Abstract
Chemoreception is a crucial biological process that is essential for the survival of animals. In insects, olfaction allows the organism to recognise volatile cues that allow the detection of food, predators and mates, whereas the sense of taste commonly allows the discrimination of soluble stimulants that elicit feeding behaviours and can also initiate innate sexual and reproductive responses. The most important proteins involved in the recognition of chemical cues comprise moderately sized multigene families. These families include odorant-binding proteins (OBPs) and chemosensory proteins (CSPs), which are involved in peripheral olfactory processing, and the chemoreceptor superfamily formed by the olfactory receptor (OR) and gustatory receptor (GR) families. Here, we review some recent evolutionary genomic studies of chemosensory gene families using the data from fully sequenced insect genomes, especially from the 12 newly available Drosophila genomes. Overall, the results clearly support the birth-and-death model as the major mechanism of evolution in these gene families. Namely, new members arise by tandem gene duplication, progressively diverge in sequence and function, and can eventually be lost from the genome by a deletion or pseudogenisation event. Adaptive changes fostered by environmental shifts are also observed in the evolution of chemosensory families in insects and likely involve reproductive, ecological or behavioural traits. Consequently, the current size of these gene families is mainly a result of random gene gain and loss events. This dynamic process may represent a major source of genetic variation, providing opportunities for FUTURE specific adaptations.
Similar content being viewed by others
Introduction
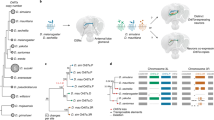
Animals have specialised sense organs that detect stimuli from the external environment, including visual, acoustic, tactile and chemical signals. Chemical signals are detected by one of two major chemosensory mechanisms: olfaction and taste. The olfactory system allows the organism to recognise volatile cues that confer the capacity to detect food, predators and mates. The sense of taste commonly allows the discrimination of soluble stimulants that elicit feeding behaviours, and it can also initiate innate sexual and reproductive responses. In insects, the early chemoreception steps—that is, those involving primary contact with chemical signals and the activation of signalling pathways—occur in porous chemosensory hairs (the sensilla) (Pelosi, 1996; Hildebrand and Shepherd, 1997; Shanbhag et al., 1999; Stengl et al., 1999) (Figure 1). These main events include: (i) the uptake of signal molecules from the external environment; (ii) transport (diffusion) through the sensory hair and (iii) interaction with the chemoreceptor, which in turn activates the cascade of events leading to spike activity in sensory neurons. The most important proteins underlying these processes are encoded in moderately sized multigene families. These families encode the odorant-binding proteins (OBPs) and chemosensory proteins (CSPs) involved in peripheral olfactory processing and a chemoreceptor superfamily formed by the olfactory receptors (ORs) and gustatory receptors (GRs).
(a) Schematic representation of the general structure of an insect olfactory hair. Gustatory sensilla have a similar structure, with only a single pore at the top of the sensory hair. (b) The first molecular steps (perireceptor events) of the insect chemosensory signalling transduction pathway. This figure depicts a general, simplified functional scheme; alternative schemes for OBP activity have been proposed (see Vogt, 2005).
Here, we review several recent comparative evolutionary analyses of the chemosensory multigene families from fully sequenced insect genomes, with a special emphasis on the 12 newly available Drosophila genomes (Figure 2). We address fundamental questions concerning the evolutionary dynamics of these gene families, such as the origin and fate of the gene repertoire, the impact of natural selection and the species-specific features of chemoreceptor evolution associated with ecological adaptation.
Accepted tree topology for the Drosophila and three other insect species surveyed. Divergence times are given in millions of years. Right: numbers of putative functional genes and pseudogenes (in parentheses). Data from Hill et al. (2002); Forêt and Maleszka (2006); Guo and Kim (2007); McBride and Arguello (2007); Nozawa and Nei (2007); Vieira et al. (2007); Engsontia et al. (2008); Gardiner et al. (2008); and Richards et al. (2008).
Insect chemosensory gene families
Odorant-binding proteins make up the majority of the protein in the sensillar fluid of insects and in the olfactory mucosa of vertebrates (Vogt and Riddiford, 1981; Pelosi, 1994). Surprisingly, in spite of their similar global function, vertebrate and insect OBPs are not homologous (Pelosi and Maida, 1995; Tegoni et al., 2000). Insect OBPs are small globular proteins (about 135–220 amino acids long) that bind and solubilise hydrophobic odorants such as pheromones. OBPs are synthesised in the accessory cells surrounding neurons and are subsequently secreted into the hydrophilic extracellular space. These proteins are characterised by a specific domain that comprises six α-helices joined by three disulphide bonds (Leal et al., 1999; Scaloni et al., 1999). Although the specific physiological roles of OBPs are not well established, it is believed that they act as molecular carriers that transport and deliver odorants to chemoreceptors located on the sensory neurons. In addition, OBPs might be involved in establishing the olfactory code (Van den Berg and Ziegelberger, 1991; Maida et al., 2003; Pophof, 2004; Matsuo et al., 2007; Laughlin et al., 2008) as well as in stimulus inactivation (Pelosi and Maida, 1995; Ziegelberger, 1995; Kaissling, 2001). Furthermore, gene expression analyses in a number of insect species indicate that OBPs are not restricted to olfactory and gustatory tissues and may also participate in other physiological functions (Pelosi et al., 2006 for a review).
Chemosensory proteins comprise another class of small soluble binding proteins (about 130 amino acids long), which are secreted into the lymph of insect chemosensory sensilla (Angeli et al., 1999). CSPs are more conserved, with a specific motif of four cysteines that form two disulphide bridges between neighbouring residues. This arrangement differs from that of OBPs, in which disulphide bridges are inter-helical and make two small loops to form a more rigid structure (Angeli et al., 1999; Leal et al., 1999; Scaloni et al., 1999). However, some CSPs can be identified by Hidden Markov Model (HMM)-based searches using information from the OBP sequence alignment as a statistical descriptor. Furthermore, the three-dimensional structures of some CSPs and OBPs can be superimposed with significant score values using structure alignment methods (Vieira and Rozas, unpublished results). Hence, in spite of differences in amino acid sequence and three-dimensional structure, CSPs and OBPs might be homologous (derived from a common ancestor). Alternatively, the current OBP–CSP similarity might result from convergent evolution. Although CSPs have been identified in insect chemosensory sensilla, there is no clear evidence that they participate in olfaction or gustation. Nevertheless, several CSPs are highly expressed in the sensillar lymph and, in vitro, are capable to bind different components of the pheromonal blends (Pelosi et al., 2006). It is again worth emphasising that not all CSPs are restricted to chemosensory organs, and it has been postulated that they are involved in carbon dioxide detection, larval development and leg regeneration (reviewed in Wanner et al., 2004).
The insect chemoreceptor superfamily comprises two distant and highly variable protein families, the ORs and GRs (Clyne et al., 1999, 2000; Gao and Chess, 1999; Vosshall et al., 1999; Fox et al., 2001; Scott et al., 2001; Hill et al., 2002; Robertson and Wanner, 2006; Engsontia et al., 2008). They are seven-transmembrane domain receptor proteins of about 400 amino acids that bind environmental compounds, thereby transforming the chemical signal into the activation of neurons in the higher processing centres in the brain, which in turn mediate the appropriate behaviour. As in OBPs, these proteins do not seem to be homologous to their functionally similar vertebrate counterparts (Hallem et al., 2006: Nei et al., 2008). In fact, insect ORs have an inverted membrane topology, with the C-terminus at the extracellular surface (Benton et al., 2006), which unlike vertebrate classic G-protein-coupled receptors, appear to form odorant-gated cation channels (Sato et al., 2008; Wicher et al., 2008). In Drosophila, ORs are expressed in antennae and maxillary palp, whereas GRs are expressed mainly in the gustatory organs (proboscis, legs and wings) but also in some olfactory structures. Interestingly, most OR neurons express two receptor genes, a specific Or and the highly conserved Or83b co-receptor, which form functional heterodimeric units in the dendritic membrane (Larsson et al., 2004). The ORs define the odorant specificity of olfactory neurons and modulate the neuronal response dynamics; the generated signal is transmitted by the axons of the sensory neurons to specific precise olfactory glomeruli in the antennal lobe (Hallem et al., 2006). The molecular function of GR proteins is less clear. Although multiple GR genes are expressed in a single GR neuron, OR83b is always absent, and it is unclear whether the GRs can function as ion channels in the absence of this co-receptor. Neurons responsive to soluble chemicals such as sugars, amino acids and repulsive (bitter-tasting) compounds, as well as carbon dioxide (Jones et al., 2007) and pheromones (Bray and Amrein, 2003), express distinct subsets of GR genes. These gustatory sensory neurons connect to different ganglia in the Drosophila brain (Amrein and Thorne, 2005).
Genomic organisation and phylogenetic analysis
The comparative genomic analysis of 12 Drosophila species (Clark et al., 2007) has provided the most exhaustive, fine-scale survey of insect chemosensory gene families to date. Putative functional and non-functional members of these families in the 12 genomes have been comprehensively identified by sequence similarity algorithms (Guo and Kim, 2007; Nozawa and Nei, 2007; Vieira et al., 2007; Gardiner et al., 2008). In some cases, the uncovered genome sequence has been confirmed by DNA re-sequencing (for example, McBride and Arguello, 2007). These studies reveal that there are an average of 49 putative functional OBPs, four CSPs, 63 ORs and 60 GRs per species across the Drosophila genus (Figure 2), with low variation in the number of genes in each family.
The chromosomal clustering of OBP genes, first observed in D. melanogaster (Galindo and Smith, 2001; Hekmat-Scafe et al., 2002; Vogt et al., 2002) occurs in all 12 Drosophila genomes (Vieira et al., 2007) (Figure 3). In Drosophila, 69% of OBP genes are arranged in 10 clusters of 2–6 genes; this organisation is significantly more conserved across the genus than expected by chance (Vieira and Rozas, unpublished results). Interestingly, the clustered arrangement is also maintained in other insect genomes (Xu et al., 2003; Forêt and Maleszka 2006). Drosophila OR and GR genes, on the other hand, appear more scattered throughout the genome, having only a few clusters (Robertson et al., 2003); this distribution differs from other insects where the receptor genes are arranged in a number of clusters (Robertson and Wanner, 2006; Bohbot et al., 2007; Engsontia et al., 2008). Indeed, the chromosomal distribution of OR genes have revealed repeated inter-chromosomal translocation events across the Drosophila phylogeny; these evolutionary events seem to be more frequent in this receptor family than in the OBP family (Guo and Kim, 2007; Conceição and Aguadé, 2008). The distinct gene expression patterns of the OBP and OR/GR families might account for these differences. In D. melanogaster, diverse OBPs are synthesised in the same supporting cells and then secreted to a restricted group of specialised sensilla. Moreover, the same OBPs can be expressed in several types of olfactory hairs and even in non-sensory tissues (Galindo and Smith, 2001; Shanbhag et al., 2001; Hekmat-Scafe et al., 2002). In contrast, olfactory and gustatory receptors are expressed in a much more precise pattern, with each sensory neuron deterministically expressing only a few specific chemoreceptor genes (generally not more than two). These distinct expression patterns might be explained by a different number, size or distribution of regulatory elements. Hence, the origin and the evolutionary fate of newly duplicated genes might be conditioned by the different regulatory architecture of OBPs and OR/GRs.
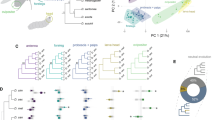
Dot-plot analysis of the chromosomal cluster that includes Obp50 paralogues. Top: phylogenetic relationships in Drosophila yakuba, Drosophila willistoni and Drosophila virilis. Left: phylogenetic relationships in D. melanogaster. Coloured and open arrows indicate OBP and non-OBP genes, respectively. Coloured boxes represent orthologous or co-orthologous OBP regions.
The Drosophila OBPs have been classified into two highly variable phylogenetic groups (or subfamilies; Vieira et al., 2007): Classic (six cysteines, one OBP domain) and Plus-C (12 cysteines and one characteristic proline, one OBP domain) (Figure 4). The Classic OBPs have been further divided into other subgroups, including the Minus-C group (usually with only four cysteines) and the Dimer OBPs (large OBP proteins formed by two consecutive Minus-C OBP domains) (Hekmat-Scafe et al., 2002). In contrast, Drosophila CSPs form a single conserved class of binding proteins with only four members in Drosophila. The OR genes of Drosophila have been sorted into 15 phylogenetic groups (Nozawa and Nei, 2007), each with a variable number of paralogous members. The OR83b subfamily, with a single member in each Drosophila species, is old and highly conserved across insects (Guo and Kim, 2007; McBride and Arguello, 2007; Nozawa and Nei, 2007; Gardiner et al., 2008). Current phylogenetic information for GRs in D. melanogaster group (McBride and Arguello, 2007) allows the classification of these genes into eight phylogenetic subfamilies. The putative volatile cabon dioxide and sweet taste receptor subfamilies form a well-supported phylogenetic clade, whereas the putative bitter taste and pheromone receptors are scattered across the GR phylogenetic tree, with representatives in several divergent subfamilies.
Phylogenetic relationships of insect OBPs. D. melanogaster and Drosophila mojavensis are depicted in red; A. gambiae, T. castaneum and A. mellifera are depicted in blue, green and orange, respectively. The scale bar represents 0.5 amino acid substitutions per site. (a) Global phylogenetic analysis. The main subfamilies are highlighted in dark grey (Classic), black (Plus-C) and light grey (Atypical). (b) Detailed phylogenetic relationships of the Obp59a orthologous group.
The integration of the genome information from Drosophila, Anopheles gambiae, Tribolium castaneum and Apis mellifera uncovers extensive lineage-specific gene duplications. Indeed, all chemosensory families exhibit clades that include multiple paralogues from the same species. Furthermore, orthologous relationships are difficult to infer, and the statistical phylogenetic support for the earlier defined Drosophila subfamilies is very low. For instance, only two OBP genes, Obp59a and Obp73a (unpublished results), have clear orthologues across insects (except in A. mellifera) (Figure 4). OBP Plus-C genes are present in flies, mosquitoes and beetles showing some lineage-specific duplications (T. castaneum has a single member). Classic OBPs are found in all surveyed genomes and also display some expansions, as the Minus-C subfamily in T. castaneum; A. mellifera, in contrast, has few OBP family members, many of which are clustered into a single, bee-specific, Minus-C clade. Interestingly, Tribolium and Apis Minus-C proteins might have originated independently from Classic OBP ancestors. Phylogenetic analysis also reveals a new, strongly supported monophyletic group within this subfamily, the Atypical OBP genes (Xu et al., 2003). Atypical OBPs are present only in Anopheles and might have evolved recently in this species.
The CSP gene family shows a similar evolutionary pattern. Dipterans and bees have a reduced number of genes (four to seven), although the family is expanded (∼20 genes) in T. castaneum (Forêt et al., 2007; Vieira and Rozas, unpublished results). The OR multigene family also has many members grouped in species-specific lineages, thereby revealing a number of gene duplication events. The only unequivocally orthologous group shared by all insects comprises the highly conserved Or83b gene, placed at the base of the OR family tree. Like the other chemosensory families, most GR families are unique and species specific. Only the carbon dioxide, sweet taste and a few orphan receptors are conserved. Compared with other insect genomes, the bee genome encodes very few GRs (only 10), distributed across seven phylogenetic lineages. Interestingly, no carbon dioxide receptor has been found in this species.
Birth-and-death evolution
Genomic evidence supports unequal crossing over as the main mechanism that generates tandem gene duplications of the chemosensory genes; in Drosophila, physically neighbouring members of these families are also phylogenetically related (Guo and Kim, 2007; Nozawa and Nei, 2007; Vieira et al., 2007). In particular, new OBP duplicates are usually detected in extant chromosomal clusters, and the most closely related genes are located in the same cluster (Figure 3) (Vieira et al., 2007). Amino acid-based phylogenetic analyses show that paralogues share common ancestors much older than those of orthologous groups, which likely predate the origin of the insects (Figure 4a). Moreover, gene trees and species trees are reconciled within the orthologous groups (Figure 4b). This strongly suggests that the genes of the chemosensory family have diverged independently since originating by gene duplication.
Across Drosophila, chemosensory families are very similar in number of genes and protein subfamily sizes; indeed, there are few examples of lineage-specific duplications occurring in short periods of time (see also; Bhutkar et al., 2007; Conceição and Aguadé, 2008). However, a comparison of chemosensory families in distantly related insects (Drosophila, Anopheles, Tribolium and Apis) reveals much more dramatic variation in gene family size and total number of genes. Although the gene copy number tends to be conserved across the chemosensory families of Drosophila, the comparative genomic analyses among these insects have identified an unexpectedly large number of gene gains/losses and pseudogenes (Robertson and Wanner, 2006; Forêt et al., 2007; Engsontia et al., 2008); these numbers are higher for the receptor superfamily than for OBPs. Interestingly, deletions and pseudogenisation events are not randomly distributed across the phylogeny. Instead, pseudogenisation events are mainly inferred in the external branches (Vieira et al., 2007), suggesting that the half lives of these pseudogenes are very short. Gene gains and losses are also unevenly distributed across Drosophila lineages. For instance, D. grimshawi has undergone the most dramatic number of episodes (27 OR and eight OBP gene gains, and 14 OR and five OBP gene losses), and D. melanogaster group has experienced a significant contraction of chemoreceptor genes (four new OR genes and 12 OR and 35 GR gene losses; McBride and Arguello, 2007).
Overall, recent genomic data clearly point to the birth-and-death (BD) model (Nei and Rooney 2005) as the major mechanism for the evolution of insect chemosensory genes (see also Roelofs and Rooney, 2003). Namely (i) many orthologous groups are identified at short-time scales (for example, for the evolution of the Drosophila genus) and fit the accepted phylogeny; (ii) there is no evidence of gene conversion between paralogues; (iii) gene families have undergone a number of gene gains and losses in many lineages, and several non-functional members (pseudogenes) can be found in many families and (iv) the distribution of phylogenetic subfamilies shows dissimilar patterns at short-time scales (across the Drosophila genus) and long-time scales (across insect species). Therefore, chemosensory genes likely originate by tandem gene duplication (resulting from unequal crossing over), evolve independently from each other and eventually would be lost from the genome by a deletion. Presumably, the observed pseudogenes would indicate that some of the inferred gene losses might have been initially triggered by a pseudogenisation event.
Analysis of gene copy-number variation across Drosophila allows an estimation of BD rates per gene and per million of years (De Bie et al., 2006; Hahn et al., 2007). Estimates of BD rates using divergence times from Tamura et al., (2004) are quite high (OBPs, λ=0.005; ORs, λ=0.006 and GRs, λ=0.011; McBride and Arguello, 2007; Vieira et al., 2007; Gardiner et al., 2008), larger than estimates for the complete genome (λ=0.0012; Hahn et al., 2007). These BD rates uncover a highly dynamic model for the evolution of chemosensory genes in which gene families are constantly renewed by the duplication of genes to replace lost or non-functional copies.
Functional diversification and natural selection
The BD process entails a progressive divergence among family members following their origin by gene duplication. The putative functional diversification associated with such sequence divergence has been studied by comparative analysis of synonymous (dS) and non-synonymous (dN) divergence (ω=dN/dS) (Forêt and Maleszka 2006; Guo and Kim, 2007; McBride, 2007; McBride and Arguello, 2007; Vieira et al., 2007; Gardiner et al., 2008). This widely used and powerful approach allows the inference of the selective pressure on protein coding sequences, and it also allows several competitive evolutionary scenarios to be explored and tested. Although ω estimates are relatively high for chemosensory genes (median values across the families range from 0.05 to 0.22), purifying selection seems to be the main force governing the evolution of chemosensory genes in the melanogaster group (Figure 5). Interestingly, there are significant selective constraint differences both among orthologous groups and across lineages, as well as some cases of episodic evolution and positive selection. CSPs are the most conserved family and, concordantly, the most constrained one. Nevertheless, there is considerable variation among orthologous groups (ω values range from 0.003 to 0.11; calculated from Vieira et al. (2007) data). There are also significant differences in functional constraints among OBP members (Vieira et al., 2007) (Figure 5). Of all Drosophila receptors, selective pressures are significantly weaker for GRs than for ORs, and they are also lower for receptor genes that have undergone duplications (Gardiner et al., 2008). The Or83b gene has the smallest ω ratio (ω=0.014), consistent with its strong conservation across insects and its essential functional role. Among GRs, the sweet taste and carbon dioxide receptors have low ω rates, whereas bitter taste members evolve rapidly (McBride and Arguello, 2007).
Distribution of the ω parameter across the main chemosensory families in the melanogaster subgroup. (a) OBPs; (b) ORs and (c) GRs. The box-plot indicates the median (waist) and the 25th and 75th percentiles (in boxes). Range bars denote data points within two standard deviations. Data from McBride and Arguello (2007) and Vieira et al. (2007).
Functional constraints are usually distributed heterogeneously along the coding region of the gene. In fact, the signal of positive selection has been inferred in some Drosophila OR and GR genes using likelihood models that account for such heterogeneity (Guo and Kim, 2007; McBride, 2007, McBride and Arguello, 2007). Although this approach did not allow the detection of positive selection in the Drosophila OBP family (Vieira et al., 2007), the use of more powerful analytical methodologies allowed to infer molecular adaptation in one member of this family (see Sánchez-Gracia and Rozas, 2008). Positive selection has also been proposed to explain the evolution of the Minus-C subfamily of A. mellifera (Forêt and Maleszka 2006). Interestingly, some of the amino acid changes predicted to have been driven by positive selection in the OBPs of Drosophila and the honeybee are located in the putative binding pocket of the protein. The impact of natural selection has also been assessed using DNA polymorphism and divergence data (McDonald and Kreitman, 1991). Using this methodology, McBride and Arguello (2007) showed that a considerable fraction of OR and GR genes deviated significantly from neutral expectations. This observation points to the action of positive selection after the D. simulans and D. melanogaster split. Once again, OR and GR genes experience dissimilar selective pressures at this short time scale; in particular, the GR family has lower values on the neutrality index (McDonald and Kreitman, 1991), suggesting a stronger impact of positive selection on GRs than ORs.
Specialist and generalist species of Drosophila also show contrasting functional constraint levels (McBride, 2007; McBride and Arguello, 2007; Vieira et al., 2007). Indeed, the strength of purifying selection on receptors and OBP genes is significantly lower in specialist species. Receptor genes in these species also exhibit higher loss rates, especially within the GR family. It is noted that the specific GR genes lost are also phylogenetically related, suggesting that the putative relaxation of selective pressure mainly affects functionally related GRs. Although ecological host specialisation might explain these findings (Matsuo et al., 2007; McBride and Arguello, 2007), they might also be promoted by species endemism (Gardiner et al., 2008). Furthermore, the differential pattern observed in the GR family along with the higher BD rates and ω values points to an elevated evolutionary rate of this family. The underlying biological explanation of this finding remains unclear, but it may be related to differences in the evolvability of taste and olfactory systems caused by their different genetic architectures. Alternatively, as taste is particularly relevant in the final recognition and selection of food, toxin avoidance and appropriate courtship behaviour, GRs might be an important target for molecular adaptation.
Concluding remarks
Altogether, the recent genomic data support the BD model for chemosensory family evolution, with progressive divergence and functional diversification among their members. In spite of high BD rates, the number of genes in each chemosensory family has remained fairly conserved across Drosophila, and a large fraction of members evolve under purifying selection. The actual family size would thereby result from a trade-off between the weak effect of the stochastic BD process (or random genomic drift; Nei, 2007) at short time scales and the maintenance of a core number of genes required for basal Drosophila chemosensory performance. The large variation in gene repertoire size observed among distantly related insects might thus be explained by genomic drift accumulated over long time scales. Indeed, gene gains and losses might provide the raw source of variation for evolutionary change. Given the crucial role of the chemosensory system in the survival and reproduction of individuals, adaptive changes likely arise in response to the demands of new environmental conditions. Molecular adaptation may entail, for instance, changes in the detection of pheromones (changes in chemical sensibility or specificity; for example, Willett, 2000), and it might be fostered by shifts in ecological interactions (for example, Matsuo et al., 2007; McBride, 2007; McBride and Arguello, 2007; Vieira et al., 2007; Gardiner et al., 2008) or even by changes in some aspects of social behaviour (for example, Krieger and Ross, 2002; Forêt and Maleszka 2006). Currently, it is very difficult to determine the evolutionary impact of random genomic drift and adaptation on the sizes of chemosensory families. Nevertheless, large gene family expansions or contractions at short time scales (especially of functionally related genes) and significant lineage-specific accelerations of the evolutionary rate likely reflect molecular adaptation. The recent genomic studies analysing closely related species with well-resolved phylogenies has provided valuable insight into evolutionary patterns and processes and illuminated features that usually are hidden in analyses using distantly related species or a small number of genes. These studies may be useful for clarifying the relative contribution of neutral mutation and natural selection to the BD evolution of chemosensory multigene families and in the molecular evolution of insects in general.
References
Amrein H, Thorne N (2005). Gustatory perception and behavior in Drosophila melanogaster. Curr Biol 15: 673–684.
Angeli S, Ceron F, Scaloni A, Monti M, Monteforti G, Minnocci A et al. (1999). Purification, structural characterization, cloning and immunocytochemical localization of chemoreception proteins from Schistocerca gregaria. Eur J Biochem 262: 745–754.
Benton R, Sachse S, Michnick SW, Vosshall LB (2006). Atypical membrane topology and heteromeric function of Drosophila odorant receptors in vivo. PLoS Biol 4: e20.
Bohbot J, Pitts RJ, Kwon H-W, Rützler M, Robertson HM, Zwiebel LJ (2007). Molecular characterization of the Aedes aegypti odorant receptor gene family. Insect Mol Biol 16: 525–537.
Bray S, Amrein H (2003). A putative Drosophila pheromone receptor expressed in male-specific taste neurons is required for efficient courtship. Neuron 39: 1019–1029.
Bhutkar A, Russo SM, Smith TF, Gelbart WM (2007). Genome-scale analysis of positionally relocated genes. Genome Res 17: 1880–1887.
Clark AG, Eisen MB, Smith DR, Bergman CM, Oliver B, Markow TA et al. (2007). Evolution of genes and genomes on the Drosophila phylogeny. Nature 450: 203–218.
Clyne PJ, Warr CG, Carlson JR (2000). Candidate taste receptors in Drosophila. Science 287: 1830–1834.
Clyne PJ, Warr CG, Freeman MR, Lessing D, Kim J, Carlson JR (1999). A novel family of divergent seven-transmembrane proteins: candidate odorant receptors in Drosophila. Neuron 22: 327–338.
Conceição IC, Aguade M (2008). High incidence of interchromosomal transpositions in the evolutionary history of a subset of Or genes in Drosophila. J Mol Evol 66: 325–332.
De Bie T, Cristianini N, Demuth JP, Hahn MW (2006). CAFE: a computational tool for the study of gene family evolution. Bioinformatics 22: 1269–1271.
Engsontia P, Sanderson AP, Cobb M, Walden KK, Robertson HM, Brown S (2008). The red flour beetle's large nose: an expanded odorant receptor gene family in Tribolium castaneum. Insect Biochem Mol Biol 38: 387–397.
Forêt S, Maleszka R (2006). Function and evolution of a gene family encoding odorant binding-like proteins in a social insect, the honey bee (Apis mellifera). Genome Res 16: 1404–1413.
Forêt S, Wanner KW, Maleszka R (2007). Chemosensory proteins in the honey bee: insights from the annotated genome, comparative analyses and expressional profiling. Insect Biochem Mol Biol 37: 19–28.
Fox AN, Pitts RJ, Robertson HM, Carlson JR, Zwiebel LJ (2001). Candidate odorant receptors from the malaria vector mosquito Anopheles gambiae and evidence of down-regulation in response to blood feeding. Proc Natl Acad Sci USA 98: 14693–14697.
Galindo K, Smith DP (2001). A large family of divergent Drosophila odorant-binding proteins expressed in gustatory and olfactory sensilla. Genetics 159: 1059–1072.
Gao Q, Chess A (1999). Identification of candidate Drosophila olfactory receptors from genomic DNA sequence. Genomics 60: 31–39.
Gardiner A, Barker D, Butlin RK, Jordan WC, Ritchie MG (2008). Drosophila chemoreceptor gene evolution: selection, specialization and genome size. Mol Ecol 17: 1648–1657.
Guo S, Kim J (2007). Molecular evolution of Drosophila odorant receptor genes. Mol Biol Evol 24: 1198–1207.
Hahn MW, Han MV, Han SG (2007). Gene family evolution across 12 Drosophila genomes. PLoS Genet 3: e197.
Hallem EA, Dahanukar A, Carlson JR (2006). Insect odor and taste receptors. Annu Rev Entomol 51: 113–135.
Hekmat-Scafe DS, Scafe CR, McKinney AJ, Tanouye MA (2002). Genome-wide analysis of the odorant-binding protein gene family in Drosophila melanogaster. Genome Res 12: 1357–1369.
Hildebrand JG, Shepherd GM (1997). Mechanisms of olfactory discrimination: converging evidence for common principles across phyla. Annu Rev Neurosci 20: 595–631.
Hill CA, Fox AN, Pitts RJ, Kent LB, Tan PL, Chrystal MA et al. (2002). G protein-coupled receptors in Anopheles gambiae. Science 298: 176–178.
Jones WD, Cayirlioglu P, Kadow IG, Vosshall LB (2007). Two chemosensory receptors together mediate carbon dioxide detection in Drosophila. Nature 445: 86–90.
Kaissling KE (2001). Olfactory perireceptor and receptor events in moths: a kinetic model. Chem Senses 26: 125–150.
Krieger MJ, Ross KG (2002). Identification of a major gene regulating complex social behavior. Science 295: 328–332.
Larsson MC, Domingos AI, Jones WD, Chiappe ME, Amrein H, Vosshall LB (2004). Or83b encodes a broadly expressed odorant receptor essential for Drosophila olfaction. Neuron 43: 703–714.
Laughlin JD, Ha TS, Jones DN, Smith DP (2008). Activation of pheromone-sensitive neurons is mediated by conformational activation of pheromone-binding protein. Cell 133: 1255–1265.
Leal WS, Nikonova L, Peng G (1999). Disulfide structure of the pheromone binding protein from the silkworm moth, Bombyx mori. FEBS Lett 464: 85–90.
Maida R, Ziegelberger G, Kaissling KE (2003). Ligand binding to six recombinant pheromone-binding proteins of Antheraea polyphemus and Antheraea pernyi. J Comp Physiol [B] 173: 565–573.
Matsuo T, Sugaya S, Yasukawa J, Aigaki T, Fuyama Y (2007). Odorant-binding proteins OBP57d and OBP57e affect taste perception and host-plant preference in Drosophila sechellia. PLoS Biol 5: e118.
McBride CS (2007). Rapid evolution of smell and taste receptor genes during host specialization in Drosophila sechellia. Proc Natl Acad Sci USA 104: 4996–5001.
McBride CS, Arguello JR (2007). Five Drosophila genomes reveal nonneutral evolution and the signature of host specialization in the chemoreceptor superfamily. Genetics 177: 1395–1416.
McDonald JH, Kreitman M (1991). Adaptive protein evolution at the Adh locus in Drosophila. Nature 351: 652–654.
Nei M (2007). The new mutation theory of phenotypic evolution. Proc Natl Acad Sci USA 104: 12235–12242.
Nei M, Niimura Y, Nozawa M (2008). The evolution of animal chemosensory receptor gene repertoires: roles of chance and necessity. Nat Rev Genet 9: 951–963.
Nei M, Rooney AP (2005). Concerted and birth-and-death evolution of multigene families. Annu Rev Genet 39: 121–152.
Nozawa M, Nei M (2007). Evolutionary dynamics of olfactory receptor genes in Drosophila species. Proc Natl Acad Sci USA 104: 7122–7127.
Pelosi P (1994). Odorant-binding proteins. Crit Rev Biochem Mol Biol 29: 199–228.
Pelosi P (1996). Perireceptor events in olfaction. J Neurobiol 30: 3–19.
Pelosi P, Maida R (1995). Odorant-binding proteins in insects. Comp Biochem Physiol B Biochem Mol Biol 111: 503–514.
Pelosi P, Zhou JJ, Ban LP, Calvello M (2006). Soluble proteins in insect chemical communication. Cell Mol Life Sci 63: 1658–1676.
Pophof B (2004). Pheromone-binding proteins contribute to the activation of olfactory receptor neurons in the silkmoths Antheraea polyphemus and Bombyx mori. Chem Senses 29: 117–125.
Roelofs WL, Rooney AP (2003). Molecular genetics and evolution of pheromone biosynthesis in Lepidoptera. Proc Natl Acad Sci USA 100: 9179–9184.
Richards S, Gibbs RA, Weinstock GM, Brown SJ, Denell R, Beeman RW et al. (2008). The genome of the model beetle and pest Tribolium castaneum. Nature 452: 949–955.
Robertson HM, Warr CG, Carlson JR (2003). Molecular evolution of the insect chemoreceptor gene superfamily in Drosophila melanogaster. Proc Natl Acad Sci USA 100: 14537–14542.
Robertson HM, Wanner KW (2006). The chemoreceptor superfamily in the honey bee, Apis mellifera: expansion of the odorant, but not gustatory, receptor family. Genome Res 16: 1395–1403.
Sanchez-Gracia A, Rozas J (2008). Divergent evolution and molecular adaptation in the Drosophila odorant-binding protein family: inferences from sequence variation at the OS-E and OS-F genes. BMC Evol Biol 8: 323.
Sato K, Pellegrino M, Nakagawa T, Nakagawa T, Vosshall LB, Touhara K (2008). Insect olfactory receptors are heteromeric ligand-gated ion channels. Nature 452: 1002–1006.
Scaloni A, Monti M, Angeli S, Pelosi P (1999). Structural analysis and disulfide-bridge pairing of two odorant-binding proteins from Bombyx mori. Biochem Biophys Res Commun 266: 386–391.
Scott K, Brady Jr R, Cravchik A, Morozov P, Rzhetsky A, Zuker C et al. (2001). A chemosensory gene family encoding candidate gustatory and olfactory receptors in Drosophila. Cell 104: 661–673.
Shanbhag SR, Hekmat-Scafe D, Kim MS, Park SK, Carlson JR, Pikielny C et al. (2001). Expression mosaic of odorant-binding proteins in Drosophila olfactory organs. Microsc Res Tech 55: 297–306.
Shanbhag SR, Müller B, Steinbrech RA (1999). Atlas of olfactory organs of Drosophila melanogaster 1. Types, external organization, innervation and distribution of olfactory sensilla. Int J Insect Morphol Embryol 28: 377–397.
Stengl M, Ziegelberger G, Boekhoff I, Krieger J (1999) Perireceptor events and transduction mechanisms in insect olfaction. In: Hansson BS (ed). Insect Olfaction. Springer: Berlin, Heidelberg, New York. pp 49–66.
Tamura K, Subramanian S, Kumar S (2004). Temporal patterns of fruit fly (Drosophila) evolution revealed by mutation clocks. Mol Biol Evol 21: 36–44.
Tegoni M, Pelosi P, Vincent F, Spinelli S, Campanacci V, Grolli S et al. (2000). Mammalian odorant binding proteins. Biochim Biophys Acta 1482: 229–240.
Van den Berg MJ, Ziegelberger G (1991). On the function of the pheromone binding protein in the olfactory hairs of Antheraea polyphemus. J Insect Physiol 37: 79–85.
Vieira FG, Sanchez-Gracia A, Rozas J (2007). Comparative genomic analysis of the Odorant-binding protein family in 12 Drosophila genomes: purifying selection and birth-and-death evolution. Genome Biol 8: R235.
Vogt RG, Riddiford LM (1981). Pheromone binding and inactivation by moth antennae. Nature 293: 161–163.
Vogt RG, Rogers ME, Franco MD, Sun M (2002). A comparative study of odorant binding protein genes: differential expression of the PBP1-GOBP2 gene cluster in Manduca sexta (Lepidoptera) and the organization of OBP genes in Drosophila melanogaster (Diptera). J Exp Biol 205: 719–744.
Vogt RG (2005). Molecular Basis of Pheromone Detection in Insects. In Comprehensive Insect Physiology, Biochemistry, Pharmacology and Molecular Biology. Gilbert LI, Iatro K, Gill S (eds). Endocrinology Vol 3, Elsevier: London. pp 753–804.
Vosshall LB, Amrein H, Morozov PS, Rzhetsky A, Axel R (1999). A spatial map of olfactory receptor expression in the Drosophila antenna. Cell 96: 725–736.
Wanner KW, Willis LG, Theilmann DA, Isman MB, Feng Q, Plettner E (2004). Analysis of the insect OS-D-like gene family. J Chem Ecol 30: 889–911.
Wicher D, Schafer R, Bauernfeind R, Stensmyr MC, Heller R, Heinemann SH et al. (2008). Drosophila odorant receptors are both ligand-gated and cyclic-nucleotide-activated cation channels. Nature 452: 1007–1011.
Willett CS (2000). Evidence for directional selection acting on pheromone-binding proteins in the genus Choristoneura. Mol Biol Evol 17: 553–562.
Xu PX, Zwiebel LJ, Smith DP (2003). Identification of a distinct family of genes encoding atypical odorant-binding proteins in the malaria vector mosquito, Anopheles gambiae. Insect Mol Biol 12: 549–560.
Ziegelberger G (1995). Redox-shift of the pheromone-binding protein in the silkmoth Antheraea polyphemus. Eur J Biochem 232: 706–711.
Acknowledgements
This work was supported by grant BFU2007-62927 from the Dirección General de Investigación Científica y Técnica (Spain) and by grant 2005SGR00166 from Comissió Interdepartamental de Recerca i Innovació Tecnològica de Catalunya (Spain).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sánchez-Gracia, A., Vieira, F. & Rozas, J. Molecular evolution of the major chemosensory gene families in insects. Heredity 103, 208–216 (2009). https://doi.org/10.1038/hdy.2009.55
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2009.55