Abstract
Linking variation in quantitative traits to variation in the genome is an important, but challenging task in the study of life-history evolution. Linkage maps provide a valuable tool for the unravelling of such trait−gene associations. Moreover, they give insight into recombination landscapes and between-species karyotype evolution. Here we used genotype data, generated from a 10k single-nucleotide polymorphism (SNP) chip, of over 2000 individuals to produce high-density linkage maps of the great tit (Parus major), a passerine bird that serves as a model species for ecological and evolutionary questions. We created independent maps from two distinct populations: a captive F2-cross from The Netherlands (NL) and a wild population from the United Kingdom (UK). The two maps contained 6554 SNPs in 32 linkage groups, spanning 2010 cM and 1917 cM for the NL and UK populations, respectively, and were similar in size and marker order. Subtle levels of heterochiasmy within and between chromosomes were remarkably consistent between the populations, suggesting that the local departures from sex-equal recombination rates have evolved. This key and surprising result would have been impossible to detect if only one population was mapped. A comparison with zebra finch Taeniopygia guttata, chicken Gallus gallus and the green anole lizard Anolis carolinensis genomes provided further insight into the evolution of avian karyotypes.
Similar content being viewed by others
Introduction
In the last decade, linkage mapping projects in the bird species have been extended from studies of agriculturally important organisms, such as chickens (Groenen et al., 2000), to a much more diverse set of species including other galliformes such as the turkey (Reed et al., 2007), anseriformes such as the mallard duck (Huang et al., 2006) and several passerine species including the collared flycatcher (Backström et al., 2008), the blue tit (Hansson et al., 2010), the zebra finch (Stapley et al., 2008; Backström et al., 2010) and the great reed warbler (Hansson et al., 2005). Although the most comprehensive whole-genome maps have been constructed for galliformes it is notable that passerines, by far the largest order in the class Aves, represent the majority of other species for which such resources have been developed. Linkage maps have been constructed with two main purposes in mind: (1) to identify regions of the genome that are responsible for genetic variation in quantitative traits, that is, to map quantitative trait loci and (2) to understand the genomic features and evolution of avian karyotypes. Often, however, particular species are well suited to investigating both types of question. In this manuscript we describe linkage maps of the great tit (Parus major) a species that is widely studied in ecology and evolutionary biology (see below).
A motivation for constructing linkage maps in wild bird species, especially passerines, which have been the focus of many ecological and evolutionary studies (Bennett and Owens, 2002), is to provide understanding of how evolution operates in natural populations. Quantitative trait loci mapping can unravel knowledge of the molecular genetic basis of ecologically relevant traits, which are usually expected to have a complex genetic architecture (Slate, 2005). Although linkage mapping is being superseded by genome-wide association studies, an approach that does not directly require a linkage map, linkage maps remain useful tools for identifying the chromosomal location and order of markers used in a genome-wide association studies.
In addition to understanding the genetic basis of ecologically important traits, linkage maps provide valuable insight into the evolution of genome organisation. For example, they help to reveal and explain the recombination landscape across the genome, how recombination rates might differ between the sexes and how species are characterised by structural genomic changes. Studies of bird genome evolution have identified a number of interesting features. First, bird karyotypes are remarkably highly conserved, with the majority of species having a haploid number of 38–42 chromosomes, and most species showing conserved synteny (Kayang et al., 2006; Backström et al., 2008; Stapley et al., 2008; Ellegren, 2010). Second, recombination is usually not evenly distributed between and within chromosomes. Macrochromosomes tend to have lower recombination rates (per Mbp) than microchromosomes, presumably because all chromosomes have an obligate crossing-over during meiosis, even if they are physically short (Groenen et al., 2000; Stapley et al., 2008). Within macrochromosomes of zebra finches, recombination rates are lower at the centre relative to telomeric regions (Stapley et al., 2008; Backström et al., 2010; Stapley et al., 2010), although this phenomenon has not been observed to the same extent in chickens (Groenen et al., 2009). Third, some bird species display heterochiasmy, that is, a difference in recombination rates between sexes.
Heterochiasmy was first believed to reflect pleiotropic consequences of selection against recombination between the sex chromosomes (the Haldane−Huxley rule) (Haldane, 1922; Huxley, 1928). More recently, several evolutionary hypotheses have been developed, where generally differences in recombination rates between sexes are expected to be due to sex differences in selection pressures. For example, in mammals recombination in males is commonly found to be lower than in females (Popa et al., 2012), possibly to reduce the possibility of recombination breaking up favourable gene combinations in the sex with more intense selection (Trivers, 1988). Other possible evolutionary explanations for heterochiasmy are that the sexes have different selective optima to balance the deleterious mutation rate and the cost of efficient repair (Ellegren, 2007). However, pronounced heterochiasmy is not a general feature in all animals and is certainly not ubiquitous in birds (Stapley et al., 2008; Groenen et al., 2009). The most striking example has been observed in the great reed warbler, where a female:male map length ratio of 2.15 was initially observed (Hansson et al., 2005), although a subsequent study with more markers revised this down (female:male ratio=1.56; Åkesson et al., 2007). In the blue tit recombination rates are slightly greater in females (female:male ratio=1.07; Hansson et al., 2010) and in a study of turkey the opposite pattern was found: (female:male ratio=0.86; Aslam et al., 2010). Apart from the overall differences in total genome map length for males and females, substantial variation in recombination rates between sexes may also exist within and between autosomes of the same species (Dunn and Bennett, 1967; Robertson, 1984). For example, in humans, levels of heterochiasmy are associated with the physical location on the chromosome relative to the centromere, where recombination rates tended to be female biased near the centromere and male biased towards the telomeres (Broman et al., 1998), with a possible relationship between heterochiasmy and GC content (Popa et al., 2012). In birds, levels of heterochiasmy have not been studied at either the within- or between-chromosome scale, which are essential starting points if trying to understand how sexual selection may cause variation in recombination rates (Ellegren, 2007).
In most species, especially of natural populations, mapping studies have been conducted on single populations only (but see Poissant et al., 2010 for an exception, albeit with a modest number of markers). In order to make further progress in understanding bird genomes, high-density linkage maps, replicated across populations of the same species are highly desirable. The initial observation that gene order was highly conserved over large evolutionary distances, for example, between passerines and galliformes, was shown to be only partially true once higher density maps with sufficient resolution to detect minor within-chromosome rearrangements became available (Backström et al., 2006; Stapley et al., 2008; Backström et al., 2010). However, alternative marker orders often have similar statistical probabilities (likelihoods) and so determining whether subtle differences in gene order are real is not straightforward. If a particular marker order is identified in two independent populations of the same species, the evidence for rearrangements becomes much greater. Similarly, the replication of linkage maps in two populations can help understand whether heterochiasmy is present on a (genomically) local scale. If particular regions of the genome display apparent heterochiasmy, then without replication, it is difficult to rule out random, non-repeatable, sex-biased sampling of the location of recombination events as the cause, rather than a conserved feature of a species’ genome. Constructing maps in two independent populations overcome this problem, because observing similar patterns in both populations provide strong support for true species-level differences in male and female recombination rates.
The great tit, a passerine of the family Paridae, is a model organism for evolutionary and ecological research in wild populations, with a wide body of work, addressing topics such as life history evolution (Garant et al., 2005; Postma and van Noordwijk, 2005), parasite infestation (Norris et al., 1994; Heeb et al., 1999), climate change (Visser et al., 1998; Charmantier et al., 2008), plumage characteristics (Evans and Sheldon, 2012), song (Slabbekoorn and Peet, 2003; Naguib et al., 2010) and personality (Dingemanse et al., 2002; Drent et al., 2003; Quinn et al., 2009; van Oers and Naguib, 2013). The development of a genome-wide linkage map is, therefore, essential to help shed light on the link between genotypic and phenotypic variation in such studies. Using chromosome painting, the karyotype of great tit has been estimated at 2n=80 (Nanda et al., 2011), similar to other passerines including the zebra finch (Itoh and Arnold, 2005). The zebra finch and great tit diverged from a common ancestor 40–45 million years ago (Barker et al., 2004), but given the highly conserved nature of avian karyotypes, a high degree of shared synteny between zebra finch and great tit genomes is expected. As with most bird species, great tit chromosomes vary in size (Nanda et al., 2011), over a range of almost two orders of magnitude. Around 10 large macrochromosomes typically have similar sizes to most mammal chromosomes and in addition there are numerous smaller microchromosomes (Ellegren, 2010).
In this study, we describe the construction and analysis of two high-density whole-genome linkage maps of the great tit. To construct the maps we used a set of SNP markers originating from a 9193 SNP chip (van Bers et al., 2012). We genotyped a captive F2 inter-cross population from selected lines on high and low early exploratory behaviour (Drent et al., 2003; van Oers et al., 2004) at the Netherlands Institute of Ecology (NIOO-KNAW), NL and birds from a wild pedigreed long-term study population (Wytham Woods) near Oxford, UK. Producing independent replicate maps for separate populations should (i) give insight into how accurate/robust linkage maps are, (ii) allow tests of whether variation in recombination rates across a chromosome are consistent across populations, (iii) allow tests of whether small differences in recombination rates between sexes (heterochiasmy) are chance departures from 1:1 or are consistent across populations and (iv) allow tests of whether minor rearrangements between bird species are genuine or an artefact of minor errors caused by alternative marker orders having very similar likelihoods. It will, thus, provide more robust insight into avian karyotype evolution, if within-species patterns are demonstrated to be repeatable.
Materials and methods
SNP genotyping
A total of 4702 birds were genotyped with an Illumina iSelect BeadChip containing 9193 SNPs, according to the manufacturers’ protocol (van Bers et al., 2012). Genotyping results were analysed using Genome Studio software (Illumina, San Diego, CA, USA). A total of 8340 SNPs were successfully genotyped in both the NL and the UK population, but only 7215 and 7203 of these SNPs were polymorphic in the NL and UK population, respectively. After excluding markers with very low minor allele frequency (MAF<0.01), 6897 and 7032 SNPs were used for map construction in the NL and UK populations, respectively. Detailed descriptions of SNP identification, SNP selection and genotyping results are given elsewhere (van Bers et al., 2010; Santure et al., 2011; van Bers et al., 2012).
Study species and mapping populations
For the construction of the linkage maps, we used 398 great tits originating from a captive population at the NIOO-KNAW, Wageningen, NL and 1656 individuals from a pedigreed long-term natural population (Wytham Woods) near Oxford, UK. The genotyped individuals in the NL population were selected from 480 genotyped individuals of the fourth generation of artificial selection on fast and slow early exploratory behaviour (Drent et al., 2003) and the F1 and F2 intercrosses between the fast and slow lines (van Oers et al., 2004). The final mapping pedigree (N=398) included all F2 intercross individuals (N=251), their F1 parents, the siblings of these F1 parents and all full-sib selection line families with their parents. These individuals were manually split into 35 subfamilies, with mean family size 15.0 (range 5–35). Sex and parentage of all individuals was confirmed from the genotypes. Owing to a small number of parent-offspring genotype mismatches, we removed five individuals from the data set, leaving 393 individuals for analysis (Supplementary Methods).
A subset of 1656 Wytham individuals with confirmed first degree pedigree links (genotyped parents and/or sibs and/or offspring) and confirmed sex were chosen for constructing the UK great tit linkage map (see Supplementary Methods for more details). For the UK population, complex pedigrees were first split into 61 subfamilies (of up to approximately 50 individuals and three generations) using the CRIGEN command (options: -size 50 and -gen 3) in Crimap (see below). Parent-offspring genotype mismatches were removed from the data set.
Predicted locations by comparative mapping
To identify and understand rearrangements in avian genomes, it is necessary to compare the positions of the homologues of typed markers on different species’ genomes. Recently, the genome of the green anole lizard A. carolinensis was sequenced and assembled (Alfoldi et al., 2011). Here we examined chromosomal rearrangements between great tit, zebra finch, chicken and the green anole; the latter provides a useful out-group to the avian species. The position of the great tit SNPs on the zebra finch, chicken and green anole lizard genomes was predicted based on the alignment of 23 to 150 bases of flanking sequence to the respective genome sequence using the default parameters in blastn (with setting a minimum expectation value (e-value) of 10−5). The three genome sequences used for comparative mapping were downloaded from the Ensembl Genome Browser; zebra finch: version Taeniopygia_guttata.taeGut3.2.4.61, www.ensembl.org/Taeniopygia_guttata/Info/Index; chicken: version Gallus_gallus.WASHUC2.56, www.ensembl.org/Gallus_gallus/Info/Index and green anole lizard: version Anolis_carolinensis.AnoCar2.0.61, www.ensembl.org/Anolis_carolinensis/Info/Index. Predicted physical map locations are the midpoints between the start and end of the mapping alignment and are, therefore, expected to be within 100 bases of the actual SNP position. Linkage groups and putative chromosomes were orientated according to the orientation of the chicken chromosomes.
Map construction
For linkage mapping we used a modified version (2.503) of the CriMap software (Green et al., 1990). We chose to present maps with three different marker densities in this manuscript: (i) framework maps, incorporating markers that could be assigned a most likely order with LOD>3, that is, the best order is 103=1000 times more likely than the next best alternative order (ii) comprehensive maps, where ‘non-framework’ markers were added by lowering levels of stringency (LOD>0.1) and (iii) parsimonious maps, adding as many markers as possible to the comprehensive map with LOD<0.1 (Supplementary Methods).
We applied two different initial strategies for linkage group construction. For the NL map, we assigned 6877 SNPs to putative great tit chromosomes based on sequence homology between SNP flanking sequences and the zebra finch genome sequence. For the UK map, we used the TWOPOINT output of CriMap to create linkage groups with the AUTOGROUP command (see Supplementary Methods for more details). In both mapping populations, the second allele of SNPs on the sex chromosome (Z), was assigned a dummy value (6) in females, to account for the hemizygosity of the female Z chromosome.
Heterochiasmy
Heterochiasmy was examined on three different scales, using the sex-specific maps option in CriMap (see section Map Construction). First, the genome-wide male and female total map lengths were compared. Second, male and female map lengths were estimated for each chromosome. Third, local levels of heterochiasmy were estimated using sex-specific map lengths in a window size of 20 markers and then sliding the window in five marker increments. For this analysis, markers common to comprehensive maps in both populations were used: Parus major (PMA) linkage groups PMA1, 1A, 2–4, 4A, 5–15, 17–21, 23, 24 and 26 contained enough shared comprehensive map markers to estimate local heterochiasmy. Following Poissant et al. (2010), heterochiasmy was measured using the size dimorphism index (SDI) developed by Lovich and Gibbons (1992). This index is considered the best estimator of sexual size dimorphism because, in contrast to ratios, it does not suffer from the fact that heterochiasmy is bound at zero in one sex and unbounded in the other sex. The SDI was obtained with the following formula:

Following convention (Lovich and Gibbons, 1992; Poissant et al., 2010), estimates were then made positive when the female value was largest and negative when the male value was largest. To distinguish between whole-chromosome/whole-genome heterochiasmy and local heterochiasmy, we use the notation SDI and SDI_local, respectively.
Statistical analysis
We used Pearson’s correlations to compare chromosome sizes between populations. We tested for the presence of genome-wide sexual dimorphism using sign tests, with the SDI of each chromosome being one data point, and tested the relationship in SDI between populations using reduced major axis regression (NLRA in PASW Statistics for Mac Version 18.0.3, SPPS Inc., Chicago, IL, USA). To calculate whether linkage group and interval SDI values fell outside the distribution, we conducted an outlier analysis. The critical values to treat a data point as an outlier were calculated by multiplying the difference between the 25% (Q1) and the 75% (Q3) quartiles by 2.2 and adding this to Q3 or subtracting it from Q1 (Hoaglin et al., 1986). Data points were classified as outliers, when they were either larger than the Q3 or small than the Q1 critical value. To analyse whether local heterochiasmy was conserved between the two populations the auto- and cross-correlation functions implemented in R version 2.13 (R Development Core Team, 2011) were used. All other statistical analyses were conducted using PASW Statistics for Mac version 18.0.3 (SPSS Inc.).
Results
Initial assignment of markers into linkage groups
For the NL population (N=393 birds), 6877 markers (mean informative meiosis±s.d.=224.6±94.4) were grouped into 32 putative great tit chromosomes based on the alignment of markers with the zebra finch genome. In the UK population (N=1656 birds), 6094 markers (mean informative meiosis±s.d.=408.0±124.2) were grouped into the same 32 putative great tit chromosomes, using AUTOGROUP on the set of autosomal SNPs. The grouping of markers by the AUTOGROUP option in CriMap demonstrated a remarkable conservation of synteny between the great tit and zebra finch genomes. Based on the very high proportion of markers within each group that aligned to a single zebra finch chromosome (range 91–100%; mean=99.7), chromosome identities could be assigned unambiguously to each linkage group from homology with the zebra finch genome. We will, therefore, refer to the homologous chromosome number when referring to the linkage groups throughout the paper. This follows the convention of Stapley et al., (2008), who assigned zebra finch linkage group names based on homology with the chicken assembly. Thus, great tit, zebra finch and chicken chromosome names are consistent. This method is in contrast to the historic approach of naming chromosomes in order of decreasing physical size. For example, from fluorescence in situ hybridization paints in great tits, PMA2 is known to be larger than PMA1 (Nanda et al., 2011).
Linkage map characteristics
The NL and UK framework maps (LOD⩾3) contained 1766 and 1674 markers, respectively. Of these, 898 markers were present on both maps. When comparing the two maps, 37 out of these 898 markers had a position on both maps that could not be predicted from the position from the other map (Supplementary Figures 1 and 2). The mean interval between the position of these markers on the NL map and the position based on the UK map was 1.91 cM (range 0.02–5.40 cM).
The NL and UK framework maps spanned 1869 cM and 1893 cM, respectively, across 32 linkage groups, homologous to TGU1 to TGU15, TGU17 to TGU28, TGUZ and the zebra finch linkage group LGE22 (Table 1). These linkage groups are homologous to GGA1 to GGA15, GGA17 to GGA28, GGAZ and two chicken linkage groups, LG2 and LGE22C19W28_E50C23. Markers that aligned with chicken chromosomes GGA1, GGA4 and GGA25 were assembled into two distinct linkage groups (PMA1 and PMA1A, PMA4 and PMA4A, and PMA25A and PMA25B) in both great tit populations. Merging the paired linkage groups into one resulted in a distance larger than 50 cM between the two groups of markers. For PMA25A/PMA25B, the two markers that were positioned closest together on the zebra finch genome, but were assigned to different linkage groups (PMA25A and PMA25B) were only 150k base pairs apart on the zebra finch genome. One pair of markers, M3554 from PMA25A and M6440 from PMA25B did have a twopoint LOD score of 2.2 in the NL population, but there was no evidence for linkage between this marker pair in the UK population. Furthermore, no other markers pointed to a possible linkage between PMA25A and PMA25B, suggesting that the two linkage groups may represent two different chromosomes.
The comprehensive maps contained 2853 markers spanning 1867 cM and 3162 markers spanning 1889 cM for the NL and UK populations, respectively. For the parsimonious map, a total of 5855 SNPs were placed on the NL map spanning 2010 cM (Supplementary Figure 1), whereas the UK map consisted of 4878 SNPs spanning 1917 cM (Supplementary Figure 2). This corresponds to an increase in size of 7.7% (NL) and 1.5% (UK) compared with the framework maps. In total 6554 unique SNP markers were present on either the NL map or the UK map. Of these, 4189 were present on both the NL and the UK map, whereas 1666 SNPs were unique to the NL map and 699 markers were present on the UK map and not on the NL map. The MAF for the 1666 markers present on the NL map, but not on the UK map (NL specific: mean±s.e.m.; 0.339±0.003), was significantly lower (t5140=4.16, P<0.001) compared with the MAF of markers present on both maps (present on both: mean±s.e.m.; 0.352±0.001). The MAF for the markers that were present on the UK map, but not on NL map did not have significantly lower MAF (UK specific: mean±s.e.m.; 0.345±0.003) than those on both maps. The number of SNPs per linkage group ranged from 5 (PMALGE22) to 664 (PMA2) for the NL population and from 3 (PMALGE22) to 624 (PMA2) for the UK population. There were no marker inconsistencies at linkage group level (that is, no marker was present on a specific linkage group on one map and present on another linkage group on the other map). Within the linkage groups, we found high consistency between the marker orders in the two populations (Supplementary Figure 3), the only rearrangements being apparent were local inversions involving a small number of markers (see discussion). Recombination rates (measured as cM/Mbp) tended to be greater at the ends of macrochromosomes than in the central parts, although unlike the situation in captive zebra finch populations, recombination ‘deserts’ were not observed in the middle of macrochromosomes (Supplementary Figure 4).
Linkage group sizes ranged from 4.58 cM to 138.15 cM in the NL population and from 3.68 cM to 139.88 cM in the UK population (Table 1), and these sizes were highly correlated between the populations (Figure 1; rp=0.97, n=32; P<0.001), indicating that the recombination rates were not population dependent, and did not differ systematically between the wild and recently-founded captive populations studied here. Based on the alignment of the flanking sequences of the most distal SNPs in great tit linkage map to the zebra finch genome, the total genome coverage was 97.1% (range 77.7–99.8 for individual linkage groups) for the NL map and 98.3% (range 28.4–99.8) for the UK map (Table 1). Assuming that our linkage map covers 32 of the expected 40 chromosomes (that is, taking PMA25A, PMA25B and LGE22 as a separate great tit chromosome), and that these three and the eight missing microchromosomes have at least one obligate recombination event during meiosis, implying a map size of at least 50 cM per chromosome, the total predicted map sizes are ∼2700 cM for the NL and ∼2600 cM for the UK populations.
Heterochiasmy
Genome wide, there was no evidence for large sex differences in map length. We found no significant levels of heterochiasmy (SDI in the NL or the UK population (Table 1; NL: x̄=−0.070, 95% C.I.=−0.166−0.0069, t27=−1.61, P=0.12; UK: x̄=−0.007, 95% C.I.=−0.067−0.051, t27=−0.21, P=0.84). At the chromosomal level SDI levels correlated strongly between populations (reduced major axis regression, y=0.72x+0.04, r2=0.57, F1,26=37.07, P<0.001; Figure 2). An outlier analysis showed that the sex-specific linkage maps of three linkage groups in the NL population significantly differed from each other: males in PMA2, PMA3 and PMA13 had longer map lengths compared with females in the NL population. Although not significant, these three linkage groups also had the lowest SDI values (that is, greater recombination rates in males) in the UK population.
Comparison of levels of heterochiasmy per linkage group for the NL and the UK populations. Heterochiasmy is indicated as the size dimorphism index (SDI). Negative SDI values indicate larger male linkage groups compared with female linkage groups. The solid line represents the y=x line, the dotted line the outcome of the reduced major axis regression (y=0.72x+0.04). Only linkage groups larger than 10 cM are plotted. Linkage groups that deviate significantly from 1:1 are indicated with *.
Because replicated maps from independent populations were constructed, it was possible to examine whether subtle, more localised differences in heterochiasmy were conserved. Most revealingly, the plots (Figure 3) and analyses of SDI_local revealed fine-scale heterochiasmy that was remarkably conserved between the two populations. Cross-correlations, in SDI_local, between the two populations were positive at all 22 chromosomes considered (range in r=0.05–0.999; median r=0.668; Supplementary Table 1). A sign test on all 22 correlations being positive is highly significant (P=4.8 × 10−7, assuming a null expectation of a correlation being equally likely to be positive or negative). At 17 of the 22 chromosomes the cross-correlation would be regarded as significantly positive, if each window was regarded as an independent data point. Importantly, the overall pattern of local heterochiasmy being highly conserved across the two populations was not dependent on window size or the interval from one marker to the next (See Supplementary Figures 5a–c, for SDI_local plots using alternative window sizes).
Size Dimorphism Index (SDI) calculated for windows of 20 SNP markers on the NL (orange) and UK (blue) framework maps. Subsequent windows were chosen by sliding the window five SNP markers along the linkage group. Positive SDI indicates that female recombination rates within the 20 SNP window was higher; negative values indicate greater recombination rates in males. Only linkage groups with at least three windows are plotted.
Comparative mapping
For 5396 of the 5855 SNPs that mapped on both the NL and the UK parsimonious linkage map, we found homology to the zebra finch genome; 1231 of the SNPs could be assigned locations on the chicken genome and 169 were aligned to the green anole lizard genome.
In general, gene order was conserved within syntenic blocks when comparing the map positions on the great tit parsimonious map with the physical position on the zebra finch (Supplementary Figure 6) and the chicken genome (Supplementary Figure 7). Two linkage groups (PMA8 and PMA19) did not show any intrachromosomal rearrangements within either the zebra finch–great tit or the chicken–great tit comparison, that is, marker order was exactly the same in great tit, zebra finch and chicken (see example in Figure 4a). For four linkage groups (PMA5, PMA9, PMA12 and PMA14) chromosomal rearrangements were similar whether comparing great tit to zebra finch or to chicken (see Figure 4b). These cases represent rearrangements that have occurred in the great tit lineage since great tits and zebra finches diverged. In four cases, rearrangements were only visible for the TGU-PMA comparison (PMA2, PMA10, PMA17 and PMA18) but not the GGA-PMA comparison (see Figure 4c). These represent rearrangements in the lineage leading to zebra finches after the great tit-zebra finch divergence. In five linkage groups (PMA4, PMA6, PMA13, PMA15 and PMA20) rearrangements were present only in the GGA-PMA comparison (see Figure 4d), and represent rearrangements that occurred after galliformes and passerines diverged, but before the divergence of great tits and zebra finches. The remaining linkage groups have undergone inversions in the great tit compared with either chicken or zebra finch, but more complex rearrangements were also observable. For example, in PMA1, represented by TGU1 and the distal part of GGA1, two inversions and a third inversion within the second inversion were present in the TGU-PMA comparison, although a rearrangement of ∼30 Mb is also present in the GGA-PMA comparison (Figure 4e). In another example, PMA7, an inversion of ∼8 Mb is present in the GGA-PMA comparison, although in the TGU-PMA comparison a large inversion of ∼25 Mb is also present in the central region of the chromosome (Figure 4f). Inference of ancestral states is more complicated for these cases, even when high-density maps from multiple passerine species are available.
Comparative maps for six example great tit linkage groups, with the great tit linkage map positions plotted against the predicted physical position on the zebra finch (TGU; left panels) and chicken (GGA; right panels) genomes. (a) PMA19 is highly conserved across avian lineages; no rearrangements relative to TGU or GGA, (b) PMA14 is an example of a rearrangement specific to the great tit lineage; an inversion relative to marker order in both TGU and GGA, (c) PMA10 is an example of a TGU-specific rearrangement; an inversion between PMA and TGU, (d) PMA6 is an example of a difference between passerines and galliformes; rearrangement relative to GGA but not TGU, (e) and (f) are chromosomes that are generally evolutionarily less stable across avian genomes, where (e) PMA1; inversions compared with TGU, and both inversions and rearrangements compared with GGA, and (f) PMA7; distinct inversions when PMA is compared with TGU and GGA.
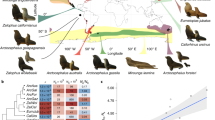
Out of the 32 great tit linkage groups, 22 contained at least one SNP that could be assigned a location on the green anole lizard genome. Thirteen of these linkage groups had shared syntenic segments, with at least two SNP markers linked on both a green anole lizard chromosome and a great tit linkage group (Figure 5). PMA1 and PMA1A aligned to different green anole lizard chromosomes, suggesting that a single chromosome 1/1A in chickens is a derived state.
Great tit-green anole lizard (Anolis carolinensis; ACA) synteny map. The horizontal bars represent the six A. carolinensis macrochromosomes ACA1–ACA6 and three microchromosomes LGa, LGb and LGc, to which aligned SNPs or blocks of SNPs could be ordered. Each coloured block represents a great tit (PMA) chromosome as is indicated in the lower part of the graph. Adjusted from (Alfoldi et al., 2011).
Discussion
In this paper we show that genetic maps are highly conserved between two great tit populations, using the same genetic markers, but different types of mapping population and different map-building methods. Owing to the construction of replicated independent genetic maps we were able to detect differences in recombination rate between sexes at a scale that has not previously been described in any bird species. Independent linkage maps also meant that genuine chromosomal rearrangements between species could be distinguished from alternative marker orders with similar likelihoods.
Heterochiasmy
According to the Haldane–Huxley rule, the recombination rate should be reduced in the heterogametic sex, that is, female maps are expected to be shorter than male maps in birds (Haldane, 1922; Huxley, 1928). Our results add to a number of birds where whole-genome map sizes between males and females do not differ, at least not dramatically (Stapley et al., 2008; Groenen et al., 2009). However, despite the relatively little genome-wide difference in male and female recombination rates, our results show that heterochiasmy is present at a more subtle, yet consistent scale both between and, especially, within chromosomes. Importantly, these findings are repeatable across populations, indicating that they are real features of the great tit genome rather than sex-biased random sampling of where recombination events were observed in the genome.
It is possible that the relative position of the centromere is important with respect to local heterochiasmy. Unfortunately, we do not have information on centromere positions, and can, therefore, only speculate on this. Several studies have shown that recombination rates in birds are lowest around the centromeres, and that they increase towards the telomeres (see for example, Backström et al., 2010), though these patterns have not previously been shown to differ between the two sexes. One intriguing possibility that requires further exploration is that local variation in heterochiasmy is related to the location of sexually antagonistic loci. It is well known that directional selection on any trait can result in an increase in recombination rate (see Otto and Lenormand, 2002). Therefore, conserved heterochiasmy on a local scale could indicate regions of the genome where selection is stronger in one sex than the other. For example, loci that explain variation in male fitness might be expected to be in regions of male-biased recombination rate and vice-versa. Measuring the fitness consequences of individual loci in wild populations remains notoriously difficult, but is perhaps possible in populations with extensive life history data and rapidly developing genomics resources, such as the great tit. The discovery of subtle, yet real, sex differences in recombination rates illustrates the advantages of replicating mapping studies in two independent mapping populations.
Population differences
Overall, our results show that the linkage maps of the two populations are very similar. We found no evidence that positions of groups of consecutive markers differed between the two populations, indicating no gene order differences between the populations. We did find order differences between single markers in 10 linkage groups, which are most likely due to power issues in assigning correct orders on a small scale rather than true rearrangements between the populations. Slightly different marker orders typically produce very similar likelihoods and total map lengths when there are insufficient recombination events to identify the true order. In domestic chicken, Groenen et al. (2009) compared maps between three mapping populations typed with the same SNP set. Though small differences were found, the authors explained this by differences in the data structure between the populations, causing variation in information content of the different markers, rather than assuming real population differences in gene order.
We used two different methods of assigning SNPs to chromosomes. AUTOGROUP and in silico mapping by BLASTing to the zebra finch genome produced near-identical linkage maps for the two populations, indicating that for avian species both these methods are robust, given a reasonably closely related genome for comparative mapping. Studies that cannot rely on reference genomes will, therefore, not be hampered by a lack of a priori information on linkage information when building maps and will most likely perform better when using the AUTOGROUP method. A disadvantage of using a reference genome for assigning markers to expected chromosomes is that blast errors may complicate the building process. Moreover, markers with low information content are unlikely to be included when assigning markers to linkage groups using AUTOGROUP, increasing build speed and accuracy. Overall, we were able to incorporate marginally more markers in the NL map (using BLAST) compared with the UK map (using AUTOGROUP).
Map lengths
The extrapolated complete genetic map sizes of 2600 and 2700 cM, are similar to map lengths of other passerines like the zebra finch (Stapley et al., 2008; Backström et al., 2010), blue tit Cyanistes caeruleus (Hansson et al., 2010) and Siberian Jay Perisoreus infaustus (Jaari et al., 2009), but they are smaller than those of chicken or turkey (∼3200 cM; Groenen et al., 2009; Aslam et al., 2010) and larger than map sizes of the great reed warbler (∼2000 cM; Åkesson et al., 2007), though this map was of lower density, probably underestimating total map length. This further demonstrates that passerines have a reduced recombination relative to domesticated galliformes (Stapley et al., 2008), as genome sizes of passerines are in general comparable with chicken, at around 1.1–1.2 Gbp (Bachmann et al., 1972; Andrews et al., 2009). Recombination is often not evenly distributed within chromosomes and in birds is often more pronounced near the telomeres. Therefore, map sizes could be underestimated when marker density is low and distal parts of chromosomes are poorly covered. The large number of markers used here (relative to other passerine maps), means that differences in map length between passerines and chickens cannot be attributed to differences in genome coverage. One possible explanation for differences in map length is that artificial selection during domestication has led to an increased recombination rate in chickens (Otto and Lenormand, 2002; Groenen et al., 2009). We found slightly larger map lengths in our captive F2-cross NL population compared with the wild UK population; these differences were, however, small compared with the between species differences, and were possibly due to the higher number of markers in the NL map, leading to inclusion of lower quality markers rather than an effect of the few generations of selection. Previous studies have demonstrated that adding error-prone markers tends to increase overall genetic map lengths (Groenen et al., 2000, 2009; Ball et al., 2010). Without maps of wild galliformes being constructed, it remains unknown whether the passerine-galliform difference is a result of domestication of chickens and turkeys or an ancestral difference between passerines and galliformes that predates domestication.
Interchromosomal rearrangements
We found only one putative interchromosomal rearrangement when comparing our map with other whole-genome high-density avian linkage maps. No previous evidence exists to expect that PMA25A and PMA25B represent two separate great tit chromosomes. A lack of power due to a lower number of markers on these linkage groups in combination with higher recombination rates in microchromosomes may, therefore, be the cause for not detecting linkage between these groups of markers. This observation is consistent with an earlier mapping study in zebra finch, where the homologues of several chicken microchromosomes were found to be split among more than one zebra finch linkage group (Stapley et al., 2008), despite chromosome painting results, indicating that these linkage groups are physically on the same chromosome (Itoh and Arnold, 2005; Nanda et al., 2011). However, when markers from PMA25A and PMA25B were mapped together in a single linkage group, there was relatively weak evidence of linkage between markers on these linkage groups in the NL data set and no evidence in the UK data set. At this stage it is unclear whether these linkage groups represent one or two chromosomes. Comparative mapping of the macrochromosomes gave identical results to a recent mapping study of the zebra finch, where markers aligning to GGA1 were found on three linkage groups (TGU1, TGU1A and TGU1B) and GGA4 was found to be homologous to two macrochromosomes (Stapley et al., 2008). When more markers were added, TGU1 and TGU1B were merged (Backström et al., 2010), which was confirmed in our study. This also confirms the results from chromosome painting studies for great tit, zebra finch and several other passerines (Itoh and Arnold, 2005; Nanda et al., 2011).
The large blocks of synteny apparent by comparing the green anole lizard and the passerine great tit genome are, in general, consistent with an earlier comparison of green anole lizard and chicken (Alfoldi et al., 2011). However, by comparing SNP locations in the great tit with their locations in the zebra finch, chicken and green anole lizard genomes, it was possible to infer the ancestral state of some of the largest differences between galliform and passerine karyotypes. Our results strongly suggest that chromosomes 1 and 1A were separate in a common avian ancestor, but became fused in the lineage that led to chickens. Similarly, PMA4 aligned to only one single Anolis carolinensis chromosome (A. carolinensis 5), whereas none of the 115 SNPs that were present on PMA4A aligned to a position on the A. carolinensis genome. Although this is not direct proof that PMA4 and PMA4A originate from two distinct ancestral chromosomes, it is consistent with the idea that these chromosomes also underwent a fusion in the chicken ancestor (Griffin et al., 2008). The ancestral states of other chromosomes cannot be inferred from our data.
Intrachromosomal rearrangements
Despite the highly conserved synteny between chickens and passerines, we found numerous intrachromosomal rearrangements. The two replicated maps were very consistent in support for the rearrangements, allowing us to conclude that they are true species-level rearrangements. Inversions and other changes in gene order are considered to be an important factor in speciation processes as they can promote reproductive isolation (Noor et al., 2001). A comparison of multiple galliform and passerine species is needed to shed further light on this aspect of avian chromosome evolution; this is an exciting area for further research, as we anticipate increasingly dense avian linkage maps to become available (see for example, Stapley et al., 2010).
In conclusion, we have constructed high-density linkage maps of two independent great tit populations. The maps will be valuable resources to aid with quantitative trait loci mapping, genome-wide association studies and chromosome partitioning of quantitative genetic variation (Santure et al., 2013; Robinson et al., 2013). More fundamentally, we show that replicated maps constructed using the same SNPs reveal new insight into bird karyotype evolution; in particular a hitherto undetected degree of fine scale heterochiasmy that opens up an exciting new opportunity to study the evolution and fitness consequences of heterochiasmy.
Data archiving
Genotype and pedigree data along with SNP information were deposited in the Dryad repository: doi:10.5061/dryad.j7260.
References
Åkesson M, Hansson B, Hasselquist D, Bensch S . (2007). Linkage mapping of AFLP markers in a wild population of great reed warblers: importance of heterozygosity and number of genotyped individuals. Mol Ecol 16: 2189–2202.
Alfoldi J, Di Palma F, Grabherr M, Williams C, Kong LS, Mauceli E et al. (2011). The genome of the green anole lizard and a comparative analysis with birds and mammals. Nature 477: 587–591.
Andrews CB, Mackenzie SA, Gregory TR . (2009). Genome size and wing parameters in passerine birds. Proc R Soc B Biol Sci 276: 55–61.
Aslam ML, Bastiaansen JWM, Crooijmans R, Vereijken A, Megens HJ, Groenen MAM . (2010). A SNP based linkage map of the turkey genome reveals multiple intrachromosomal rearrangements between the Turkey and Chicken genomes. BMC Genomics 11: 647.
Bachmann K, Harington BA, Craig JP . (1972). Genome size in birds. Chromosoma 37: 405–416.
Backström N, Brandström M, Gustafsson L, Qvarnström A, Cheng H, Ellegren H . (2006). Genetic mapping in a natural population of collared flycatchers (Ficedula albicollis): conserved synteny but gene order rearrangements on the avian Z chromosome. Genetics 174: 377–386.
Backström N, Forstmeier W, Schielzeth H, Meelenius H, Nam K, Bolund E et al. (2010). The recombination landscape of the zebra finch taeniopygia guttata genome. Genome Res 20: 485–495.
Backström N, Karaiskou N, Leder EH, Gustafsson L, Primmer CR, Qvarnstrom A et al. (2008). A gene-based genetic linkage map of the collared flycatcher (Ficedula albicollis) reveals extensive synteny and gene-order conservation during 100 million years of avian evolution. Genetics 179: 1479–1495.
Ball AD, Stapley J, Dawson DA, Birkhead TR, Burke T, Slate J . (2010). A comparison of SNPs and microsatellites as linkage mapping markers: lessons from the zebra finch (Taeniopygia guttata). BMC Genomics 11: 218.
Barker FK, Cibois A, Schikler P, Feinstein J, Cracraft J . (2004). Phylogeny and diversification of the largest avian radiation. Proc Natl Acad Sci USA 101: 11040–11045.
Bennett PM, Owens IPF . (2002) Evolutionary Ecology of Birds: Life Histories, Mating Systems and Extinction. Oxford University Press: Oxford.
Broman KW, Murray JC, Sheffield VC, White RL, Weber JL . (1998). Comprehensive human genetic maps: individual and sex-specific variation in recombination. Am J Hum Genet 63: 861–869.
Charmantier A, McCleery RH, Cole LR, Perrins C, Kruuk LEB, Sheldon BC . (2008). Adaptive phenotypic plasticity in response to climate change in a wild bird population. Science 320: 800–803.
Dingemanse NJ, Both C, Drent PJ, van Oers K, van Noordwijk AJ . (2002). Repeatability and heritability of exploratory behaviour in great tits from the wild. Animal Behav 64: 929–938.
Drent PJ, van Oers K, van Noordwijk AJ . (2003). Realized heritability of personalities in the great tit (Parus major). Proc R Soc Lond B Biol Sci 270: 45–51.
Dunn LC, Bennett D . (1967). Sex differences in recombination of lined genes in animals. Genet Res 9: 211–220.
Ellegren H . (2007). Characteristics, causes and evolutionary consequences of male-biased mutation. Proc R Soc B Biol Sci 274: 1–10.
Ellegren H . (2010). Evolutionary stasis: the stable chromosomes of birds. Trends Ecol Evol 25: 283–291.
Evans SR, Sheldon BC . (2012). Quantitative genetics of a carotenoid-based color: heritability and persistent natal environmental effects in the great tit. Am Nat 179: 79–94.
Garant D, Kruuk LEB, Wilkin TA, McCleery RH, Sheldon BC . (2005). Evolution driven by differential dispersal within a wild bird population. Nature 433: 60–65.
Green P, Falls K, Crooks S . (1990) Documentation for CRI-MAP In. Washington University: St. Louis.
Griffin DK, Robertson LB, Tempest HG, Vignal A, Fillon V, Crooijmans RP et al. (2008). Whole genome comparative studies between chicken and turkey and their implications for avian genome evolution. BMC Genomics 9: 168.
Groenen MAM, Cheng HH, Bumstead N, Benkel BF, Briles WE, Burke T et al. (2000). A consensus linkage map of the chicken genome. Genome Res 10: 137–147.
Groenen MAM, Wahlberg P, Foglio M, Cheng HH, Megens HJ, Crooijmans R et al. (2009). A high-density SNP-based linkage map of the chicken genome reveals sequence features correlated with recombination rate. Genome Res 19: 510–519.
Haldane JBS . (1922). Sex ratio and unisexual sterility in hybrid animals. J Genet 12: 101–109.
Hansson B, Akesson M, Slate J, Pemberton JM . (2005). Linkage mapping reveals sex-dimorphic map distances in a passerine bird. Proc R Soc Lond B Biol Sci 272: 2289–2298.
Hansson B, Ljungqvist M, Dawson DA, Mueller JC, Olano-Marin J, Ellegren H et al. (2010). Avian genome evolution: insights from a linkage map of the blue tit (Cyanistes caeruleus). Heredity 104: 67–78.
Heeb P, Werner I, Mateman AC, Kolliker M, Brinkhof MWG, Lessells CM et al. (1999). Ectoparasite infestation and sex-biased local recruitment of hosts. Nature 400: 63–65.
Hoaglin DC, Iglewicz B, Tukey JW . (1986). Performance of some resistant rules for outlier labeling. J Am Stat Assoc 81: 991–999.
Huang YH, Zhao YH, Haley CS, Hu SQ, Hao JP, Wu CX . (2006). A genetic and cytogenetic map for the duck (Anas platyrhynchos). Genetics 173: 287–296.
Huxley JS . (1928). Sexual differences of linkage Gammarus chevreuxi. J Genet 20: 145–156.
Itoh Y, Arnold AP . (2005). Chromosomal polymorphism and comparative painting analysis in the zebra finch. Chromosome Res 13: 47–56.
Jaari S, Li MH, Merila J . (2009). A first-generation microsatellite-based genetic linkage map of the Siberian jay (Perisoreus infaustus): insights into avian genome evolution. BMC Genomics 10: 1.
Kayang BB, Fillon V, Inoue-Murayama M, Miwa M, Leroux S, Feve K et al. (2006). Integrated maps in quail (Coturnix japonica) confirm the high degree of synteny conservation with chicken (Gallus gallus) despite 35 million years of divergence. BMC Genomics 7: 101.
Lovich JE, Gibbons JW . (1992). A review of techniques for quantifying sexual size dimorphism. Growth Dev Aging 56: 269–281.
Naguib M, Kazek A, Schaper SV, van Oers K, Visser ME . (2010). Singing activity reveals personality traits in great tits. Ethology 116: 763–769.
Nanda I, Benisch P, Fetting D, Haaf T, Schmid M . (2011). Synteny conservation of chicken macrochromosomes 1-10 in different avian lineages revealed by cross-species chromosome painting. Cytogenet Genome Res 132: 165–181.
Noor MAF, Grams KL, Bertucci LA, Reiland J . (2001). Chromosomal inversions and the reproductive isolation of species. Proc Natl Acad Sci USA 98: 12084–12088.
Norris K, Anwar M, Read AF . (1994). Reproductive effort influences the prevalence of haematozoan parasites in great tits. J Animal Ecol 63: 601–610.
Otto SP, Lenormand T . (2002). Resolving the paradox of sex and recombination. Nat Rev Genet 3: 252–261.
Poissant J, Hogg JT, Davis CS, Miller JM, Maddox JF, Coltman DW . (2010). Genetic linkage map of a wild genome: genomic structure, recombination and sexual dimorphism in bighorn sheep. BMC Genomics 11: 524.
Popa A, Samollow P, Gautier C, Mouchiroud D . (2012). The sex-specific impact of meiotic recombination on nucleotide composition. Genome Biol Evol 4: 412–422.
Postma E, van Noordwijk AJ . (2005). Gene flow maintains a large genetic difference in clutch size at a small spatial scale. Nature 433: 65–68.
Quinn JL, Patrick SC, Bouwhuis S, Wilkin TA, Sheldon BC . (2009). Heterogeneous selection on a heritable temperament trait in a variable environment. J Animal Ecol 78: 1203–1215.
Reed KM, Chaves LD, Mendoza KM . (2007). An integrated and comparative genetic map of the turkey genome. Cytogenet Genome Res 119: 113–126.
Robertson DS . (1984). Different frequency in the recovery of crossover products from male and female gametes of plants hypoploid for B-a translocations in maize. Genetics 107: 117–130.
Robertson MR, Santure AW, DeCauwer I, Sheldon BC, Slate J . (2013). Partitioning of genetic variation across the genome using multimarker methods in a wild bird population. Mol Ecol 22: 3963–3980.
Santure AW, DeCauwer I, Robinson MR, Poissant J, Sheldon BC, Slate J . (2013). Genomic dissection of variation in clutch size and egg mass in a wild great tit (Parus major) population. Mol Ecol 22: 3949–3962.
Santure AW, Gratten J, Mossman JA, Sheldon BC, Slate J . (2011). Characterisation of the transcriptome of a wild great tit Parus major population by next generation sequencing. BMC Genomics 12: 283.
Slabbekoorn H, Peet M . (2003). Ecology: birds sing at a higher pitch in urban noise - great tits hit the high notes to ensure that their mating calls are heard above the city's din. Nature 424: 267–267.
Slate J . (2005). Quantitative trait locus mapping in natural populations: progress, caveats and future directions. Mol Ecol 14: 363–379.
Stapley J, Birkhead TR, Burke T, Slate J . (2008). A linkage map of the zebra finch Taeniopygia guttata provides new insights into avian genome evolution. Genetics 179: 651–667.
Stapley J, Birkhead TR, Burke T, Slate J . (2010). Pronounced inter- and intrachromosomal variation in linkage disequilibrium across the zebra finch genome. Genome Res 20: 496–502.
Team R.D.C. (2011) R Foundation for Statistical Computing URL http://www.R-project.org In. R foundation for Statistical Computing Vienna: Austria.
Trivers R . (1988). Sex differences in rates of recombination and sexual selection. In: The Evolution of Sex (eds Michod RL, Levin BR, Sinauer Press Sunderland: Massachusetts. pp 270–286.
van Bers NEM, Santure A, van Oers K, De Cauwer I, Dibbits BW, Mateman AC et al. (2012). The design and cross-population application of a genome-wide SNP chip for the great tit Parus major. Mol Ecol Resour 12: 753–770.
van Bers NEM, van Oers K, Kerstens HHD, Dibbits BW, Crooijmans RPMA, Visser ME . (2010). Genome-wide SNP detection in the great tit Parus major using high throughput sequencing. Mol Ecol 19: 89–99.
van Oers K, Drent PJ, De Jong G, van Noordwijk AJ . (2004). Additive and nonadditive genetic variation in avian personality traits. Heredity 93: 496–503.
van Oers K, Naguib M . (2013). Avian Personality. In: Animal Personalities: Behavior, Physiology, and Evolution (eds Carere C, Maestripieri D, The University of Chicago Press: Chicago and London. pp 66–95.
Visser ME, van Noordwijk AJ, Tinbergen JM, Lessells CM . (1998). Warmer springs lead to mistimed reproduction in great tits (Parus major). Proc R Soc Lond B Biol Sci 265: 1867–1870.
Acknowledgements
We thank the many people who have collected data and blood samples from the great tit populations used in this study over the past decades. Matt Robinson extracted DNA of a substantial number of the UK samples and Christa Mateman and Bert Dibbits extracted DNA from the NL samples and contributed to SNP genotyping in the NL population. Our thanks to Christa Mateman, Martin Elferink and Winnie Raey for their help in building the NL map. This work was funded by the European Research Council (ERC Starting Grant GA202487–Avian EGG to JS), by the Netherlands Genomics Initiative (NGI-HORIZON) and by an NWO-VICI grant to MEV.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
van Oers, K., Santure, A., De Cauwer, I. et al. Replicated high-density genetic maps of two great tit populations reveal fine-scale genomic departures from sex-equal recombination rates. Heredity 112, 307–316 (2014). https://doi.org/10.1038/hdy.2013.107
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2013.107
Keywords
This article is cited by
-
From molecules to populations: appreciating and estimating recombination rate variation
Nature Reviews Genetics (2020)
-
Exploring the unmapped DNA and RNA reads in a songbird genome
BMC Genomics (2019)
-
Avian ecological epigenetics: pitfalls and promises
Journal of Ornithology (2019)
-
Karyotypes and recombination patterns of the Common Swift (Apus apus Linnaeus, 1758) and Eurasian Hobby (Falco subbuteo Linnaeus, 1758)
Avian Research (2018)
-
Gene and transposable element methylation in great tit (Parus major) brain and blood
BMC Genomics (2016)