Abstract
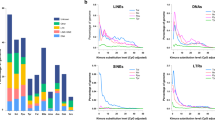
Members of the cys-loop ligand-gated ion channel (cysLGIC) superfamily mediate chemical neurotransmission and are studied extensively as potential targets of drugs used to treat neurological disorders, such as Alzheimer's disease. Insect cys-loop LGICs also have central roles in the nervous system and are targets of highly successful insecticides. Here, we describe the cysLGIC superfamily of the parasitoid wasp, Nasonia vitripennis, which is emerging as a highly useful model organism and is deployed as a biological control of insect pests. The wasp superfamily consists of 26 genes, which is the largest insect cysLGIC superfamily characterized, whereas Drosophila melanogaster, Apis mellifera and Tribolium castaneum have 23, 21 and 24, respectively. As with Apis, Drosophila and Tribolium, Nasonia possesses ion channels predicted to be gated by acetylcholine, γ-amino butyric acid, glutamate and histamine, as well as orthologues of the Drosophila pH-sensitive chloride channel (pHCl), CG8916 and CG12344. Similar to other insects, wasp cysLGIC diversity is broadened by alternative splicing and RNA A-to-I editing, which may also serve to generate species-specific receptor isoforms. These findings on N. vitripennis enhance our understanding of cysLGIC functional genomics and provide a useful basis for the study of their function in the wasp model, as well as for the development of improved insecticides that spare a major beneficial insect species.
Similar content being viewed by others
Introduction
Members of the cys-loop ligand-gated ion channel (cysLGIC) superfamily mediate both fast excitatory and inhibitory synaptic transmission in the nervous system of vertebrates and invertebrates. In insects, the superfamily includes cation-permeable nicotinic acetylcholine receptors (nAChRs) (Raymond-Delpech et al., 2005; Sattelle et al., 2005), γ-amino butyric acid (GABA)-gated anion channels (Ffrench-Constant et al., 1993; Buckingham et al., 2005), glutamate-gated chloride channels (GluCls) (Cully et al., 1996) and histamine-gated chloride channels (HisCls) (Gisselmann et al., 2002; Zheng et al., 2002). Studies of Drosophila melanogaster and Apis mellifera have shown that cysLGICs mediate important aspects of behaviour, such as escape response (Fayyazuddin et al., 2006), sleep (Agosto et al., 2008), learning and memory (El Hassani et al., 2005; Gauthier et al., 2006; Liu et al., 2007). CysLGICs are also of considerable interest as they are targets of widely used insecticides (Raymond-Delpech et al., 2005). For example, nAChRs are targets of neonicotinoids, which have been the fastest-growing class of insecticides in modern crop protection (Jeschke and Nauen, 2008). In addition, GABA receptors, GluCls and HisCls are targets of fipronil and avermectins (Iovchev et al., 2002; Bloomquist, 2003).
The parasitoid wasp, Nasonia vitripennis, is emerging as a useful model organism because of its ease of handling in the laboratory, as well as the possibility to apply RNA interference (Lynch and Desplan, 2006) and haplo-diploid genetics to study biological processes (Pultz and Leaf, 2003). In addition, Nasonia is used in the biological control of insects, including vectors of human, animal and plant disease, and agricultural pests (Werren et al., 2010). ‘In the United States alone, biological control programmes using parasitoid wasps save approximately US$20 billion annually in crop loss to newly invasive species, a figure excluding the vast savings from biological control of native pests’ (Werren et al., 2010). The recent sequencing of the N. vitripennis genome (Werren et al., 2010) will therefore likely provide a valuable basis for studying important biological processes and lead to methods for further enhancing their utility in controlling insect pests. We report here, as part of the Nasonia genome-sequencing effort, the N. vitripennis cysLGIC gene superfamily, which represents a critical step in identifying key components of the wasp nervous system.
Materials and methods
Identification of cysLGIC subunits in the N. vitripennis genome
To identify putative cysLGIC subunits, we screened the N. vitripennis genome (assembly version 1.0) (http://www.hgsc.bcm.tmc.edu/projects/nasonia/) with cDNA sequences of every member of the A. mellifera cysLGIC superfamily using the tBLASTn algorithm (Altschul et al., 1990). Candidate wasp cysLGIC subunits were identified based on their considerable sequence homology with previously characterized subunits (sequence with lowest similarity had E-value 8e−8), particularly in the N-terminal ligand-binding domain and the four transmembrane regions. The highly variable N-terminal signal peptides, which are a feature of cysLGIC subunits, were identified from the GLEAN consensus set of predicted genes (Werren et al., 2010), the EST data available at NCBI (http://www.ncbi.nlm.nih.gov/) and by RACE PCR. Reverse transcriptase-PCRs were carried out (see Supplementary Table 1 for sequences of primers used) to verify and correct the open reading frame sequences of each subunit.
Reverse transcription and PCR
Total RNA was extracted from 12 N. vitripennis adult wasps homogenized in Trizol (Invitrogen, Paisley, UK) using the RNeasy Mini Kit (Qiagen, Crawley, UK). First-strand cDNA was synthesized from 1 μg total RNA using Superscript III First-Strand Synthesis Super Mix (Invitrogen). N-terminal signal peptides not identified in predicted protein sequences or EST data were obtained by 5′ RACE PCR using the 5′/3′ RACE Kit, 2nd Generation (Roche, Burgess Hill, UK). We were unable to determine the N-terminal sequences for some subunits (Figures 1 and 2). Nested reverse transcriptase-PCR reactions were carried out to amplify transcripts of wasp cysLGIC subunits and to detect transcript variants arising from alternative splicing. Primer pairs (see Supplementary Table 1) recognizing different exons were used to allow the identification of cDNA-specific products. The PCR reactions were carried out in a total volume of 50 μl containing Taq polymerase and 1 × PCR buffer (Sigma, Gillingham, UK), 0.2 mM dNTP mix (Roche), 0.4 μM of each primer and 2 μl first-strand cDNA template. The PCR reaction conditions were 35 cycles of 95 °C for 30 s, 50–55 °C for 30 s and 72 °C for 90 s. The first PCR was used at a final dilution of 1 in 500 as template for the second nested PCR reaction. DNA sequence chromatograms for each cysLGIC subunit were analysed using Chromas 2 (Technelysium Pty Ltd, Tewantin, Australia) to detect single nucleotide polymorphisms (SNPs) or RNA-editing sites as shown by mixed signal peaks. No SNPs were observed, and the putative RNA-editing sites detected in Nvitα6 were verified by amplifying and sequencing genomic DNA present in the extracted total RNA, which was first treated with DNase-free RNase (Roche), using primers recognizing intron DNA (see Supplementary Table 3 for primers used). Sequence chromatograms showing a defined region of mixed peaks indicated differential splicing. The corresponding cysLGIC PCR products were cloned into the pGEM-T Easy vector (Promega, Southampton, UK) and between 10 and 20 transformants were sequenced to identify individual subunit isoforms. All PCR products were analysed by electrophoresis in a TAE gel and then purified using the QIAquick Gel Extraction Kit (Qiagen), whereas subunits cloned in pGEM-T Easy were purified using the QIAprep Spin Miniprep Kit (Qiagen). Purified DNA was sequenced by the dye termination method at Cogenics (Essex, UK, http://www.cogenics.com/index.cfm).
Protein sequence alignment of N. vitripennis nAChR subunits. Dα1 of D. melanogaster is included for comparison. N-terminal signal leader peptides are shown in lower case and the loops implicated in ligand binding (LpA-F) as well as the four transmembrane regions (TM1-4) are indicated. Residues preceding TM2, which are important for ion charge selectivity, are indicated by exclamation marks. The two cysteines forming the cys-loop are marked by asterisks and the vicinal cysteines characteristic of α subunits are highlighted by black shading. Putative N-glycosylation sites are boxed and potential phosphorylation sites are highlighted as follows: tyrosine kinase phosphorylation (encircled), protein kinase C phosphorylation (light grey shading), casein kinase II phosphorylation (dark grey shading) and cAMP- and cGMP-dependent protein kinase phosphorylation (dark grey shading with white text). nAChR, cation-permeable nicotinic acetylcholine receptor.
Protein sequence alignment of N. vitripennis cysLGIC subunits other than nAChRs. Rdl of D. melanogaster is included for comparison. N-terminal signal leader peptides are shown in lower case and the loops implicated in ligand binding (LpA–F) as well as the four transmembrane regions (TM1–4) are indicated. Residues preceding TM2, which are important for ion charge selectivity, are indicated by exclamation marks. The two cysteines forming the cys-loop are marked by asterisks, whereas the putative second cys-loop in LpC is highlighted by black shading. Putative N-glycosylation sites are boxed and potential phosphorylation sites are highlighted as follows: tyrosine kinase phosphorylation (encircled), protein kinase C phosphorylation (light grey shading), casein kinase II phosphorylation (dark grey shading) and cAMP- and cGMP-dependent protein kinase phosphorylation (dark grey shading with white text). HisCl1 and HisCl2 have been abbreviated to His1 and His2, respectively. cysLGIC, cys-loop ligand-gated ion channel; HisCl, histamine-gated chloride channel; nAChR, cation-permeable nicotinic acetylcholine receptor.
Sequence analysis
The multiple protein sequence alignments were constructed with ClustalX (Thompson et al., 1997) using the slow-accurate mode with a gap-opening penalty of 10 and a gap-extension penalty of 0.1 and applying the Gonnet 250 protein weight matrix. The protein alignments were viewed using GeneDoc (http://www.nrbsc.org/gfx/genedoc/index.html). Identity values between subunit sequences were calculated using the GeneDoc program. The neighbour-joining method and bootstrap resampling, available with the ClustalX program, were used to construct a phylogenetic tree, which was then displayed using the TreeView application (Page, 1996). Signal-peptide cleavage sites were predicted using the SignalP 3.0 server (Bendtsen et al., 2004) and membrane-spanning regions were predicted using the TMpred program (http://www.ch.embnet.org/software/TMPRED_form.html). The PROSITE database (Hulo et al., 2006) was used to identify potential N-glycosylation and phosphorylation sites.
Results
The N. vitripennis cysLGIC superfamily consists of 26 subunit members
Using tBLASTn (Altschul et al., 1990), 26 candidate cysLGIC subunit genes were identified in the N. vitripennis genome and manually annotated. This is the fourth complete insect cysLGIC superfamily to be described after those of D. melanogaster (Dent, 2006), A. mellifera (Jones and Sattelle, 2006) and Tribolium castaneum (Jones and Sattelle, 2007), and the largest known to date as the fruit fly, honey bee and red beetle possess 23, 21 and 24 subunits, respectively. Reverse transcriptase–PCR (see Supplementary Table 1 for primers used) was used to show that all of the Nasonia cysLGIC subunit genes are transcribed, with one exception, the nAChR subunit Nvitβ4. An alignment of their protein sequences shows that the wasp subunits possess features common to members of the cysLGIC superfamily (Sine and Engel, 2006) (Figures 1 and 2). These include: (i) an extracellular N-terminal domain containing distinct regions (loops A–F) that form the ligand-binding site (Corringer et al., 2000); (ii) the dicysteine loop (cys-loop) consisting of two disulphide bond-forming cysteines separated by 13 amino-acid residues; (iii) four transmembrane regions (TM1–4), the second of which (TM2) contributes most of the channel-lining residues; (iv) a highly variable intracellular loop between TM3 and TM4. As with other cysLGIC subunits, the Nasonia sequences also possess potential N-glycosylation sites in the extracellular N-terminal domain, which can affect receptor maturation, channel desensitization and conductance (Gehle et al., 1997; Nishizaki, 2003), and putative phosphorylation sites in the TM3–TM4 intracellular loop, which regulates several aspects of receptor function, such as desensitization and aggregation (Hopfield et al., 1988; Borges and Ferns, 2001).
A comparison of sequence identities between N. vitripennis, A. mellifera and D. melanogaster cys-loop LGIC subunits (Tables 1 and 2), as well as the use of a phylogenetic tree (Figure 3), indicates orthologous relationships between the wasp, honey bee and fruit fly subunits. To facilitate comparisons between the three species, Nasonia subunits were named after their Drosophila counterparts as previously done with Apis and Tribolium subunits (Jones and Sattelle, 2006, 2007). For example, the N. vitripennis orthologues of Drosophila Dα1, RDL and CG8916 were designated Nvitα1, Nvit_RDL and Nvit_8916, respectively.
Unrooted tree showing relationships of N. vitripennis, A. mellifera and D. melanogaster cys-loop LGIC subunit protein sequences. Numbers at each node signify bootstrap values with 100 replicates and the scale bar represents substitutions per site. The subunits shown in the tree are as follows: A. mellifera Amelα1 (DQ026031), Amelα2 (AY540846), Amelα3 (DQ026032), Amelα4 (DQ026033), Amelα5 (AY569781), Amelα6 (DQ026035), Amelα7 (AY500239), Amelα8 (AF514804), Amelα9 (DQ026037), Amelβ1 (DQ026038), Amelβ2 (DQ026039), Amel_GluCl (DQ667185), Amel_GRD (DQ667183), Amel_HisCl1 (DQ667187), Amel_HisCl2 (DQ667188), Amel_LCCH3 (DQ667184), Amel_pHCl (DQ667189), Amel_RDL (DQ667182), Amel_6927 (DQ667195), Amel_8916 (DQ667193), Amel_12344 (DQ667194); D. melanogaster Dα1 (CAA30172), Dα2 (CAA36517), Dα3 (CAA75688), Dα4 (CAB77445), Dα5 (AAM13390), Dα6 (AAM13392), Dα7 (AAK67257), Dβ1 (CAA27641), Dβ2 (CAA39211), Dβ3 (CAC48166), GluCl (AAG40735), GRD (Q24352), HisCl1 (AAL74413), HisCl2 (AAL74414), LCCH3 (AAB27090), the putative cysLGIC subunit Ntr (AF045471), pHCl (NP_001034025), RDL (AAA28556), CG6927 (AAF45992), CG7589 (AAF49337), CG8916 (BT022901), CG11340 (AAF57144), CG12344 (AAF58743); N. vitripennis Nvitα1 (FJ821429), Nvitα2 (FJ821430), Nvitα3 (FJ821431), Nvitα4 (FJ821432), Nvitα5 (FJ821434), Nvitα6 (FJ821435), Nvitα7 (FJ821438), Nvitα8 (FJ821439), Nvitα9 (FJ821440), Nvitα10 (FJ821441), Nvitα11 (FJ821442), Nvitα12 (FJ821443), Nvitβ1 (FJ821444), Nvitβ2 (FJ821445), Nvitβ3 (FJ821446), Nvitβ4 (FJ821447), Nvit_HisCl1 (FJ851089), Nvit_HisCl2 (FJ851090), Nvit_pHCl (FJ851091), Nvit_GRD (FJ851092), Nvit_8916 (FJ851093), Nvit_12344 (FJ851094), Nvit_CLGC1 (FJ851095), Nvit_LCCH3 (FJ851096), Nvit_RDL (FJ851097), Nvit_GluCl (FJ851099). cysLGIC, cys-loop ligand-gated ion channel; GluCl, glutamate-gated chloride channel; HisCl, histamine-gated chloride channel.
Nasonia nicotinic acetylcholine receptor subunits
We identified 16 candidate nAChR subunit-encoding genes in the N. vitripennis genome. This is larger than other insect nAChR gene families described, which are of D. melanogaster (10 subunits), Anopheles gambiae (10), A. mellifera (11), Bombyx mori (12) and T. castaneum (12) (Sattelle et al., 2005; Jones et al., 2005a, 2006; Jones and Sattelle, 2007; Shao et al., 2007). Twelve of the Nasonia nAChR subunits possess the two adjacent cysteine residues in loop C (Figure 1), which are important for acetylcholine (ACh) binding (Corringer et al., 2000), defining them as α subunits. The remaining four subunits were designated β as they lack the vicinal cysteines. For Nvitβ4, only a partial sequence was found, possibly because of incomplete genome information, with loops D to F missing (Figure 1). Nvitβ4 is likely to be a pseudogene as a frameshift in exon 1 (Supplementary Figure 1) results in a premature stop codon and its peptide sequence is likely to be spread over two reading frames. As PCR with primers specific to different exons failed to amplify Nvitβ4 cDNA products, we conclude that the subunit is unlikely to be transcribed. To enable Nvitβ4 to be included in the protein sequence analysis, we assumed that the subunit was encoded by a single reading frame (Figure 1, Supplementary Figure 1).
Nasonia possesses the typical core groups of nAChR subunits that are highly conserved between different insect species (Jones et al., 2007). Thus, subunit equivalents of Dα1–7, Dβ1 and Dβ2, are evident in the wasp genome (Figure 3). As is the case for other non-drosophilid insects (Jones et al., 2005b), the Nasonia orthologue of Dβ2 is of the α type (Nvitα8). Nvitα5, along with its orthologues in Apis, Bombyx and Tribolium, depart strongly from the Dipteran orthologues, Dα5 (Figure 3), Agamα5 and Mdα5, of Musca domestica (Gao et al., 2007b). As is the case for Dα1, Dα2, Dα3, Dα4, Dβ2 and their orthologues in other insect species, the corresponding Nasonia subunits (Nvitα1–4 and Nvitα8) have an insertion in loop F (Figure 1), which may contribute to interactions with the neonicotinoid, imidacloprid (Shimomura et al., 2004). The Dα1, Dα2 and Dβ2 genes, as well as their Anopheles orthologues, Agamα1, Agamα2 and Agamα8, are similarly arranged and tightly clustered within 200 and 220 kb, respectively (Jones et al., 2005a). Immunohistochemical and coimmunoprecipitation studies show that Dα1, Dα2 and Dβ2 are integral components of certain nAChR subtypes, leading to the suggestion that gene clustering may favour coordinated expression and co-assembly of these nAChR subunits (Chamaon et al., 2002). In Nasonia, as with Apis, Nvitα1 and Nvitα2 are clustered on scaffold 1 but are separated from the wasp Dβ2 orthologue, Nvitα8, which is located on scaffold 9. The separation of these genes may thus result in diversification of receptor expression and co-assembly. Another subunit, Nvitα5, is however clustered with Nvitα1 and Nvitα2, being located within 220 kb of both subunits. The clustering of the honey bee subunits Amelα7 and Amelβ1 (Jones et al., 2006), which is also seen in the genomes of Anopheles and Tribolium (Jones et al., 2005a; Jones and Sattelle, 2007), is not conserved in Nasonia, as Nvitα7 and Nvitβ1 are located on scaffolds 31 and 8, respectively.
Insect nAChR gene families possess at least one divergent subunit that shares relatively low sequence identity with other nAChR subunits (Jones et al., 2007). The five insect nAChR gene families previously described (those of A. gambiae, A. mellifera, B. mori, D. melanogaster and T. castaneum) contain up to three divergent subunits, each with a different complement of α and β subunits (Lansdell and Millar, 2002; Sattelle et al., 2005; Jones et al., 2005a, 2006; Jones and Sattelle, 2007; Shao et al., 2007). We found a considerably larger number of divergent subunits in the Nasonia genome consisting of four α and three β subunits (Figure 3). As with one of the Tribolium divergent subunits, Tcasα9, Nvitα11 possesses an atypical FxCC amino-acid motif (Figure 1), instead of the highly conserved YxCC found in loop C, which may result in unusual ligand-binding properties (Galzi et al., 1991). In addition, the Nasonia divergent subunits lack the GEK motif characteristic of nAChR subunits, which precedes TM2 (Figure 1), and has an important role in ion permeation and selectivity (Jensen et al., 2005). This raises the possibility that subunits of this type may form nAChRs with distinct ion channel characteristics. Nvitα11 and Nvitα12 are present within 300 kb of each other on scaffold 4 of the Nasonia genome, whereas Nvitβ2 and Nvitβ3 are clustered within 15 kb on scaffold 164, indicating that both pairs of subunits arose from a recent duplication event from a common gene. Nvitα9 and Nvitβ4 are also clustered together within 10 kb on scaffold 69. In the case of these two subunits, a duplication event may have caused a frameshift, resulting in Nvitβ4 being a pseudogene (Supplementary Figure 1).
Two Nasonia nAChR subunits, Nvitα4 and Nvitα6, have alternatively spliced exons most likely arising from tandem exon duplication (Kondrashov and Koonin, 2001). Thus, as with Dα4 and orthologues of several other insect species (Lansdell and Millar, 2000; Jones et al., 2005a, 2006; Jones and Sattelle, 2007; Shao et al., 2007), Nvitα4 possesses two alternatives for exon 4 (denoted exon4 and exon4′) (Figure 4a), and similar to Amelα6 (Jin et al., 2007), Nvitα6 has three alternatives for exon 8 (Figure 4b). Analysis of sequence chromatograms shows that both alternatives for Nvitα4 exon4 are transcribed and reverse transcriptase–PCR (see Supplementary Table 2 for primers used) detected all three possible combinations of alternate exons for Nvitα6. Alternative splicing of Nvitα4 exon4 substitutes residues in loop E, which may affect ligand binding (Corringer et al., 2000; Amiri et al., 2008), as well as residues in the vicinity of the cys-loop, which may affect receptor assembly (Lansdell and Millar, 2000). For Nvitα6, alternative splicing of exon 8 changes residues in the TM2 domain, which may alter the ion channel properties of the receptor (Grauso et al., 2002). The alternative exons of both Apis and Nasonia α4 and α6 subunits are identical, with the only exception that an alanine residue in Nvitα6 exon 8c is replaced by an asparagine in the equivalent honey bee exon (Figure 4b).
Alternative splicing of exons in Nasonia cys-loop LGIC subunits. Equivalent alternate exons of N. vitripennis and A. mellifera cys-loop LGIC subunits are aligned. (a) Exon 4 splice variants in Nvitα4 and Amelα4. The cysteine residues forming the cys-loop are marked by asterisks. (b) Splice variants of exon 8 in Nvitα6 and Amelα6. Similar to Apis, but unlike Drosophila, no alternatives for exon 3 were found. (c) Splice variants of exons 3 and 6 in both Nvit_RDL and Amel_RDL. (d) Exon 3 splice variants in Nvit_GluCl and Amel_GluCl. Throughout the figure, Nasonia residues, which differ from those of the orthologous Apis exon, are highlighted in bold, and loops B–F, as well as TM2, are indicated. GluCl, glutamate-gated chloride channel.
In analysing the cDNA sequences of the Nasonia cysLGICs, we detected potential RNA A-to-I editing (Jepson and Reenan, 2007) in one subunit, Nvitα6 (Figure 5). Sequencing of the corresponding genomic DNA using primers listed in Supplementary Table 3 confirmed that the nucleotide changes occur at the RNA level. Nvitα6 is edited at five sites, three of which correspond to sites 4, 5 and 6 in Dα6 (Grauso et al., 2002), which are conserved in several diverse insect species such as A. mellifera (Jones et al., 2006), B. mori (Jin et al., 2007), D. melanogaster (Grauso et al., 2002), Heliothis virescens (Grauso et al., 2002), M. domestica (Gao et al., 2007a) and T. castaneum (Jones and Sattelle, 2007). The remaining two editing sites have so far been observed only in Nasonia and they may generate receptor isoforms with characteristics particular to the wasp as both sites alter the amino-acid residues in the vicinity of the ligand-binding domain and the cys-loop.
RNA A-to-I editing in Nvitα6. Arrows highlight the mixed adenosine/guanosine peak in the cDNA sequence indicating RNA editing and the resulting amino-acid change. The corresponding genomic DNA (gDNA) sequence, which lacks this mixed A/G signal, is also shown. Editing sites 4, 5 and 6 (Grauso et al., 2002), are indicated. The edited region is included with loop E indicated and a cysteine, which is part of the cys-loop, marked by an asterisk. An N-glycosylation site, which is removed by editing at sites 4 and 5, is boxed. Amino acids targeted by editing are highlighted in bold.
A comparison of Nasonia and Apis nAChR gene structures shows that four orthologue pairs (Nvitα1 and Amelα1, Nvitα4 and Amelα4, Nvitα8 and Amelα8, Nvitβ1 and Amelβ1) share identical sets of exon–intron boundaries (Figure 6). As noted for Amelα5, Amelα6 and Amelα7 (Jones et al., 2006), the orthologous Nasonia subunits, Nvitα5, Nvitα6 and Nvitα7, in addition to closely resembling the vertebrate's α7 subunits (up to 48% identity), possess exon–intron boundaries found in mammalian, bird and fish α7 and the closely related α8 subunits.
Exon composition of N. vitripennis and A. mellifera nAChR subunits. The N-terminal signal peptide is shown as a bar, the cys-loop is denoted as a star, and the four transmembrane regions are marked as white boxes. Conserved exon–intron boundaries are indicated by broken lines. Boundaries highly conserved in nAChR genes of invertebrates and vertebrates are highlighted by arrows, and boundaries particular to the vertebrate α7 subunits are also indicated. nAChR, cation-permeable nicotinic acetylcholine receptor.
Nasonia GABA-gated anion and cation channels
The Nasonia genome contains orthologues of the three known D. melanogaster GABA-gated ion channels (Figure 3 and Table 2), RDL, GRD and LCCH3 (Buckingham et al., 2005). As is the case for Drosophila, Apis and Tribolium (Jones and Sattelle, 2006, 2007), Nvit_RDL possesses a PAR sequence before TM2 (Figure 2), which is important for forming anion channels (Jensen et al., 2005), whereas Nvit_GRD and Nvit_LCCH3 lack this sequence, and thus may form cation-permeable channels as shown for Drosophila GRD and LCCH3 expressed in oocytes of Xenopus laevis (Gisselmann et al., 2004). However, whereas electrophysiology has clearly shown that GABA induces inhibitory chloride channels in insects, it remains to be established whether GABA-gated cation channels function in vivo (Buckingham et al., 2005).
As is the case for Drosophila and Apis RDL, exons 3 and 6 are alternatively spliced in Nvit_RDL (Figure 4c) with two alternatives for each exon. Exons 3a and 3b are completely conserved in both the Hymenoptera species. In contrast, exons 6a and 6b of the wasp and honey bee have different amino-acid residues in loops C and F, which may give rise to receptor variants that have distinct ligand-binding characteristics in the two insect species. The gene structures of Nasonia and Apis GABA subunits (Figure 7) differ in the Nvit_RDL and Nvit_LCCH3; each possess one more exon than their honey bee orthologues, whereas Nvit_GRD has no introns from TM3 to its C-terminus, and Amel_GRD has one in the TM3–TM4 intracellular loop. All three subunits possess up to four of the five exon–intron boundaries (labelled a, b, c, d and e) observed to be highly conserved in the invertebrate and vertebrate GABA receptor-like genes (Tsang et al., 2007) (Figure 7). In fact, boundaries b and d are present in all Apis and Nasonia non-nAChR cysLGIC subunits (Figure 7).
Exon composition of N. vitripennis and A. mellifera non-nAChR subunits. The N-terminal signal peptide is shown as a bar, the cys-loop is denoted as a star, and the four transmembrane regions are marked as white boxes. Conserved exon–intron boundaries are indicated by broken lines. Boundaries b–e, which are highly conserved in GABA receptor-like genes of invertebrates and vertebrates (Tsang et al., 2007) are highlighted by arrows. GABA, γ-amino butyric acid.
Nasonia glutamate and HisCls
As with D. melanogaster, A. mellifera and T. castaneum, N. vitripennis has one glutamate-gated chloride channel (Nvit_GluCl) and two histamine-gated chloride channel (Nvit_HisCl1 and Nvit_HisCl2) orthologues (Figure 3). Consistent with their putative function as anion channels, Nvit_GluCl, Nvit_HisCl1 and Nvit_HisCl2 all have the PAR motif preceding TM2 (Figure 2) (Jensen et al., 2005). Out of the ligand-gated anion channels, HisCl2 is the most highly conserved between Nasonia and Apis, sharing 95% identity (Table 2).
Nvit_GluCl has two alternatives for exon 3 (Figure 4d), as is the case for Amel_GluCl (Jones and Sattelle, 2006), with both exons being identical in the honey bee and wasp. The gene structures of Nvit_GluCl and Amel_GluCl are also the same, as are the gene structures of Nvit_HisCl1 and Amel_HisCl (Figure 7), whereas Nvit_HisCl2 possesses an additional exon to Amel_HisCl2 in the N-terminal extracellular domain.
Nasonia pHCl and uncharacterized cys-loop LGIC subunits
The N. vitripennis genome possesses an orthologue of the pH-sensitive chloride channel (Nvit_pHCl, Figures 2 and 3), which was first identified in Drosophila (Schnizler et al., 2005). The Drosophila pHCl has several splice variants, of which Variant 3 introduces an insertion and a protein kinase C phosphorylation site in the intracellular region between TM3 and TM4. We detected a similar variant in Nvit_pHCl transcripts (Figure 8a), where the peptide insertion differs by only one amino-acid residue from that of Drosophila pHCl and is completely identical to the equivalent insertion in Amel_pHCl (Jones and Sattelle, 2006). The novel insertion in loop C found in Amel_pHCl (Variant 4) (Jones and Sattelle, 2006) is also present in Nvit_pHCl, which differs from the bee insertion by four amino-acid residues (Figure 8b). The gene structures of Amel_pHCl and Nvit_pHCl differ, with the Nasonia gene having two extra exons near the N-terminus and an extra exon in the TM3–TM4 intracellular loop (Figure 7).
Differential splicing in Nvit_pHCl. (a) Alignment of Variant 3 of Drosophila pHCl with the equivalent Nasonia and Apis variants. The variants are caused by the differential use of splice sites, which inserts stretches of amino acids (underlined). A potential kinase C phosphorylation site is highlighted in grey shading. (b) Alignment of loop C (LpC) sequences of Nvit_pHCl Variant 4 and a similar variant in Apis (Amel_pHCl Variant 4) where use of differential splice sites introduces an insertion (underlined). Residues that differ between the equivalent splice variants of the two species are highlighted in bold. The N-glycosylation sites are boxed.
Five Drosophila cysLGIC subunits have yet to be functionally characterized. These are CG6927, CG7589, CG8916, CG11340 and CG12344. CG8916 and CG12344 appear to be closely related to GRD and HisCls, respectively (Figure 3), whereas CG6927, CG7589 and CG11340 form a distinct subfamily of cysLGIC subunits, which we have denoted Insect Group 1 of cysLGIC subunits (Jones and Sattelle, 2007). Nasonia possesses candidate orthologues of CG8916 and CG12344 and only one member (Nvit_CLGC1) of Insect Group 1 (Figure 3). A. mellifera also has one Insect Group 1 subunit called Amel_6927 (Jones and Sattelle, 2006), which shares high sequence identity with Nvit_CLGC1 of 77% (Table 2) as well as a similar gene structure (Figure 7). The finding of a single Group 1 subunit in both these Hymenoptera species is consistent with the suggestion that ‘gene duplication occurred after the emergence of the Hymenoptera to give rise to the three subunits present in both Tribolium (Coleoptera) and Drosophila (Diptera)’ (Jones and Sattelle, 2007).
Discussion
As part of the Nasonia genome-sequencing project (Werren et al., 2010), we have described the wasp cysLGIC gene superfamily, which encodes for receptors that have major roles in the nervous system and are also targets of highly successful insecticides. This is the fourth complete insect cysLGIC superfamily to be described after those of the Dipteran D. melanogaster (Littleton and Ganetzky, 2000), the Coleopteran T. castaneum (Jones and Sattelle, 2007) and the Hymenopteran A. mellifera (Jones and Sattelle, 2006). Thus, we report here the first comparison of two cysLGIC superfamilies from insects of the same order. Considering that the cysLGIC superfamilies of Drosophila, Apis and Tribolium, which represent diverse insect species of different orders, are compact with only minor changes in gene numbers (Jones and Sattelle, 2007), we find in the Nasonia genome a striking expansion in the number of nAChR subunits. Although A. gambiae (Jones et al., 2005a), A. mellifera (Jones et al., 2006), B. mori (Shao et al., 2007), D. melanogaster (Lansdell and Millar, 2002) and T. castaneum (Jones and Sattelle, 2007) possess one, two, three, one and two divergent subunits, respectively, N. vitripennis has seven (Figures 1 and 3). One of these is a pseudogene (Supplementary Figure 1), whereas the remaining six are potentially functional. It will be of interest to determine the functional roles of insect divergent nAChR subunits and why different species possess their own distinct complement. The non-nAChR subunits of Apis and Nasonia appear to be under a higher degree of evolutionary constraint with both species possessing an equal number of subunits, each with obvious orthologous relationships (Figure 3, Table 2).
Nasonia wasps are of agricultural importance as they control crop-damaging insect pests. The characterization of their cysLGICs provides an important basis for the future rational design of insecticides that control pests though sparing beneficial species. The use of heterologous expression systems such as X. laevis oocytes (Buckingham et al., 2006) has allowed the functional characterization of several Drosophila cysLGICs, such as RDL (Ffrench-Constant et al., 1993), GRD and LCCH3 (Gisselmann et al., 2004), GluCl (Cully et al., 1996), HisCl1 and HisCl2 (Gisselmann et al., 2002; Zheng et al., 2002) and pHCl (Schnizler et al., 2005). Similar studies of heterologously expressed ion channels from other insect species, including N. vitripennis, in combination with the use of three-dimensional models such as those based on the crystal structure of the molluscan acetylcholine-binding protein (Smit et al., 2003), will likely prove useful in the search for novel compounds that show selectivity for receptors of certain insect species as well as in determining the mechanisms of insecticide interaction with cysLGICs. For insect nAChRs, functional expression in heterologous systems has so far proven elusive, although α subunits of Drosophila and other insect species, such as Nilaparvata lugens, can form robust functional channels when coexpressed with a vertebrate β2 subunit and studies on such hybrid receptors have provided insights into the selectivity of neonicotinoids for insect nAChRs over those of vertebrates, regions of subunit proteins involved in imidacloprid interactions and the actions of different neonicotinoids (Matsuda et al., 2005; Liu et al., 2009).
Nasonia provides a powerful tool for the study of cysLGIC function. The possibility to use haplo-diploid genetics, according to which unfertilized eggs develop as haploid males, facilitates screening for recessive mutations affecting a biological process of interest, which may include cysLGIC physiology, pharmacology, trafficking or assembly. To add to the utility of Nasonia as a model organism, parental RNA interference, as a result of which RNA interference arising from double-stranded RNA introduced into pupae or adults also spreads to their progeny, is effective in N. vitripennis (Lynch and Desplan, 2006). Thus, RNA interference could be used to elucidate the roles of Nasonia cysLGIC subunits in various stages of development, different behaviours and responses to insecticides.
References
Agosto J, Choi JC, Parisky KM, Stilwell G, Rosbash M, Griffith LC (2008). Modulation of GABAA receptor desensitization uncouples sleep onset and maintenance in Drosophila. Nat Neurosci 11: 354–359.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Amiri S, Shimomura M, Vijayan R, Nishiwaki H, Akamatsu M, Matsuda K et al. (2008). A role for Leu118 of loop E in agonist binding to the alpha 7 nicotinic acetylcholine receptor. Mol Pharmacol 73: 1659–1667.
Bendtsen JD, Nielsen H, von Heijne G, Brunak S (2004). Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340: 783–795.
Bloomquist JR (2003). Chloride channels as tools for developing selective insecticides. Arch Insect Biochem Physiol 54: 145–156.
Borges LS, Ferns M (2001). Agrin-induced phosphorylation of the acetylcholine receptor regulates cytoskeletal anchoring and clustering. J Cell Biol 153: 1–12.
Buckingham SD, Biggin PC, Sattelle BM, Brown LA, Sattelle DB (2005). Insect GABA receptors: splicing, editing, and targeting by antiparasitics and insecticides. Mol Pharmacol 68: 942–951.
Buckingham SD, Pym L, Sattelle DB (2006). Oocytes as an expression system for studying receptor/channel targets of drugs and pesticides. Methods Mol Biol 322: 331–345.
Chamaon K, Smalla KH, Thomas U, Gundelfinger ED (2002). Nicotinic acetylcholine receptors of Drosophila: three subunits encoded by genomically linked genes can co-assemble into the same receptor complex. J Neurochem 80: 149–157.
Corringer PJ, Le Novere N, Changeux JP (2000). Nicotinic receptors at the amino acid level. Annu Rev Pharmacol Toxicol 40: 431–458.
Cully DF, Paress PS, Liu KK, Schaeffer JM, Arena JP (1996). Identification of a Drosophila melanogaster glutamate-gated chloride channel sensitive to the antiparasitic agent avermectin. J Biol Chem 271: 20187–20191.
Dent JA (2006). Evidence for a diverse Cys-loop ligand-gated ion channel superfamily in early bilateria. J Mol Evol 62: 523–535.
El Hassani AK, Dacher M, Gauthier M, Armengaud C (2005). Effects of sublethal doses of fipronil on the behavior of the honeybee (Apis mellifera). Pharmacol Biochem Behav 82: 30–39.
Fayyazuddin A, Zaheer MA, Hiesinger PR, Bellen HJ (2006). The nicotinic acetylcholine receptor Dalpha7 is required for an escape behavior in Drosophila. PLoS Biol 4: e63.
Ffrench-Constant RH, Rocheleau TA, Steichen JC, Chalmers AE (1993). A point mutation in a Drosophila GABA receptor confers insecticide resistance. Nature 363: 449–451.
Galzi JL, Bertrand D, Devillers-Thiery A, Revah F, Bertrand S, Changeux JP (1991). Functional significance of aromatic amino acids from three peptide loops of the alpha 7 neuronal nicotinic receptor site investigated by site-directed mutagenesis. FEBS Lett 294: 198–202.
Gao JR, Deacutis JM, Scott JG (2007a). The nicotinic acetylcholine receptor subunit Mdalpha6 from Musca domestica is diversified via post-transcriptional modification. Insect Mol Biol 16: 325–334.
Gao JR, Deacutis JM, Scott JG (2007b). The nicotinic acetylcholine receptor subunits Mdalpha5 and Mdbeta3 on autosome 1 of Musca domestica are not involved in spinosad resistance. Insect Mol Biol 16: 691–701.
Gauthier M, Dacher M, Thany SH, Niggebrugge C, Deglise P, Kljucevic P et al. (2006). Involvement of alpha-bungarotoxin-sensitive nicotinic receptors in long-term memory formation in the honeybee (Apis mellifera). Neurobiol Learn Mem 86: 164–174.
Gehle VM, Walcott EC, Nishizaki T, Sumikawa K (1997). N-glycosylation at the conserved sites ensures the expression of properly folded functional ACh receptors. Brain Res Mol Brain Res 45: 219–229.
Gisselmann G, Plonka J, Pusch H, Hatt H (2004). Drosophila melanogaster GRD and LCCH3 subunits form heteromultimeric GABA-gated cation channels. Br J Pharmacol 142: 409–413.
Gisselmann G, Pusch H, Hovemann BT, Hatt H (2002). Two cDNAs coding for histamine-gated ion channels in D. melanogaster. Nat Neurosci 5: 11–12.
Grauso M, Reenan RA, Culetto E, Sattelle DB (2002). Novel putative nicotinic acetylcholine receptor subunit genes, Dalpha5, Dalpha6 and Dalpha7, in Drosophila melanogaster identify a new and highly conserved target of adenosine deaminase acting on RNA-mediated A-to-I pre-mRNA editing. Genetics 160: 1519–1533.
Hopfield JF, Tank DW, Greengard P, Huganir RL (1988). Functional modulation of the nicotinic acetylcholine receptor by tyrosine phosphorylation. Nature 336: 677–680.
Hulo N, Bairoch A, Bulliard V, Cerutti L, De Castro E, Langendijk-Genevaux PS et al. (2006). The PROSITE database. Nucleic Acids Res 34: D227–D230.
Iovchev M, Kodrov P, Wolstenholme AJ, Pak WL, Semenov EP (2002). Altered drug resistance and recovery from paralysis in Drosophila melanogaster with a deficient histamine-gated chloride channel. J Neurogenet 16: 249–261.
Jensen ML, Schousboe A, Ahring PK (2005). Charge selectivity of the Cys-loop family of ligand-gated ion channels. J Neurochem 92: 217–225.
Jepson JE, Reenan RA (2007). Genetic approaches to studying adenosine-to-inosine RNA editing. Methods Enzymol 424: 265–287.
Jeschke P, Nauen R (2008). Neonicotinoids-from zero to hero in insecticide chemistry. Pest Manag Sci 64: 1084–1098.
Jin Y, Tian N, Cao J, Liang J, Yang Z, Lv J (2007). RNA editing and alternative splicing of the insect nAChR subunit alpha6 transcript: evolutionary conservation, divergence and regulation. BMC Evol Biol 7: 98.
Jones AK, Brown LA, Sattelle DB (2007). Insect nicotinic acetylcholine receptor gene families: from genetic model organism to vector, pest and beneficial species. Invert Neurosci 7: 67–73.
Jones AK, Grauso M, Sattelle DB (2005a). The nicotinic acetylcholine receptor gene family of the malaria mosquito, Anopheles gambiae. Genomics 85: 176–187.
Jones AK, Marshall J, Blake AD, Buckingham SD, Darlison MG, Sattelle DB (2005b). Sgbeta1, a novel locust (Schistocerca gregaria) non-alpha nicotinic acetylcholine receptor-like subunit with homology to the Drosophila melanogaster Dbeta1 subunit. Invert Neurosci 5: 147–155.
Jones AK, Raymond-Delpech V, Thany SH, Gauthier M, Sattelle DB (2006). The nicotinic acetylcholine receptor gene family of the honey bee, Apis mellifera. Genome Res 16: 1422–1430.
Jones AK, Sattelle DB (2006). The cys-loop ligand-gated ion channel superfamily of the honeybee, Apis mellifera. Invert Neurosci 6: 123–132.
Jones AK, Sattelle DB (2007). The cys-loop ligand-gated ion channel gene superfamily of the red flour beetle, Tribolium castaneum. BMC Genomics 8: 327.
Kondrashov FA, Koonin EV (2001). Origin of alternative splicing by tandem exon duplication. Hum Mol Genet 10: 2661–2669.
Lansdell SJ, Millar NS (2000). Cloning and heterologous expression of Dalpha4, a Drosophila neuronal nicotinic acetylcholine receptor subunit: identification of an alternative exon influencing the efficiency of subunit assembly. Neuropharmacology 39: 2604–2614.
Lansdell SJ, Millar NS (2002). Dbeta3, an atypical nicotinic acetylcholine receptor subunit from Drosophila: molecular cloning, heterologous expression and coassembly. J Neurochem 80: 1009–1018.
Littleton JT, Ganetzky B (2000). Ion channels and synaptic organization: analysis of the Drosophila genome. Neuron 26: 35–43.
Liu X, Krause WC, Davis RL (2007). GABAA receptor RDL inhibits Drosophila olfactory associative learning. Neuron 56: 1090–1102.
Liu Z, Han Z, Zhang Y, Song F, Yao X, Liu S et al. (2009). Heteromeric co-assembly of two insect nicotinic acetylcholine receptor alpha subunits: influence on sensitivity to neonicotinoid insecticides. J Neurochem 108: 498–506.
Lynch JA, Desplan C (2006). A method for parental RNA interference in the wasp Nasonia vitripennis. Nat Protoc 1: 486–494.
Matsuda K, Shimomura M, Ihara M, Akamatsu M, Sattelle DB (2005). Neonicotinoids show selective and diverse actions on their nicotinic receptor targets: electrophysiology, molecular biology, and receptor modeling studies. Biosci Biotechnol Biochem 69: 1442–1452.
Nishizaki T (2003). N-glycosylation sites on the nicotinic ACh receptor subunits regulate receptor channel desensitization and conductance. Brain Res Mol Brain Res 114: 172–176.
Page RD (1996). TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12: 357–358.
Pultz MA, Leaf DS (2003). The jewel wasp Nasonia: querying the genome with haplo-diploid genetics. Genesis 35: 185–191.
Raymond-Delpech V, Matsuda K, Sattelle BM, Rauh JJ, Sattelle DB (2005). Ion channels: molecular targets of neuroactive insecticides. Invert Neurosci 5: 119–133.
Sattelle DB, Jones AK, Sattelle BM, Matsuda K, Reenan R, Biggin PC (2005). Edit, cut and paste in the nicotinic acetylcholine receptor gene family of Drosophila melanogaster. Bioessays 27: 366–376.
Schnizler K, Saeger B, Pfeffer C, Gerbaulet A, Ebbinghaus-Kintscher U, Methfessel C et al. (2005). A novel chloride channel in Drosophila melanogaster is inhibited by protons. J Biol Chem 280: 16254–16262.
Shao YM, Dong K, Zhang CX (2007). The nicotinic acetylcholine receptor gene family of the silkworm, Bombyx mori. BMC Genomics 8: 324.
Shimomura M, Yokota M, Matsuda K, Sattelle DB, Komai K (2004). Roles of loop C and the loop B-C interval of the nicotinic receptor alpha subunit in its selective interactions with imidacloprid in insects. Neurosci Lett 363: 195–198.
Sine SM, Engel AG (2006). Recent advances in Cys-loop receptor structure and function. Nature 440: 448–455.
Smit AB, Brejc K, Syed N, Sixma TK (2003). Structure and function of AChBP, homologue of the ligand-binding domain of the nicotinic acetylcholine receptor. Ann NY Acad Sci 998: 81–92.
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997). The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25: 4876–4882.
Tsang SY, Ng SK, Xu Z, Xue H (2007). The evolution of GABAA receptor-like genes. Mol Biol Evol 24: 599–610.
Werren JH, Richards S, Desjardins CA, Niehuis O, Gadau J, Colbourne JK et al. (2010). Functional and evolutionary insights from the genomes of three parasitoid Nasonia species. Science 327: 343.
Zheng Y, Hirschberg B, Yuan J, Wang AP, Hunt DC, Ludmerer SW et al. (2002). Identification of two novel Drosophila melanogaster histamine-gated chloride channel subunits expressed in the eye. J Biol Chem 277: 2000–2005.
Acknowledgements
We are indebted to the Nasonia Genome Project (Human Genome Sequencing Center), which provided the starting point for this study. We thank Jack Werren for kindly providing us with N. vitripennis wasps. We thank Michael Williamson and Cornelis Grimmelikhuijzen for providing extra cDNA. This work was funded by the Medical Research Council (MRC), Dupont and the Wellcome Trust Integrative Physiology Initiative in Ion Channels (OXION).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Heredity website (http://www.nature.com/hdy)
Rights and permissions
About this article
Cite this article
Jones, A., Bera, A., Lees, K. et al. The cys-loop ligand-gated ion channel gene superfamily of the parasitoid wasp, Nasonia vitripennis. Heredity 104, 247–259 (2010). https://doi.org/10.1038/hdy.2009.97
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2009.97
Keywords
This article is cited by
-
Transcriptome profiling analysis of vesicular acetylcholine transporter (vacht) functions and signaling pathways in Tribolium castaneum
International Journal of Tropical Insect Science (2023)
-
The genome of the stable fly, Stomoxys calcitrans, reveals potential mechanisms underlying reproduction, host interactions, and novel targets for pest control
BMC Biology (2021)
-
In silico identification and assessment of insecticide target sites in the genome of the small hive beetle, Aethina tumida
BMC Genomics (2020)
-
Identification of transcriptome and fluralaner responsive genes in the common cutworm Spodoptera litura Fabricius, based on RNA-seq
BMC Genomics (2020)
-
The whole genome sequence of the Mediterranean fruit fly, Ceratitis capitata (Wiedemann), reveals insights into the biology and adaptive evolution of a highly invasive pest species
Genome Biology (2016)