Abstract
Powerful and accurate detection of first-generation (F1) hybrids and backcrosses in nature is needed to achieve a better understanding of the function and dynamics of introgression. To document the frequency of ongoing interspecific gene exchange between two Mediterranean evergreen oaks, the cork oak (Quercus suber) and the holm oak (Q. ilex), we analyzed 1487 individuals originating from across the range of the two species using eight microsatellite loci and two Bayesian clustering approaches (implemented in the programs STRUCTURE and NEWHYBRIDS). Simulated data were used to assess the differences between the two clustering methods and to back up the choice of the threshold value for the posterior probability to discriminate admixed from pure individuals. We found that the use of STRUCTURE resulted in the highest power to detect hybrids, whereas NEWHYBRIDS provided the highest accuracy. Irrespective of the approach, the two species were clearly distinguished as independent genetic entities without any prior information. In contrast with previous reports, we found no evidence for unidirectional introgression. The overall hybridization rate was very low (<2% of introgressed individuals). Only two individuals were identified as F1 hybrids and five as early backcrosses. This work shows that the combined application of the two complementary Bayesian approaches and their systematic validation with simulations, fit for the case at hand, helps gain resolution in the identification of admixed individuals.
Similar content being viewed by others
Introduction
Natural hybridization and introgression are widespread phenomena in plants, with important evolutionary implications (Rieseberg and Carney, 1998). The movement of genes across species boundaries can promote the appearance of new lineages (Seehausen, 2004), adaptive solutions (Rieseberg et al., 2003) or colonization abilities (Potts and Reid, 1988; Petit et al., 2004). Measuring the frequency of hybrids and describing their geographic distribution should help focus measures directed to conservation or breeding programs (Burgess et al., 2005; Kothera et al., 2007). Different types of molecular markers can inform on different spatial and temporal scales of the hybridization–introgression dynamics. Chloroplast and mtDNA have been used to describe past episodes of introgression (Palmé et al., 2004; Heuertz et al., 2006) whereas nuclear loci have been useful to infer contemporary rates of interspecific gene exchange (Lexer et al., 2005; Fernández-Manjarrés et al., 2006). However, identifying hybrid individuals in nature using molecular markers still represents an important challenge. Availability of hypervariable codominant markers (for example, microsatellites) and powerful statistical procedures (that is, Bayesian clustering methods, which do not rely on a priori morphological classification) has facilitated the detection of first-generation (F1) hybrids and backcrosses. However, the choice of the method that will provide the best resolution needs to be established for a given situation.
Oaks represent good models for such studies. Interspecific hybridization is the most frequently invoked mechanism to account for the existence of plants morphologically and ecologically intermediate between extant oak species (Jensen et al., 1993; Howard et al., 1997; González-Rodríguez et al., 2004) and to interpret the extensive local sharing of organelle and nuclear genes between species (Whittemore and Schaal, 1991; Howard et al., 1997; Petit et al., 1997; Dumolin-Lapègue et al., 1999). However, in some cases, interspecific gene exchanges have been detected with molecular markers in the absence of obvious morphologically intermediate forms (Whittemore and Schaal, 1991; Dodd and Afzal-Rafii, 2004). Moreover, the possibility that shared alleles represent ancestral segregating polymorphisms rather than the outcome of hybridization has been suggested (Muir and Schlötterer, 2005; but see Lexer et al., 2006). Environmental variation, disturbance as well as the degree of contact between species can affect the frequency and the spatial distribution of hybrids in natural oak populations (Nason, 1992; Rushton, 1993; Howard et al., 1997; Dumolin-Lapègue et al., 1999; Dodd and Afzal-Rafii, 2004; Tovar-Sanchez and Oyama, 2004; Curtu et al., 2007; Valbuena-Carabaña et al., 2007). Although hybridization between some oak species, such as the closely related species Quercus robur and Q. petraea, has been analyzed extensively for nuclear, chloroplast and mitochondrial variation, our understanding of the underlying processes is still unclear.
In this study we focus on two distantly related oak species, Q. suber (cork oak) and Q. ilex (holm oak), which have partially overlapping geographic distributions in the western part of the Mediterranean basin. The two evergreen species have a major ecological function in many Mediterranean woody ecosystems and constitute key elements of seminatural systems of high economical and social importance (for example, cork extraction and silvopastoral uses; Plieninger et al., 2003; Martín Vicente and Fernández Alés, 2006). Cork oaks and holm oaks are easily discriminated by a few morphological traits, including bark (that is, cork layer is found exclusively in Q. suber), leaf and fruit features (Amaral Franco, 1990). Some concerns exist about the effect of hybridization on cork quality and on breeding programs of Q. suber (Oliveira et al., 2007). Within the section Cerris (subgenus Quercus), Q. suber and Q. ilex belong to different clades (groups Cerris and Ilex, respectively), which are thought to have diverged during the middle Tertiary (Manos et al., 2001). Despite their deep phylogenetic divergence, clearly supported by internal transcribed spacer, amplified fragment length polymorphisms and isozyme variation (Manos et al., 1999; Toumi and Lumaret, 2001; Bellarosa et al., 2005; López de Heredia et al., 2007b), hybridization has been inferred on the basis of morphological and molecular markers (Elena-Rosselló et al., 1992; Toumi and Lumaret, 1998; Lumaret et al., 2002; Oliveira et al., 2003; Bellarosa et al., 2005). Furthermore, extensive surveys of chloroplast DNA diversity of both species and of other relatives (such as Q. coccifera) across the whole distribution range have demonstrated widespread cytoplasmic introgression, mainly localized along a northeast-southwest line, from French Catalonia and eastern Iberia to Morocco (reviewed in Lumaret et al., 2005). Interspecific exchanges seem to be limited to introgression of Q. ilex cpDNA and mtDNA into Q. suber, with only very few cases of Q. suber cpDNA introgressing into Q. ilex (Belahbib et al., 2001; Lumaret et al., 2002; Jiménez et al., 2004; Staudt et al., 2004). Because organelle DNA is maternally inherited in Quercus (Dumolin et al., 1995), this asymmetry implies that Q. ilex has acted predominantly as the maternal species in interspecific crosses. Boavida et al. (2001) provided experimental support for this hypothesis by showing that F1 hybrids are more easily produced when Q. suber is the pollen donor. In addition, unidirectional mating can be favored by phenology (Q. ilex flowers earlier) combined with protandry (that is, male flowers appear earlier than female flowers; Varela and Valdiviesso, 1996).
To date, no data are available on mating preferences in later hybrid generations, as hybrid individuals with known pedigree remain extremely rare in oaks. In such a context, identifying F1 hybrids and backcrosses would be important, particularly when the proportion of hybrid individuals is low and when they are morphologically cryptic (as seems to be the case for Q. suber and Q. ilex; Lumaret et al., 2002; Staudt et al., 2004). We present here a broad-scale survey of molecular variation across the overlapping range of Q. suber and Q. ilex to explore the extent and pattern of nuclear introgressive hybridization, using a panel of eight highly discriminating microsatellite loci. Our specific aims are (1) to assess the effectiveness of two Bayesian clustering approaches to distinguish hybrid individuals without knowledge of their pedigree and (2) to document the frequency of contemporary interspecific gene exchange in natural populations of cork and holm oaks, and hence evaluate previously proposed hybridization scenarios. For these purposes, we use admixture analysis of multilocus microsatellite genotypes from a range-wide sample of sympatric and allopatric populations of the two species. Furthermore, we simulate hybrid genotypes to assess the performance and the limits of the procedure used to detect hybrid individuals and to distinguish among hybrid classes.
Materials and methods
Sampling strategy
We sampled 597 Q. suber and 515 Q. ilex from 13 populations across the distribution range of cork oak and the overlapping range of holm oak (Figure 1). Five mixed woods were more intensively sampled (775 individuals). Two of them (Castilla-La Mancha and Sicily) include part of the individuals used in Soto et al. (2007) and Burgarella et al. (2007). In the mixed population of Minorca, the sample includes all existing cork oaks on the island (67 individuals). As additional reference, another set of 375 cork oaks have been included, sampled from an international provenance trial established in 1998 in the frame of the Q. suber network from the European Programme for the Conservation of Forest Genetic Resources (EUFORGEN), which covered the complete distribution range of the species (35 provenances). Reference codes, geographic allocations and sampling sizes are given in Table 1. Individuals were tentatively assigned to each species according to their morphology.
Sampling sites. Light gray, distribution range of Quercus suber (modified from http://www.bioversityinternational.org/networks/euforgen/); dashed line, distribution range of Q. ilex; triangles, populations included in the field trial and dark gray circles, Q. ilex, Q. suber and mixed stands (see Table 1 for population code).
Microsatellite typing
Individuals were genotyped at eight microsatellite loci: MSQ4, MSQ13 (Dow et al., 1995), QpZAG9, QpZAG15, QpZAG36, QpZAG46 (Steinkellner et al., 1997), QrZAG11 and QrZAG20 (Kampfer et al., 1998). A detailed description of the protocols has been published elsewhere (Soto et al., 2003, 2007). At MSQ13, 25% of Q. ilex genotypes had three or four alleles, possibly due to gene duplication in this species. On the contrary, Q. suber showed a normal banding pattern. MSQ13 is a highly informative locus, because allele sizes do not overlap between the two species (Soto et al., 2003). To include this locus in the following analyses, we pooled the alleles typical of Q. ilex. To identify them, we defined the pure genotype pool of each species with the other seven loci, performing a preliminary clustering analysis with STRUCTURE (same settings described below).
Information content of microsatellites and genetic differentiation
Deviation from Hardy–Weinberg equilibrium and linkage disequilibrium (LD) was tested using FSTAT 2.9.3.2 (Goudet, 2001). To assess the diagnostic power of each marker, we estimated the allele frequency differential between the two species, δ (Shriver et al., 1997). For a given locus, δ is calculated as half the sum of the absolute value of allele frequency differences between species. F-statistics were also estimated for both species in each mixed population and in the whole set of individuals following the weighted analysis of variance method of Weir and Cockerham (1984). All analyses were carried out only with putative purebred individuals, selected after a preliminary screening for potential hybrids, as explained below.
Nuclear admixture analysis for hybrid identification
To identify hybrid individuals and estimate population-level hybridization, we carried out admixture analyses using two different Bayesian clustering approaches, as implemented in the programs STRUCTURE version 2 (Pritchard et al., 2000) and NEWHYBRIDS version 1.1 beta (Anderson and Thompson, 2002). Both methods were used to assign probabilistically individual multilocus genotypes to categories (clusters) by jointly inferring the parameters corresponding to each cluster and the cluster membership of each individual (that is, without a priori knowledge of the allele frequencies in the separate clusters). A Markov chain Monte Carlo simulation procedure provides the estimates from the posterior distribution reflecting the membership of each individual. In the STRUCTURE model, the posterior probability (q) describes the proportion of an individual genotype originating from each of K categories. In our case, setting K=2 corresponds to the assumption of two species contributing to the gene pool of the sample. Instead, NewHybrids model assumes that the sample is drawn from a mixture of pure individuals and hybrids (Anderson and Thompson, 2002). Under this model, q describes the probability that an individual belongs to each of different genotype frequency classes (in our case: parental purebreds, F1 hybrid and the two first backcrosses categories). Analyses were carried out for all individuals jointly and for each of the mixed populations separately. In all cases, no prior species information was used. With STRUCTURE, calculations were carried out under the admixture model assuming independent allele frequencies, given the high interspecific differentiation (see results). A burn-in of 50 000 steps followed by 100 000 iterations was used with each program, after verifying that results do not vary significantly across multiples runs and with longer cycles of burn-in/iterations.
When using these assignment approaches, an important decision is the choice of the optimal threshold value (Tq) for the q associated with the classification of each individual into purebred or hybrid (Vähä and Primmer, 2006). We used threshold values of 0.90 (Pritchard et al., 2000; Vähä and Primmer, 2006) and 0.75. With STRUCTURE, a value of q higher or equal to the threshold indicates a purebred genotype and a value of q lower than the threshold indicates an introgressed genotype. With NEWHYBRIDS, the threshold values can be used in three ways. In the most restrictive way (criterion 1) the threshold value is applied to each category (pure species, F1 hybrids, backcrosses) separately, by assigning only the individuals with q⩾Tq and leaving the others unassigned (Oliveira et al., 2007). Alternatively, q values for all hybrid categories (F1 hybrids, backcrosses) can be combined (Vähä and Primmer, 2006) to distinguish hybrids regardless of their category (criterion 2). A third option (criterion 3), the most relaxed, is to apply the threshold only to the purebred category, assuming that individuals with q⩾Tq are purebreds and that all others are hybrids (this is the only case where no individual remains unassigned).
Performance of the two admixture analyses
We used simulated data to assess which method provides the most reliable results with our experimental system (as suggested by Vähä and Primmer, 2006). Specifically, we tried to identify the Tq for the q to distinguish hybrids from purebreds. We also tested which of the criteria suggested for hybrid identification with NEWHYBRIDS performs best, and we evaluated the effect of different sample sizes.
Allele frequencies for parental species were estimated from the whole sample after taking out potentially introgressed individuals identified in preliminary runs of both STRUCTURE and NEWHYBRIDS (these are the individuals with q<0.90 for pure species categories, which corresponds to the criterion 3 for NEWHYBRIDS). Ten thousand purebred genotypes were then generated with HYBRIDLAB 1.0 (Nielsen et al., 2006) for each species using these allele frequencies. In addition, three hybrid sets of 10 000 genotypes each were generated by randomly drawing alleles (random mating assumed) from each of the simulated purebred genotypes for the F1 set and from simulated purebred genotypes and simulated F1 genotypes for each backcross set. Genotypes were sampled without replacement from the five simulated sets with POPTOOLS 2.6 (Hood, 2005) to create samples of 150 and 1500 individuals with two different proportions of hybrids (HP): 0 and 2%. The first figure corresponds to the complete lack of hybrids in the sample, whereas HP=2% corresponds to 3 hybrids (one F1 and two F1 backcrosses to each parent species) and 30 hybrids (10 F1 and 10 of each of the two backcrosses), respectively, for N=150 and 1500. Sample sizes and HPs have been chosen to represent the actual population samples. For each HP, 100 replicate data sets were generated for N=150 and 10 replicates for N=1500. Each simulated data set was analyzed with STRUCTURE and NEWHYBRIDS with the same setting conditions, threshold values and criteria described before.
The following measures were used to evaluate the performance of the methods:
-
1)
the hybrid proportion: number of individuals classified as hybrids over the total number of individuals in the sample;
-
2)
the power to detect the true hybrid/purebred status of individuals (‘efficiency’ sensu Vähä and Primmer, 2006): number of correctly identified individuals for a category over the actual number of individuals of that category in the sample;
-
3)
the accuracy (sensu Yang et al., 2005 and Vähä and Primmer, 2006): number of correctly identified individuals for a category over the total number of individuals assigned to that category; and
-
4)
the type I error: number of individuals wrongly identified as hybrids over the total number of actual purebreds in the sample.
Finally, we compared the power and accuracy of the clustering algorithms as a function of the number of molecular markers examined. We considered two sets of three combinations of molecular markers (2, 4 and 6 loci), with N=1500 simulated genotypes. The first set was composed of three combinations of loci with decreasing value of δ, starting with the two most discriminating, MSQ13 and QpZAG9 (Table 2). The second set was composed of three combinations of loci with increasing value of δ, starting with the two with the least discriminatory power (that is, QpZAG36 and QrZAG20, Table 2). This provided approximate upper and lower bounds of the power and accuracy for different combinations of loci.
Results
Information content of microsatellites and species differentiation
Although some loci showed significant homozygous excess (18 tests out of 144 with P-value <0.05) and LD (10 tests out of 504 with P-value <0.05), no consistent pattern was found across all populations and species (data not shown). All marker loci have high discriminatory power over the whole sample, with allele frequency differential ranging from δ=0.62 to δ=1 (Table 2). After removing putative hybrids to calculate δ, MSQ13 appears to be fully diagnostic. High and significant genetic differentiation between the two species was found over the whole sample as well as in each region (range wide θ=0.41, P-value=0.001; minimum θ=0.40, Minorca; maximum θ=0.44, Spanish Catalonia). For comparison, intraspecific differentiation is 10 times lower (Q. suber θ=0.05; Q. ilex θ=0.06).
Hybrid detection and performance of the admixture analysis
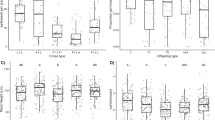
Results of simulations performed with all eight loci for each sample size scenario (that is, 150 and 1500) were quite similar across methods (that is, STRUCTURE versus NEWHYBRIDS) and thresholds (that is, 0.90 versus 0.75). Nevertheless, higher power and accuracy and lower error rates were reached with the larger sample size (data not shown). Thus, results presented here refer exclusively to analyses of real data performed with all 1487 individuals jointly and of simulated data with the 1500 samples. With NEWHYBRIDS, criterion 2 (hybrid probability: sum of probabilities for F1 and backcrosses) was selected because it showed the best performance using simulated data (results not shown).
In the absence of hybrids, both Bayesian approaches used to infer the individual admixture proportions perform well, although STRUCTURE provides a small proportion of false hybrids with the 0.90 threshold (Table 3). On the contrary, when the simulated sample contains hybrid individuals, the best HP estimate is found with STRUCTURE and the 0.90 threshold; a slight underestimate is obtained with NEWHYBRIDS for both threshold values, and a strong underestimate with STRUCTURE and the 0.75 threshold (Table 3). Likewise, the power to correctly classify purebreds is higher than 99% in all cases, but the highest proportion of correctly identified hybrids is achieved when STRUCTURE is used with the 0.90 threshold (92%), followed by NEWHYBRIDS with thresholds of 0.75 and of 0.90. Compared to STRUCTURE, detection ability is lower with NEWHYBRIDS, because some individuals remain unassigned (for the empirical data set, nine genotypes are unassigned with Tq=0.90 and four with Tq=0.75), but accuracy in identifying hybrids is improved (>99% for a power >86% using both thresholds; Table 3). Thus, STRUCTURE provides power whereas NEWHYBRIDS provides accuracy.
As expected, both the power and accuracy increase with the number of loci (Figure 4). This increase is higher for the identification of hybrids than for the identification of purebreds (results not shown). The simulations show that the four most discriminant loci suffice to reach high power in identifying hybrids with STRUCTURE and high accuracy with NEWHYBRIDS, values comparables with those obtained using eight loci (Figure 4). However, a higher number of individuals remain unassigned with NEWHYBRIDS when only four loci are used (112, including 39 hybrids, compared to 71, including 22 hybrids, with all eight markers).
Power (a) and accuracy (b) in detecting hybrid individuals for 10 simulated samples of N=1500, analyzed with STRUCTURE (black line) and NEWHYBRIDS (gray line), as a function of the number of microsatellite markers (Tq=0.90). Loci have been combined according to their decreasing (solid line) and increasing (dashed line) value of frequency differential δ (see text).
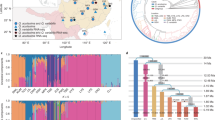
When applied to our experimental data set, both methods separated the 1487 individuals examined into two well-defined groups congruent with the observed suber and ilex phenotypes. Both methods also identified a very low total number of putative hybrids, most of them in mixed populations. Some differences were found between both methods, in agreement with the results of the simulations. With STRUCTURE, 17 potential hybrids were detected with a threshold Tq=0.90 (that is, an HP=1.1%), but this estimate drops to 4 with Tq=0.75 (HP=0.03%; Figure 2a). All remaining individuals have a very high probability to belong to the purebred species (Q. suber: range 0.903–0.998; Q. ilex: range 0.925–0.998). With NEWHYBRIDS, five individuals were identified as hybrids with Tq=0.90 (HP=0.20%) and seven with Tq=0.75 (HP=0.34%; Figure 2b). Again, putative purebreds present high q-values (Q. suber: range 0.901–1.000; Q. ilex: range 0.960–1.000). Surprisingly, three individuals morphologically identified as Q. suber, from Minorca (one) and from Sicily (two), have been classified by molecular analysis as pure Q. ilex.
Posterior probability (q) for all individuals identified as putative hybrids by at least one of the method–threshold (Tq) combination. Each individual is represented by a vertical bar partitioned into segments, the length of which describes (a) the estimated membership proportions for each parental species (Q. suber and Q. ilex) by STRUCTURE and (b) the estimated probability of belonging to the parental species and the three hybrid classes (F1, first backcross with each of the parental species) by NEWHYBRIDS. Individuals are identified by a population code (see Table 1) and ID number. †Classified as hybrid with Tq=0.90, ‡Classified as hybrid with Tq=0.75.
Genetic composition of hybrid/introgressed individuals
STRUCTURE detected a total of 17 individuals with q between 0.10 and 0.90 (Figure 2a); 8 of them had been classified in the field as Q. suber and 9 as Q. ilex. However, NEWHYBRIDS assigns six of them to purebred categories with q>0.95 (two Q. suber and four Q. ilex, matching field identification) (Figure 2b). In view of the high accuracy provided by NEWHYBRIDS and the false positive rate associated with STRUCTURE (when HP=2%, type I error=0.001; Table 3), the hybrid nature of those six individuals is uncertain. In contrast, the hybrid nature of the remaining 11 trees appears more consistent and for 7 of them very well supported. Only two individuals, one from the SCa population (suber SCa70) and one from the MN population (ilex MN36), showed intermediate proportions compatible with an F1 genotype with both methods (Figure 2), although a backcross status cannot be excluded. In fact, simulations showed that all F1 hybrids are always correctly classified as hybrids (that is, none was assigned to any pure species) whichever method and threshold is used (data not shown), but some of them present a pattern of admixture indistinguishable from that of backcrosses (Figure 3). The remaining nine individuals (SCa95, MN32, MN39, MN45, TU2 suber morphotype, CLM48, SCa36, SCa84 and SI2 ilex morphotype) probably result from one or more generations of backcross. Among them, SCa95, SCa36 and SI2 have the phenotype of one species despite having a large assignment probability to the other species (Figure 2).
Posterior probability (q) for simulated hybrid individuals analyzed in 10 repetitions of simulated samples with N=1500 and 2% hybrid proportions each with (a) STRUCTURE and (b) NEWHYBRIDS. Number of hybrids: 100 backcrosses to Q. suber (Bxs), 100 first-generation hybrids (F1) and 100 backcrosses to Q. ilex (Bxi).
Discussion
Evidence and rate of hybridization between cork and holm oaks
The microsatellite loci chosen for this work were highly differentiated between species (θ=0.41) and had good diagnostic power (δ=0.62–1.0). In fact, both Bayesian clustering approaches used (implemented in STRUCTURE and NEWHYBRIDS) assigned nearly all individuals with high probability to each of two genetically defined groups, resulting in an almost perfect match with the observed morphotypes. Very few hybrid genotypes have been detected (0.027–1.14% of the total sample, using the most and least restrictive conditions, respectively; Figure 2). Using simulated data, we have quantified the resolution level achieved and the uncertainty attached to the experimental system and threshold values for two posterior probabilities (0.90 and 0.75). These results indicate that, although the correct identity of hybrid individuals cannot be guaranteed in all cases, it is possible to get a good estimate of the actual proportion of hybrids in our sample (see estimated and simulated HP in Table 3). Simulations also showed that we could achieve similar results with half of the loci (Figure 4) by selecting those with the highest discriminatory power, in agreement with Boecklen and Howard (1997). This may suggest a rapid method to distinguish hybrids from pure holm oaks and cork oaks. However, this conclusion has to be taken with caution, because simulations relies on simplifying assumptions (for example, symmetrical introgression, limited type of backcross categories) likely not fulfilled by natural populations. Hence, we consider a worth effort increasing the number of molecular markers to improve the level of resolution, even if highly diagnostic markers are available.
The low frequency (<2%) of contemporary gene exchange detected between Q. suber and Q. ilex is consistent with the available knowledge on nuclear variability for the species. A low number of hybrids has been reported in previous surveys of isozyme diversity (Elena-Rosselló et al., 1992; Toumi and Lumaret, 1998; Lumaret et al., 2002; Staudt et al., 2004). Nevertheless, the extensive sharing of chloroplast DNA haplotypes between Q. suber and Q. ilex in some regions has led some authors to hypothesize widespread introgressive hybridization events in the past (Belahbib et al., 2001; Lumaret et al., 2002; Jiménez et al., 2004; López de Heredia et al., 2005). Such findings are not incompatible, given that even a low fraction of hybrids can have considerable evolutionary impact because of the cumulative effect of introgression through time (Ellstrand et al., 1996; Mallet, 2005) and the possibility for introgressed genes to become amplified by demographic growth (Currat et al., 2008). In this respect, López de Heredia et al. (2007a) suggested that the acidophilous Q. suber was able to colonize the calcareous area of eastern Iberia (where chloroplast introgression has been reported), thanks to the hybridization with Q. ilex, which is largely indifferent to soil nature. It is noteworthy that we found a higher proportion of early generation hybrids in Catalonia and Minorca, located within the area of chloroplast introgression and where soils are mostly formed on more or less decarbonated calcarenites and dolomites, unfavorable to cork oak. This would be consistent with the ‘environmental emasculation’ hypothesis proposed by Williams et al. (2001), according to which environmental stress, at the margins of the suitable habitat of a species, can lead to a decrease in the competitive ability of its pollen, thus favoring hybridization. Alternatively, the process could be driven (exclusively or complementarily) by demographic factors, due to demographic imbalance during colonization, as suggested by Currat et al. (2008). In Minorca, for instance, Q. suber population size is limited to the 67 individuals we sampled.
As shown by results from controlled crosses (Boavida et al., 2001), Q. suber likely acts as the pollen donor in interspecific mating events with Q. ilex. This finding is supported by the discovery of widespread introgression of ilex-type cpDNA in Q. suber populations (Belahbib et al., 2001; Jiménez et al., 2004; Lumaret et al., 2005), whereas the opposite situation (that is, Q. ilex trees showing suber chlorotypes) is considerably less frequent. However, no evidence of unidirectional gene flow has been found in this study, because we detected a similar number of backcrosses to each species (Figure 2). Artificial crosses involving F1s and the parental species would help determine the direction of introgression and the nature of barriers to random mating. For instance, Olrik and Kjaer (2007) showed that Q. robur–Q. petraea unidirectional hybridization does not imply necessarily asymmetric backcrossing to the parental species, after performing controlled crosses with an F1 tree of known pedigree.
Hybrid identity
Among the 17 putative hybrids, we could distinguish at least 2 putative F1s (suber SCa70 and ilex MN36) and 5 backcrosses (suber MN32, MN45, ilex SCa36, SCa84 and SI2) with very high probability (Figure 2). The reliability of their hybrid identity is supported by the coincident assignation with two different methods and by the high accuracy and low error observed in data-based simulations with two different thresholds values (Table 3). Although we can be reasonably sure that these seven individuals are not purebreds, and that any actual F1 present in the sample would not have been classified as purebred, we cannot exclude that the two putative F1s are backcrosses. Similarly, some uncertainty is involved in the identity of the 10 remaining individuals (Figure 2). Our simulations indicated that the identification of backcrosses is more problematic than that of F1 hybrids, because they can be confused with pure individuals and vice versa, as already shown with other hybrid systems (Barilani et al., 2007). The extent of incorrect classification can be minimized by choosing an appropriate threshold to improve accuracy (and reduce error), whereas accepting the associated trade off of loss of power (Vähä and Primmer, 2006). We found through simulations that Tq=0.90 is an appropriate threshold for this purpose (Table 3). With STRUCTURE, we obtain a good estimate of the proportion of hybrids in the sample (>90%) with a very low associated error. With NEWHYBRIDS, reliable results on true hybrid identity (accuracy=1.000) are obtained with virtually no error. Hence, the joint use of these Bayesian approaches is suggested to improve the resolution in hybrid identification, especially for studies relying on the prior identification of hybrid plants (for example, controlled crosses or detailed phenotypic observations of hybrids compared to parental species). We note that the present study was based on a very limited number of loci. In admixture zones that are already many generations old, both power and accuracy of hybrid detection will increase greatly if a much larger, genome-wide panel of diagnostic marker loci is used, especially if linkage between loci is accounted for during the estimation of hybrid ancestry (Falush et al., 2003).
No general rule about morphological features of hybrid individuals between Q. suber and Q. ilex can be deduced from previous studies. Putative hybrids with parental morphology (Toumi and Lumaret, 1998; Belahbib et al., 2001; Staudt et al., 2004), intermediate morphology (Toumi and Lumaret, 1998; Lumaret et al., 2002; Bellarosa et al., 2005) or leaf morphology skewed toward Q. ilex (Staudt et al., 2004) have been reported. Bark cannot be used as a discriminating feature because F1s are considered to lack cork and, thus, they could be confused with pure Q. ilex (Lumaret et al., 2002; Bellarosa et al., 2005). The existence of morphologically cryptic hybrids seems to be the only certainty. In any case, results from the studies cited above are hardly comparable among them and with the present one, due to the different sample designs and type of genome variability observed. Moreover, in all of these studies the identification of genetic diagnostic elements is dependent on the morphological determination of pure species. In contrast, the Bayesian approach used here allows us to define the genetic boundaries of pure species independently from any feature other than genetic data, thus allowing more accurate estimates of species status (Duminil et al., 2006) and gene exchange. We found a similar proportion of each parental morphotype among the putative hybrid individuals (Figure 2) and very good correspondence between morphotype and molecular-based assignation for the putative purebreds. However, discrepancy was detected for a few individuals, because three trees identified in the field as Q. suber were assigned to pure ilex using microsatellites and three putative hybrid individuals (suber SCa95 and ilex SCa36, SI2) were morphologically similar to one species but assigned with greater probability to the other species (Figure 2). In contrast, there was no ambiguous assignment with simulated data; that is, backcrosses to Q. suber (Bxs) were never assigned to Q. ilex with q>0.50 by STRUCTURE (Figure 3a), and they were never assigned to Q. ilex nor to backcrosses with Q. ilex (Bxi) with q>0.10 by NewHybrids (Figure 3b). The same was found with backcrosses to Q. ilex (Figures 3a and b). Thus, we consider that the discrepancy mentioned above is not due to the lack of resolution of the methods but reveals instead either the lack of correspondence between the phenotype and nuclear genotype (expected after several backcrosses, that is, ‘advanced’ introgressed individuals) or mislabeling of samples during their collection and processing (although this is unlikely for backcrosses, given their extremely low frequency in the sample).
Conclusions
The strength of our approach relied on the combination of two complementary Bayesian methods and on their validation by systematic simulations precisely adjusted to the empirical data investigated. The whole procedure is recommended to gain precision and accuracy in the identification of F1 hybrids and backcrosses for every real-case study, regardless of the level of hybridization. We expect that future studies of hybrids in natural populations will achieve even greater accuracy and power by increasing genomic coverage and accounting for linkage between loci. In the case of Q ilex and Q. suber, the identification of hybrid types has been addressed for the first time in this study. Our results suggest a very low rate of bidirectional gene flow between Q. ilex and Q. suber. Further studies are required to understand the geographic distribution and possible adaptive function of hybridization between these two species through time and space. Powerful and accurate detection of adult hybrid and introgressed individuals will be particularly valuable to address the adaptive differences among hybrid classes and the reproductive behavior of hybrid individuals.
References
Amaral Franco J (1990). Quercus L. In: Castroviejo S (ed). Flora Ibérica. Plantas vasculares de la Península Ibérica e Islas Baleares. Real Jardín Botánico, CSIC: Madrid. vol. II.
Anderson EC, Thompson EA (2002). A model-based method for identifying species hybrids using multilocus genetic data. Genetics 160: 1217–1229.
Barilani M, Sfougaris A, Giannakopoulos A, Mucci N, Tabarroni C, Randi E (2007). Detecting introgressive hybridisation in rock partridge populations (Alectoris graeca) in Greece through Bayesian admixture analyses of multilocus genotypes. Conserv Genet 8: 343–354.
Belahbib N, Pemonge MH, Ouassou A, Sbay H, Kremer A, Petit RJ (2001). Frequent cytoplasmic exchanges between oak species that are not closely related: Quercus suber and Q. ilex in Morocco. Mol Ecol 10: 2003–2012.
Bellarosa R, Simeone MC, Papini A, Schirone B (2005). Utility of ITS sequence data for phylogenetic reconstruction of Italian Quercus spp. Mol Phylogenet Evol 34: 355–370.
Boavida LC, Silva JP, Feijó JA (2001). Sexual reproduction in the cork oak (Quercus suber L.). II. Crossing intra- and interspecific barriers. Sex Plant Reprod 14: 143–152.
Boecklen WJ, Howard DJ (1997). Genetic analysis of hybrid zones: numbers of markers and power of resolution. Ecology 78: 2611–2616.
Burgarella C, Navascués M, Soto A, Lora González A, Fici S (2007). Narrow genetic base in forest restoration with holm oak (Quercus ilex L.) in Sicily. Ann Forest Sci 64: 757–763.
Burgess KS, Morgan M, DeVerno LL, Husband CB (2005). Asymmetrical introgression between two Morus species (M. alba, M. rubra) that differ in abundance. Mol Ecol 14: 3471–3483.
Currat M, Ruedi M, Petit R, Excoffier L (2008). The hidden side of invasions: massive introgression by local genes. Evolution 62: 1908–1920.
Curtu AL, Gailing O, Finkedley R (2007). Evidence for hybridization and introgression within a species-rich oak (Quercus spp.) community. BMC Evol Biol 7: 218.
Dodd RS, Afzal-Rafii Z (2004). Selection and dispersal in a multispecies oak hybrid zone. Evolution 58: 261–269.
Dow B, Ashley M, Howe H (1995). Characterization of highly variable (GA/CT)n microsatellites in the bur oak, Quercus macrocarpa. Theor Appl Genet 91: 137–141.
Duminil J, Caron H, Scotti I, Cazal S-O, Petit RJ (2006). Blind population genetics survey of tropical rainforest trees. Mol Ecol 15: 3505–3513.
Dumolin-Lapègue S, Kremer A, Petit RJ (1999). Are chloroplast and mitochondrial DNA variation species independent in oaks? Evolution 53: 1406–1413.
Dumolin S, Demesure B, Petit R (1995). Inheritance of chloroplast and mitochondrial genomes in pedunculate oak investigated with an efficient PCR method. Theor Appl Genet 91: 1253–1256.
Elena-Rosselló JA, Lumaret R, Cabrera E, Michaud H (1992). Evidence for hybridization between sympatric holm-oak and cork oak in Spain based on diagnostic enzyme markers. Vegetatio 99–100: 115–118.
Ellstrand NC, Whitkus R, Rieseberg LH (1996). Distribution of spontaneous plant hybrids. Proc Natl Acad Sci 93: 5090–5093.
Falush D, Stephens M, Pritchard JK (2003). Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164: 1567–1587.
Fernández-Manjarrés JF, Gerard PR, Dufour J, Raquin C, Frascaria-Lacoste N (2006). Differential patterns of morphological and molecular hybridization between Fraxinus excelsior L. and Fraxinus angustifolia Vahl (Oleaceae) in eastern and western France. Mol Ecol 15: 3245–3257.
González-Rodríguez A, Arias DM, Valencia S, Oyama K (2004). Morphological and RAPD analysis of hybridization between Quercus affinis and Q. laurina (Fagaceae), two Mexican red oaks. Am J Bot 91: 401–409.
Goudet J (2001). FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). Available at http://www2.unil.ch/popgen/softwares/fstat.htm.
Heuertz M, Carnevale S, Fineschi S, Sebastiani F, Hausman JF, Paule L et al. (2006). Chloroplast DNA phylogeography of European ashes, Fraxinus sp. (Oleaceae): roles of hybridization and life history traits. Mol Ecol 15: 2131–2140.
Hood GM (2005). PopTools version 2.6.6. Available at http://www.cse.csiro.au/poptools.
Howard DJ, Preszler RW, Williams JH, Fenchel S, Boecklen WJ (1997). How discrete are oak species? Insights form a hybrid zone between Quercus grisea and Quercus gambelii. Evolution 51: 747–755.
Jensen RJ, Hokanson SC, Isebrands JG, Hancock JF (1993). Morphometric variation in oaks of the Apostle Islands in Wisconsin: evidence of hybridization between Quercus rubra and Q. ellipsoidalis. Am J Bot 80: 1358–1366.
Jiménez MP, Lopez de Heredia U, Collada C, Lorenzo Z, Gil L (2004). High variability of chloroplast DNA in three Mediterranean evergreen oaks indicates complex evolutionary history. Heredity 93: 510–515.
Kampfer S, Lexer C, Glössl J, Steinkellner H (1998). Characterization of (GA)n microsatellite loci from Quercus robur. Hereditas 129: 183–186.
Kothera L, Ward SM, Carney SE (2007). Assessing the threat from hybridization to the rare endemic Physaria bellii Mulligan (Brassicaceae). Biol Conserv 140: 110–118.
Lexer C, Fay MF, Joseph JA, Nica MS, Heinze B (2005). Barrier to gene flow between two ecologically divergent Populus species, P. alba (white poplar) and P. tremula (European aspen): the role of ecology and life history in gene introgression. Mol Ecol 14: 1045–1057.
Lexer C, Kremer A, Petit RJ (2006). Shared alleles in sympatric oaks: recurrent gene flow is a more parsimonious explanation than ancestral polymorphism. Mol Ecol 15: 2007–2012.
López de Heredia U, Carrión JS, Jiménez P, Collada C, Gil L (2007a). Molecular and palaeobotanical evidence for multiple glacial refugia for evergreen oaks on the Iberian Peninsula. J Biogeogr 34: 1505–1517.
López de Heredia U, Jiménez P, Collada C, Simeone MC, Bellarosa R, Schirone B et al. (2007b). Multi-marker phylogeny of three evergreen oaks reveals vicariant patterns in the Western Mediterranean. Taxon 56: 1199–1209.
López de Heredia U, Jiménez P, Díaz-Fernández PM, Gil L (2005). The Balearic Islands: a reservoir of cpDNA genetic variation for evergreen oaks. J Biogeogr 32: 939–949.
Lumaret R, Mir H, Michaud H, Raynal V (2002). Phylogeographical variation of chloroplast DNA in holm oak (Quercus ilex L.). Mol Ecol 11: 2327–2336.
Lumaret R, Tryphon-Dionnet M, Michaud H, Sanuy A, Ipotesi E, Born C et al. (2005). Phylogeographical variation of chloroplast DNA in cork oak (Quercus suber). Ann Bot 96: 853–861.
Mallet J (2005). Hybridization as an invasion of the genome. Trends Ecol Evol 20: 229–237.
Manos PS, Doyle JJ, Nixon KC (1999). Phylogeny, biogeography, and processes of molecular differentiation in Quercus subgenus Quercus (Fagaceae). Mol Phylogenet Evol 12: 333–349.
Manos PS, Zhou Z, Cannon CH (2001). Systematics of Fagaceae: phylogenetic tests of reproductive trait evolution. Int J Plant Sci 162: 1361–1379.
Martín Vicente Á, Fernández Alés R (2006). Long term persistence of dehesas. Evidences from history. Agroforest Syst 67: 19–28.
Muir G, Schlötterer C (2005). Evidence for shared ancestral polymorphism rather than recurrent gene flow at microsatellite loci differentiating two hybridizing oaks (Quercus spp.). Mol Ecol 14: 549–561.
Nason JD (1992). Patterns of hybridization and introgression in populations of oaks, manzanitas, and irises. Am J Bot 79: 101–111.
Nielsen EE, Bach LA, Kotlick P (2006). HYBRIDLAB (version 1.0): a program for generating simulated hybrids from population samples. Mol Ecol Notes 6: 971–973.
Oliveira P, Custódio AC, Branco C, Reforço I, Rodrigues F, Varela MC et al. (2003). Hybrids between cork oak and holm oak: isoenzyme analysis. Forest Genet 10: 283–297.
Oliveira R, Godinho R, Randi E, Ferrand N, Alves P (2007). Molecular analysis of hybridisation between wild and domestic cats (Felis silvestris) in Portugal: implications for conservation. Conserv Genet 9: 1–11.
Olrik DC, Kjaer ED (2007). The reproductive success of a Quercus petraea x Q. robur F1-hybrid in back-crossing situations. Ann Forest Sci 64: 37–45.
Palmé AE, Su Q, Palsson S, Lascoux M (2004). Extensive sharing of chloroplast haplotypes among European birches indicates hybridization among Betula pendula, B. pubescens and B. nana. Mol Ecol 13: 167–178.
Petit RJ, Bialozyt R, Garnier-Gere P, Hampe A (2004). Ecology and genetics of tree invasions: from recent introductions to Quaternary migrations. Forest Ecol Manag 197: 117–137.
Petit RJ, Pineau E, Demesure B, Bacilieri R, Ducousso A, Kremer A (1997). Chloroplast DNA footprints of postglacial recolonization by oaks. Proc Natl Acad Sci 94: 9996–10001.
Plieninger T, Pulido FJ, Konold W (2003). Effects of land-use history on size structure of holm oak stands in Spanish dehesas: implications for conservation and restoration. Environ Conserv 30: 61–70.
Potts B, Reid JB (1988). Hybridization as a dispersal mechanism. Evolution 42: 1245–1255.
Pritchard JK, Stephens M, Donnelly P (2000). Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
Rieseberg LH, Carney SE (1998). Plant hybridization. New Phytol 140: 599–624.
Rieseberg LH, Raymond O, Rosenthal DM, Lai Z, Livingstone K, Nakazato T et al. (2003). Major ecological transitions in wild sunflowers facilitated by hybridization. Science 301: 1211–1216.
Rushton BS (1993). Natural hybridization within the genus Quercus L. Ann Sci Forest 50 (Suppl 1): 73–90.
Seehausen O (2004). Hybridization and adaptive radiation. Trends Ecol Evol 19: 198–207.
Shriver MD, Smith MW, Jin L, Marcini A, Akey JM, Deka R et al. (1997). Ethnic-affiliation estimation by use of population-specific DNA markers. Am J Hum Gen 60: 957–964.
Soto A, Lorenzo Z, Gil L (2003). Nuclear microsatellites markers for the identification of Quercus ilex L. and Quercus suber L. hybrids. Silvae Genet 52: 63–66.
Soto A, Lorenzo Z, Gil L (2007). Differences in fine-scale genetic structure and dispersal in Quercus ilex L. and Q. suber L.: consequences for regeneration of Mediterranean open woods. Heredity 99: 601–607.
Staudt M, Mir C, Joffre R, Rambal S, Bonin A, Landais D et al. (2004). Isoprenoid emissions of Quercus spp. (Q.suber and Q.ilex) in mixed stands contrasting in interspecific genetic introgression. New Phytol 163: 573–584.
Steinkellner H, Fluch S, Turetschek E, Lexer C, Streiff R, Kremer A et al. (1997). Identification and characterization of (GA/CT)n—microsatellite loci from Quercus petraea. Plant Mol Biol 3: 1093–1096.
Toumi L, Lumaret R (1998). Allozyme variation in cork oak (Quercus suber L.): the role of phylogeography and genetic introgression by other Mediterranean oak species and human activities. Theor Appl Genet 97: 647–656.
Toumi L, Lumaret R (2001). Allozyme characterization of four Mediterranean evergreen oak species. Biochem Syst Ecol 29: 799–817.
Tovar-Sanchez E, Oyama K (2004). Natural hybridization and hybrid zones between Quercus crassifolia and Quercus crassipes (Fagaceae) in Mexico: morphological and molecular evidence. Am J Bot 91: 1352–1363.
Vähä J-P, Primmer CR (2006). Efficiency of model-based Bayesian methods for detecting hybrid individuals under different hybridization scenarios and with different numbers of loci. Mol Ecol 15: 63–72.
Valbuena-Carabaña M, Gonzalez-Martinez SC, Hardy OJ, Gil L (2007). Fine-scale spatial genetic structure in mixed oak stands with different levels of hybridization. Mol Ecol 16: 1207–1219.
Varela MC, Valdiviesso T (1996). Phenological phases of Quercus suber L. flowering. Forest Genet 3: 93–102.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Whittemore AT, Schaal BA (1991). Interspecific gene flow in sympatric oaks. Proc Natl Acad Sci 88: 2540–2544.
Williams JH, Boecklen WJ, Howard DJ (2001). Reproductive processes in two oak (Quercus) contact zones with different levels of hybridization. Heredity 87: 680–690.
Yang BZ, Zhao H, Kranzler HR, Gelernter J (2005). Practical population group assignment with selected informative markers: characteristics and properties of Bayesian clustering via STRUCTURE. J Epidemiol 28: 302–312.
Acknowledgements
This work was partially funded by the EU Project QLRT-2001-01594 (CREOAK) and by the Spanish Ministry of Environment (DGB) through the covenant ‘Evaluación y conservación de los recursos genéticos de los Quercus esclerófilos mediterráneos en España’. We thank Stella Mérola, Unai Lopez de Heredia and Carmen García Barriga for their help with field and laboratory work, and P Montoya for English revision. We also thank Miguel Navascués for constructive discussions during the development of the work and on earlier versions of the paper. Suggestions by anonymous reviewers and editor were much appreciated.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Burgarella, C., Lorenzo, Z., Jabbour-Zahab, R. et al. Detection of hybrids in nature: application to oaks (Quercus suber and Q. ilex). Heredity 102, 442–452 (2009). https://doi.org/10.1038/hdy.2009.8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2009.8
Keywords
This article is cited by
-
Haplotype diversity patterns in Quercus suber (Fagaceae) inferred from cpDNA sequence data
Plant Systematics and Evolution (2023)
-
Characterization of pollen tube development in distant hybridization of Chinese cork oak (Quercus variabilis L.)
Planta (2023)
-
Counteracting forces of introgressive hybridization and interspecific competition shape the morphological traits of cryptic Iberian Eptesicus bats
Scientific Reports (2022)
-
Development of Quercus acutissima (Fagaceae) pollen tubes inside pistils during the sexual reproduction process
Planta (2022)
-
Evidence for extensive hybridisation and past introgression events in feather grasses using genome-wide SNP genotyping
BMC Plant Biology (2021)