from others, at least at a rate likely to offset harvesting pressures, and might result in an irreversible loss of genetic diversity from the gene pool. Thus, in combination with earlier studies (Jørstad et al., 1994; Turan et al., 1998) the present study supports the current status of Icelandic summer-spawners and Barents Sea Norwegian spring-spawners as distinct management units.
References
Bentzen, P., Harris, A. and Wright, J. M. (1991). Cloning of hypervariable minisatellite and simple sequence microsatellite repeats for DNA fingerprinting of important aquacultural species. In: Burke, T., Dolf, G. A., Jeffreys, A. J. and Wolf, R. (eds) DNA Fingerprinting: Approaches and Applications, pp. 243–262. Birkhäuser, Basel.
Brooker, A. L., Cook, D., Bentzen, P., Wright, J. M. and Doyle, R. W. (1994). Organisation of microsatellites differs between mammals and cold-water teleost fishes. Can J Fish Aquat Sci. 51: 1959–1966.
Carr, S. M. and Marshall, H. D. (1991). Detection of intraspecific DNA-sequence variation in the mitochondrial cytochrome-B gene of Atlantic cod (Gadus morhua) by the polymerase chain reaction. Can J Fish Aquat Sci. 48: 48–52.
Dahle, G. and Eriksen, A. G. (1990). Spring and autumn spawners of herring (Clupea harengus) in the North Sea, Skagerrak and Kattegat, population genetic analysis. Fish Res. 9: 131–141.
Debarros, P. and Holst, J. C. (1995). Identification of geographic origin of Norwegian spring-spawning herring (Clupea harengus L.) based on measurements of scale annuli. ICES J Mar Sci. 52: 863–872.
Domanico, M. J., Phillips, R. B. and Schweigert, J. F. (1996). Sequence variation in ribosomal DNA of Pacific (Clupea pallasi) and Atlantic (Clupea harengus) herring. Can J Fish Aquat Sci. 53: 2418–2423.
Estoup, A., Rousset, F., Michalakis, Y., Cornuet, J. -M., Adriamanga, M. and Guyomard, R. (1998). Comparative analysis of microsatellite and allozyme markers: a case study investigating microgeographic differentiation in brown trout (Salmo trutta). Mol Ecol. 7: 339–353.
Excoffier, L., Smouse, P. E. and Quattro, J. M. (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA data. Genetics. 131: 479–491.
Fisher, R. A. (1954). Statistical Methods for Research Workers 12th edn. Oliver & Boyd, Edinburgh.
Fridriksson, A. (1958). The tribes of the north coast herring of Iceland with special reference to the period, 1948–55. Rapp Proces-Verb Réunions; Cons Int Explor Mer Copenhag. 143: 36–44.
Garcíadeleon, F. J., Chikhi, L. and Bonhomme, F. (1997). Microsatellite polymorphism and population subdivision in natural populations of European sea bass, Dicentrarchus labrax (Linnaeus, 1958). Mol Ecol. 6: 51–62.
Garza, J. C., Slatkin, M. and Freimer, N. B. (1995). Evolution and mutation of microsatellites in humans and chimpanzees. Am J Hum Genet. 57: 929–929.
Goldstein, D. B., Ruizlinares, A., Cavalli-Sforza, L. L. and Feldman, M. W. (1995). An evaluation of genetic distances for use with microsatellite loci. Genetics. 139: 463–471.
Goodman, S. J. (1997). RST CALC: a collection of computer programs for calculating unbiased estimates of genetic differentiation and gene flow from microsatellite data and determining their significance. Mol Ecol. 6: 881–885.
Goudet, J. (1996). F-Stat (version 1.2): a computer program to calculate F-statistics. J Hered. 86: 485–486.
Goudet, J., Raymond, M., Demeeus, T. and Rousset, F. (1996). Testing differentiation in diploid populations. Genetics. 139: 463–471.
Grant, W. S. (1986). Biochemical genetic divergence between Atlantic, Clupea harengus and Pacific, Clupea pallasi herring. Copeia. 3: 714–719.
Grant, W. S. and Utter, F. M. (1984). Biochemical genetics of Pacific herring (Clupea pallasi). Can J Fish Aquat Sci. 41: 856–864.
Grant, W. S., Clark, A. M. and Bowen, B. W. (1998). Why restriction fragment length polymorphism analysis of mitochondrial DNA failed to resolve sardine (Sardinops) biogeography: insights from mitochondrial DNA cytochrome b sequences. Can J Fish Aquat Sci. 55: 2539–2547.
Haegele, C. W. and Schweigert, J. F. (1985). Distribution and characteristics of herring spawning grounds and description of spawning behaviour. Can J Fish Aquat Sci. 42 (Suppl.), 39–55.
Hauser, L. and Ward, R. D. (1998). Population identification in pelagic fish: the limits of molecular markers. In: Carvalho, G. R. (ed.) Advances in Molecular Ecology, pp. 191–224. NATO Science Series IOS Press, Amsterdam.
Hedgecock, D. (1994). Temporal and spatial genetic structure of marine animal populations in the California current. California Cooperative Oceanic Fish Invest Reports. 35: 73–81.
Hourston, A. S. (1982). Homing by Canada’s west coast herring to management units and divisions as indicated by tagging recoveries. Can J Fish Aquat Sci. 39: 1414–1422.
Jarne, P. and Lagoda, P. J. L. (1996). Microsatellites, from molecules to populations and back. Trends Ecol Evol. 11: 424–429.
Jørstad, K. E., King, D. P. F. and Nævdal, G. (1991). Population structure of Atlantic herring, Clupea harengus L. J Fish Biol. 39: 43–52.
Jørstad, K. E., Dahle, G. and Paulsen, O. I. (1994). Genetic comparison between Pacific herring (Clupea pallasi) and a Norwegian fjord stock of Atlantic herring (Clupea harengus). Can J Fish Aquat Sci. 51: 233–239.
King, D. P. F., Ferguson, A. and Moffett, I. J. J. (1987). Aspects of population genetics of herring, Clupea harengus around the British Isles and in the Baltic Sea. Fish Res. 6: 35–52.
Kornfield, I. and Bogdanowicz, S. M. (1987). Differentiation of mitochondrial-DNA in Atlantic herring Clupea harengus. Fish Bull. 85: 561–568.
Mcquinn, I. H. (1997). Metapopulations and the Atlantic herring. Rev Fish Biol Fish. 7: 297–329.
Mork, J., Ryman, N., Ståhl, G., Utter, F. and Sundnes, G. (1985). Genetic variation in Atlantic cod (Gadus morhua) throughout its range. Can J Fish Aquat Sci. 42: 1580–1587.
Nei, M. (1972). Genetic distances between populations. Am Nat. 106: 283–292.
O’Connell, M. and Wright, J. M. (1997). Microsatellite DNA in fishes. Rev Fish Biol Fish. 7: 331–363.
O’Connell, M., Danzmann, R. G., Cornuet, J. -M., Wright, J. M. and Ferguson, M. M. (1997). Differentiation of rainbow trout (Oncorhynchus mykiss) populations in Lake Ontario and the evaluation of the stepwise mutation and infinite allele mutation models using microsatellite variability. Can J Fish Aquat Sci. 54: 1391–1399.
O’Connell, M., Dillon, M. C., Wright, J. M., Bentzen, P., Merkouris, S. and Seeb, J. (1998). Genetic structuring among Alaskan Pacific herring (Clupea pallasi) populations identified using microsatellite variability. J Fish Biol. 53: 150–163.
Raymond, M. and Rousset, F. (1995). Genepop (Version-1.2)—Population genetics software for exact tests and ecumenicism. J Hered. 86: 248–249.
Rice, W. R. (1989). Analyzing tables of statistical tests. Evolution. 43: 223–225.
Rico, C., Ibrahim, K. M., Rico, I. and Hewitt, G. M. (1997). Stock composition in North Atlantic populations of whiting using microsatellite markers. J Fish Biol. 51: 462–475.
Rousset, F. and Raymond, M. (1995). Testing heterozygote excess and deficiency. Genetics. 140: 1413–1419.
Ruzzante, D. E. (1998). A comparison of several measures of genetic distances and population structure with microsatellite data: bias and sampling variance. Can J Fish Aquat Sci. 55: 1–14.
Ruzzante, D. E., Taggart, C. T. and Cook, D. (1998). A nuclear DNA basis for shelf- and bank-scale population structure in NW Atlantic cod (Gadus morhua): Labrador to Georges Bank. Mol Ecol. 7: 1663–1680.
Ryman, N., Lagercrantz, U., Andersson, L., Chakraborty, R. and Rosenberg, R. (1984). Lack of correspondence between genetic and morphologic variability patterns in Atlantic herring (Clupea harengus). Heredity. 53: 687–704.
Shaw, P. W., Pierce, G. and Boyle, P. R. (1999). Subtle population structuring within a highly vagile marine invertebrate, the Veined squid Loligo forbesi (Cephalopoda: Loliginidae) uncovered using microsatellite DNA markers. Mol Ecol. 8: 407–417.
Slatkin, M. (1995). A measure of population subdivision based on microsatellite allele frequencies. Genetics. 139: 457–462.
Smith, P. J., Jamieson, A. and Birley, A. J. (1990). Electrophoretic studies and the stock concept in marine teleosts. Cons Int Explor Mer. 47: 231–245.
Stenevik, E. K., Fossum, P., Johannessen, A. and Folkvord, A. (1996). Identification of Norwegian spring spawning herring (Clupea harengus L.) larvae from spawning grounds off western Norway applying otolith microstructure analysis. Sarsia. 80: 285–292.
Svetovidov, A. N. (1963). Fauna of U.S.S.R. Fishes, 2. Clupeidae Israel Program for Scientific Translations, Jerusalem.
Taggart, J. B., Hynes, R. A., Prodöhl, P. A. and Ferguson, A. (1992). A simplified protocol for routine total DNA isolation from salmonid fishes. J Fish Biol. 40: 963–966.
Turan, C. (1997). Population Structure of Atlantic Herring Clupea harengus L., in the Northeast Atlantic using Phenotypic and Molecular Approaches Ph.D. Thesis, University of Hull, Hull.
Turan, C., Carvalho, G. R. and Mork, J. (1998). Molecular genetic analysis of Atlanto-Scandian herring (Clupea harengus) populations using allozymes and mitochondrial DNA markers. J Mar Biol Ass UK. 78: 269–283.
Valdes, A. M., Slatkin, M. and Freimer, N. B. (1993). Allele frequencies at microsatellite loci: the stepwise mutation model revisited. Genetics. 133: 737–749.
Waples, R. S. (1998). Separating the wheat from the chaff: patterns of genetic differentiation in high gene flow species. J Hered. 89: 438–450.
Weir, B. S. and Cockerham, C. C. (1984). Estimating F-statistics for the analysis of population structure. Evolution. 38: 1358–1370.
Wheeler, J. P. and Winters, G. H. (1984). Homing of Atlantic herring (Clupea harengus harengus) in Newfoundland waters as indicated by tagging data. Can J Fish Aquat Sci. 41: 108–117.
Acknowledgements
We thank Dr J. Jakobsson and Ms A. Gardarsdottir for providing Icelandic herring samples, Professor J. Mork for Norwegian samples and Dr Doug Hay for the Pacific herring sample. We gratefully acknowledge the allocation of a studentship (C.T.) and financial support from the T. C. University of Mustafa Kemal, Faculty of Fisheries, Iskenderun, Hatay in Turkey.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shaw, P., Turan, C., Wright, J. et al. Microsatellite DNA analysis of population structure in Atlantic herring (Clupea harengus), with direct comparison to allozyme and mtDNA RFLP analyses. Heredity 83, 490–499 (1999). https://doi.org/10.1038/sj.hdy.6885860
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6885860
Keywords
This article is cited by
-
The current knowledge status of the genetic population structure of the European sardine (Sardina pilchardus): uncertainties to be solved for an appropriate fishery management
Reviews in Fish Biology and Fisheries (2022)
-
Genetic diversity and population structure analysis of Lateolabrax maculatus from Chinese coastal waters using polymorphic microsatellite markers
Scientific Reports (2021)
-
Genetic mapping and QTL analysis in European hazelnut (Corylus avellana L.)
Molecular Breeding (2016)
-
Development and characterization of nine novel microsatellite markers for the milkfish Chanos chanos
Conservation Genetics Resources (2015)
-
Population genetic structure of the sea bass (Lateolabrax japonicus) in Korea based on multiplex PCR assays with 12 polymorphic microsatellite markers
Genes & Genomics (2014)