Abstract
Horizontal transfer (HT), defined as the transfer of genetic material between species, is considered to be an essential step in the ‘life cycle’ of transposable elements. We present a broad overview of suspected cases of HT of transposable elements in Drosophila. Hundred-one putative events of HT have been proposed in Drosophila for 21 different elements (5.0% refer to non-long terminal repeat (LTR) retrotransposons, 42.6% to LTR retrotransposons and 52.4% to DNA transposons). We discuss the methods used to infer HT, their limits and the putative vectors of transposable elements. We outline all the alternative hypotheses and ask how we can be almost certain that phylogenetic inconsistencies are due to HT.
Similar content being viewed by others
Introduction
One of the outstanding traits of transposable elements (TEs) is their ability to cross species boundaries and invade new genomes. This process, named horizontal transfer (HT), and defined as transfer of genetic material between species, has been proposed as an essential step in the DNA transposons ‘life cycle’ and, therefore, in genome evolution. A successful HT requires a TE transfer into the germ line of the recipient species, followed by a high transposition rate leading to a rapid propagation into the genome, as well as into the population by vertical transmission (Le Rouzic and Capy, 2005). The transpositional activity is then regulated or suppressed by various mechanisms, and the frequency of functional copies decreases as they are subjected to random mutations, excision, purifying selection and stochastic losses. HT of a functional element to a naïve genome or reintroduction into the primary host genome can therefore prevent its extinction. This fact was illustrated by Hartl et al. (1997) for the mariner element.
A growing body of evidence has shown that this phenomenon may not be very rare in eukaryotes. However, although several results seem to be robust enough to be ascribed to HT, in other cases alternative hypotheses can be frequently proposed (Capy et al., 1994; Cummings, 1994). This work aims to present an overview of the cases of HT already suspected in Drosophila and the arguments used to infer their robustness. The Drosophilidae, including about 3700 species with well-documented phylogenetic relationships, is certainly the best family of eukaryotes to define the features of HTs (detection of HT, conditions and frequency for successful HT) in connection with environmental and populational dynamics. First, we present the facts generally considered to infer HT and the phenomena leading to a misinterpretation. Second, we present the mechanisms and vectors possibly involved in the HT process. Finally, an evaluation of the main aspects that led the authors to their conclusions is proposed.
How many putative cases of HT have been described in Drosophila?
An increasing amount of experimental data suggests that many TEs have been horizontally transferred among Drosophila species. The first report of such an event in Drosophila came from the recent invasion of Drosophila melanogaster by P and I elements (Brégliano and Kidwell, 1983; Kidwell, 1983). Beginning with the analyses of hybrid dysgenesis between strains of D. melanogaster, Margaret Kidwell made the first step toward proving one of the most convincing cases of HT, the P element transfer from D. willistoni to D. melanogaster. This was later demonstrated molecularly by Daniels et al. (1990b) and by Kidwell and co-workers (Clark et al., 1995, 2002; Clark and Kidwell, 1997). Starting from the very few HT cases proposed during the 1980s, the number increased over the next decades. During the 1990s, at least 21 papers were published on this subject, and 20 in the first 6 years of this decade. To date, at least 101 putative HT events in the Drosophilidae have been proposed from the analysis of 21 elements (Table 1 and Supplementary Tables 1 and 2).
Analysis of the process of HT raises several questions. The first is about what type of TEs are involved in the transfer events. Of the 101 cases listed in Table 1, 5.0% refer to non-long terminal repeat (LTR) retrotransposons, 42.6% to LTR retrotransposons and 52.4% to DNA transposons. These frequencies are partially in agreement with the proposition of Silva et al. (2004) regarding the preponderance of HTs involving DNA transposons over LTR and non-LTR retrotransposons. They pointed out that this gradient of HT may reflect the presence of DNA intermediates during the transposition. Indeed, from the frequency of HTs detected in several types of organisms, Silva et al. concluded that they are more common for DNA transposons for which the DNA intermediate is the only one present during the transposition process. For LTR retrotransposons, there is a DNA intermediate only after the reverse transcription of an RNA copy, and for non-LTR retrotransposons there are no DNA intermediates, since they are directly reverse transcribed into the target site (Luan et al., 1993). The DNA transposons>LTR retrotransposons>non-LTR retrotransposons gradient observed in the Drosophilidae is consistent with the analysis carried out on a larger spectrum of species (Silva et al., 2004). In Drosophila, the number of HTs suspected for class II elements and LTR retrotransposons is fairly similar, and the low frequency of HTs for the non-LTR retrotranposons is confirmed, suggesting that this particular subclass of TE is less prone to HT.
There is some controversy about the possibility of HT involving non-LTR retrotransposons. Malik et al. (1999), who analyzed divergence versus age of various lineages of non-LTR retrotransposons of eukaryotes, did not find any reliable evidence for HTs for these elements during the past 600 Myr. However, Kordis and Gubensek (1998) have shown that Bov-B Line, an ancient component of Squamata genomes (snakes and lizards) since the Mesozoic era, was horizontally transferred to the ancestor of Ruminantia about 40–50 Myr ago.
In Drosophila, four different non-LTR retrotransposons have been pointed as implicated in HT (Table 1 and Supplementary Table 1). Based on its patchy distribution, the I element was suggested as having been horizontally transferred to D. melanogaster in the pioneer studies using genetic analyses and Southern blot (Brégliano and Kidwell, 1983; Kidwell, 1983). Now, with the availability of several genomes, it is possible to see that D. simulans, D. sechellia and D. melanogaster possess almost identical I elements (data not shown), reinforcing the suggested HT. Moreover, the elements jockey, F and Doc seem to be involved in HT (Mizrokhi and Mazo, 1990; Sánchez-Gracia et al., 2005). Taken together, these data suggest that, although less frequently, non-LTR elements can also be horizontally transferred.
How can we infer HTs?
In addition to minimum requirements such as geographic, temporal and ecological overlap between donor and recipient species, three different kinds of evidence are generally used to infer HT of TEs: (i) high sequence similarity between TEs of very distantly related species (Daniels et al., 1990a, 1990b; Robertson and Lampe, 1995; Brunet et al., 1999, ii) incongruence between host and TE phylogenies (Robertson and Lampe, 1995; Terzian et al., 2000; Almeida and Carareto, 2005) and (iii) discontinuous occurrence (patchy distribution) of a TE across a group of species (Daniels et al., 1990a, 1990b; Arca and Savakis, 2000; Loreto et al., 2001).
Even though such phenomena have been observed, to accept HT it is necessary to exclude all possibilities of vertical transmission, since ancestral polymorphism, differential evolutionary rates according to the activity of different sequences, high selective constraints in small parts of the ancestral element and stochastic losses can all occur (Capy et al., 1994; Cummings, 1994). The similarity between TEs, for instance, can be puzzling because many paralogous copies can co-propagate with varying degrees of success within a lineage. Let us suppose that two copies of the same TE family diverge within species lineages and that these copies are vertically transmitted through speciation events. Different copies may survive (or may be sampled in each species) and, because the sequences compared are, in a sense, paralogous, the TE would not match the species tree (Goodman et al., 1979). Hence, a comparison of two paralogous sequences in two related species would inflate the actual divergence time between orthologous elements in those species. Using such estimates as indicators of divergence, other comparisons would indicate a lesser divergence than expected, which would then mistakenly be suggested as HT (Malik et al., 1999). So sequence similarity deserves careful analysis, since genomes of several species have been shown to harbor more than one subfamily of a particular TE, such as mariner (Robertson and MacLeod, 1993; Robertson and Lampe, 1995; Hartl et al., 1997), gypsy (Hochstenbach et al., 1996; Martínez-Sebastián et al., 2002; Heredia et al., 2004) and P element (Clark and Kidwell, 1997), among others.
Finally, a last point seldom if ever discussed is that the author's conclusions can be affected by the number of species used. Well-supported inferences will be reinforced and powerless inferences may be rejected when a larger number of species are studied. The best examples are provided by the analyses of hobo and P elements. This can be illustrated by the comparison of the analyses published by Maruyama and Hartl (1991) and Brunet et al. (1999) about the HT between the species of the Zaprionus genus and those of the melanogaster subgroup.
The strength of the inference
Regarding the strength of the inferences, several questions can be asked. Perhaps the most important question concerns the methodologies used to infer HT. As previously described, HTs are generally inferred from sequence similarity, tree incongruence and/or patchy distribution. Different methodologies have been used to this end (Table 2).
HT can be inferred when TE divergence is significantly lower than that of the host genes, assuming similar or higher levels of selective constraints on the latter. For recent HT events, when the similarity between TE is high, such an approach is sufficient. It has been used by several authors for P, copia and mariner elements (Maruyama and Hartl, 1991; Clark et al., 1994, 1995; Robertson and Lampe, 1995; Clark and Kidwell, 1997; Jordan et al., 1999). The classical example is the P element, which is 3 kb long and differs in just one nucleotide (position 32 in the 5′ UTR) in D. melanogaster and D. willistoni (Daniels et al., 1990b). Such a situation is exceptional and there is no doubt about its interpretation. However, when the similarity between TEs is high compared to the divergence time of their host, the existence of strong selective pressure on TEs, at least to maintain their activity, cannot be excluded. To solve this problem, several authors have used the estimates of synonymous (kS) and non-synonymous (kA) rates of substitution and particularly the kA/kS ratio to infer selective constraints (Robertson and Lampe, 1995; Terzian et al., 2000; Maside et al., 2003; Heredia et al., 2004). Because of degeneracy of the genetic code, a proportion of nucleotide substitutions in protein-coding sequences are expected to be silent, leading to no amino-acid substitution. King and Jukes (1969) predicted that these synonymous nucleotide substitutions should be more or less neutral and should evolve at a similar rate to non-coding regions of genomes. Many evolutionary analyses have used such an approach (Nei, 2005). However, it has been shown recently that synonymous sites could not be neutral. Different causes may be responsible for such a phenomenon, for example, the genetic code bias, the sites required in splicing mechanisms or those involved in RNA secondary structure and other aspects related to functional RNAs (Parmley et al., 2006; Xing and Lee, 2006). Nevertheless, these constraints in evolutionary rates of synonymous sites do not substantively affect the kS–kA metrics, and the major conclusions based on this method remain valid.
Silva and Kidwell (2000) proposed a new approach based on the comparison of the kA/kS ratio of TE with those of host genes. Such a method was applied to the P element in comparison to the Adh and per genes of host species. They observed that kS of TE can be low compared to that of host genes, suggesting that this could be explained by a smaller divergence time due to an HT.
Since the codon usage can differ widely from one species to another, several authors have proposed to use this bias to infer HT. Indeed, a recent HT between species having different codon usage should be easily detectable since the codon usage of the transferred TE would be different from that of the new host species. Given this assumption, it is presumed that TE and host genes should have the same codon usage. To test this hypothesis, Lerat et al. (2000) have compared the codon usage of different Drosophila species containing P elements. No correlation was evidenced, and it was clearly shown that TEs are mainly AT rich. This is a general feature of all TEs, whatever their mode of transposition (see, for instance, Lerat et al., 2002). Such an observation cannot therefore be taken alone as a strong argument in favor of HT.
Recently, phylogenetic reconciliation using TreeMap, originally used to analyze species biogeography (Page, 1988), resolution of orthologous and paralogous gene lineages (Goodman et al., 1979; Page and Charleston, 1997, 1998) and host–parasite phylogenies (Paterson and Poulin, 1999; Skerikova et al., 2001; Jackson and Charleston, 2004), was used to infer historical associations between TEs and species (Almeida and Carareto, 2005). This method is based on the mapping of TE phylogeny onto the host one. This allows the outcome that maximizes the number of co-speciation events to be selected. The statistical significance of this number is then tested against a null distribution of co-speciation events obtained by randomizing the host phylogeny. The possibility of defining the HT direction (donor versus recipient species) is thus a strong argument for using the historical association methodology as a complementary approach alongside the ‘classical’ phylogenetic one. The Minos transposon HTs between species of the repleta and saltans groups inferred by this approach (Almeida and Carareto, 2005) were in agreement with those using sequence similarity and parsimony, attesting that it could be an additional strategy for this kind of study. So the concordance between the ‘classical’ analysis and the historical associations could be indicative of the strength of the claimed HT cases.
The patchy distribution of an element, that is, the presence of a specific TE in one or a few species inside a phylogenetic cluster lacking this element, can also be evidence of HT. This was the initial argument used in several works based on Southern blot, PCR and in situ hybridization (Daniels et al., 1990a, 1990b; Maruyama and Hartl, 1991). However, this is a weak demonstration, since stochastic losses or an elevated evolutionary rate of TEs in some lineages can also lead to patchy distributions (Table 2).
The inference of HT events summarized in Table 1 (Supplementary Tables 1 and 2) is based on the occurrence of one, two or three of the types of evidence (sequence similarity, patchy distribution and tree incongruence) considered as essential by Silva et al. (2004). In 13 cases, HT was inferred from a single type of evidence (sequence similarity: 5; patchy distribution: 6; tree incongruence: 2), 16 cases used a combination of two arguments (sequence similarity+patchy distribution: 5; sequence similarity+tree incongruence: 10; patchy distribution+tree incongruence: 1) and 16 used the three types of evidence or combinations of two of them, including a complementary analysis such as π, kA/kS and D statistics. Silva et al. (2004) proposed that the stronger cases of HT are those confirmed by the three types of evidence. Under this assumption, only 15.8% of the 97 putative cases listed in Table 1 and Supplementary Tables 1 and 2 could be considered as actual HTs. This does not mean that the remaining cases are not HTs, but it would be more prudent to apply several tests before making a conclusion. Genetic material can indeed be easily transferred between closely related species by intogression. However, several distortions might also be due to an inappropriate sampling of TEs or of species.
The possible mechanisms of HT
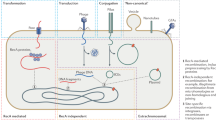
Several mechanisms and vectors have been proposed to explain how some genetic material could jump from one species to another. They vary from very simple ones, such as a direct transfer, to more complex systems involving intermediate vectors (Figure 1).
Different horizontal transfer mechanisms and vectors that have been suggested: (a) some TEs, such as long terminal repeat retrotransposons, are able to produce virus-like particles (VLPs) that may work as a vector; (b) TEs could be transported by DNA virus; (c) some parasites and parasitoids supposedly transfer TE DNA directly from the donor to the host; (d) TEs could hitchhike in the genomes of intracellular symbiotic bacteria (ISB), becoming its vector, or (e) bacteriophage could be an intermediate vector to transfer TE to intracellular symbiotic bacterial genomes. But in this case, the intracellular symbiotic bacteria (ISB) would be the final vector; or (f) the cycle would be similar to panel e, but the ISB is transferred among different host species by parasites or parasitoids.
The simplest mechanism of HT can be attributed to LTR retrotransposons (Figure 1a), since some of these elements, such as gypsy and copia elements, are able to produce virus-like particles (Miyake et al., 1987; Syomin et al., 1993; Lecher et al., 1997). In D. melanogaster, the gypsy virus-like particles are capable of efficiently infecting the germ line of strains devoid of active gypsy, and a high level of insertion activity is observed in their progeny (Song et al., 1994). Its infectious properties result from the expression of the env gene, encoding a protein responsible for its infectivity (Kim et al., 1994; Song et al., 1994; Teysset et al., 1998; Chalvet et al., 1999). Therefore, gypsy can potentially be transmitted as intracellular virus-like particles without the need of any vector. This element is widely distributed in the Drosophila genus. While it is likely to be an old component of these genomes, many cases of HT, attributed to their infectious properties, have been described (Alberola and de Frutos, 1996; Terzian et al., 2000; Vázquez-Manrique et al., 2000; Heredia et al., 2004).
Other virus-mediated HT processes involve DNA viruses. In this case, TEs take a viral shuttle (Figure 1b). For instance, it has been shown that the baculovirus isolated from the Lepidoptera Trichoplusia ni frequently harbors the piggyBac element (Fraser et al., 1985). This process may also provide a powerful means for horizontal transmission of DNA transposons among species (Miller and Miller, 1982; Fraser, 2001).
Drosophila parasites and parasitoids, such as mites and wasps, have also been pointed out as possible vectors of TE (Figure 1c). The mite Procteolaelaps regalis, for example, was proposed as the specific vector of P element transmission from D. willistoni to D. melanogaster (Houck et al., 1991). Drosophila DNA was recovered from this mite, while it was not integrated in its genome. The feeding behavior of the mite—piercing and sucking eggs and larvae—could be a mechanism for transferring DNA between species. Although a potential vector does not need to integrate the sequence transferred into its own genome, this point raises questions about the validity of such a mechanism, since the germ line of the recipient species must be reached. The P element was recovered in the gut of the mite, but there is no evidence that a transfer to D. melanogaster germ line was possible. A less specific but more convincing example is the HT of mariner from the moth Adoxophyes honmai to the parasitoid wasp Ascogaster reticulates. This was deduced from the high sequence similarity between the moth and the wasp mariners and the lack of this element in congeneric wasps (Yoshiyama et al., 2001).
Intracellular symbiotic bacteria, such as Wolbachia and spiroplasms, have also been considered as possible vectors at the intracellular level, since they live inside germ cells. These bacteria are widespread among the Drosophila genus. Moreover, Kondo et al. (2002) have shown the presence of Wolbachia-like genes in the bean beetle Callosobruchus chinensis genome. While Wu et al. (2004) did not identify any specific relationships for any gene between the genomes of D. melanogaster and its Wolbachia symbiont, in particular for TEs, it has been shown recently that a widespread lateral gene transfer from a Wolbachia to its Drosophila host is possible (Hotopp et al., 2007). However, it is worth recalling that an HT does not require a TE integration into the vector genome. The endosymbiont can only be a shuttle that offers a ‘hitchhike’ to TE.
Recently, Gavotte et al. (2007) have shown that bacteriophage infection is a common feature of Wolbachia, with 70% of the tested strains containing the parasite. Moreover, the authors showed an absence of congruence between phages and Wolbachia phylogenies, suggesting that these parasites can successfully transfer themselves horizontally. Considering that phages are implicated in transduction, a mechanism allowing genetic transfer between bacterial cells, they are relevant potential mediators of TE among intracellular symbiotic bacteria (Figure 1e). It is interesting to note that Wolbachia bacteriophage WO is able to infect free-living bacteria, notably increasing their potential as an HT vector. For example, the closest relative of Wolbachia WO is a phage found in the plant pathogen Xylella fastidiosa, which is transmitted by the Wolbachia-infected Glassy-winged sharpshooter (Simpson et al., 2000). While less information is available about the intracellular bacteria Spiroplasma, it has also been shown that they can be horizontally transferred by mites between Drosophila species (Jaenike et al., 2007). A model involving intracellular symbiotic bacteria or even DNA virus as primary TE vectors, with parasites or parasitoids, such as mites or wasps, as secondary vectors (Figure 1f) could therefore be relevant. However, it is important to stress that HTs are not restricted to a single mechanism, and all the models proposed are not mutually exclusive.
Finally, HT can also result from an introgression. This concerns closely related species between which hybrids can be partly fertile (in general, the heterogametic sex, according to the Haldane's rule). Such a phenomenon has been reported between D. bifasciata and D. imaii (Haring et al., 1995), the species of simulans complex (Lachaise et al., 1988), the species of the groups willistoni (reviewed by Bock, 1984) and saltans (Bicudo, 1973, 1979; Bicudo and Prioli, 1978) and, D. serido and D. buzzatii (Madi-Ravazzi et al., 1997). This may explain the sequence similarity of TEs between closely related species observed by several authors (Brunet et al., 1994; Haring et al., 1995; Silva and Kidwell, 2000; Almeida and Carareto, 2005). In such a case, an analysis of mitochondrial DNA polymorphism could be useful to confirm the introgression.
Donor and recipient species
The direction of HT is also of interest. Which is the donor and which is the recipient species? In 41 (40.6%) out of the 101 HT events proposed (Table 1 and Supplementary Tables 1 and 2), there is an indication of the donor species or at least of a group of donor species. In 27 (26.7%) of them, the species involved in the transfer are known, but the direction of the transfer could not be established. Finally, in 33 (32.7%) of them, the origin of the element could not be inferred. Among the 55 species considered as donor or recipient species, D. melanogaster (21 events) and D. simulans (10 events) have the largest participation. This prevalence seems to result from the fact that they are the most studied species of the genus Drosophila. The cosmopolitan nature of both species could be another factor responsible for the higher frequency of HTs. Indeed, colonization of new habitats may facilitate the encounter with new species and favor horizontal transmission (Biémont et al., 1999). The difference of HT frequency observed between these two species could be due to the older worldwide dispersal of D. melanogaster from tropical Africa compared to that of D. simulans (Capy et al., 1993). This suggests that D. simulans has had less time and fewer opportunities to experience HTs. However, alternative explanations cannot be ruled out. For instance, the proportion of TEs in D. simulans genome is lower than that of D. melanogaster (5 versus 15%; Dowsett and Young, 1982). One explanation for this could be the more recent world invasion of D. simulans, but another could be its higher Ne. Indeed, a higher Ne should lead to a more efficient elimination of TEs from the genome (see, for instance, Lynch and Conery, 2003). Alternative explanations, which will not be developed here, have also been proposed by Capy et al. (1994), Arnault and Dufournel (1994) and Silva and Kidwell (2000). They strongly underline the difficulties in explaining the differences observed between species as a result of the interaction of several evolutionary processes (populational, genomics, and so on.). More intensive investigations should indicate the reality or otherwise of the differences between the two species, on the one hand, and the prevalence of HT in cosmopolitan species, on the other hand.
Conclusions and perspectives
HT of TEs is a phenomenon frequently assumed in Drosophila, and the number of putative cases has increased during the last decades. However, several questions remain open: (i) the frequency of HTs; (ii) the mechanisms of transfer from one species to another; and (iii) when and why HT has occurred.
It is clear that HT frequency is not the same for different classes of TEs. This phenomenon is more frequent for DNA transposons, intermediate for LTR retrotransposons and rare but not null for non-LTR retrotransposons. Given that over a hundred families of TEs have been described in Drosophila and that only 21 elements are putatively involved in HTs, this suggests that successful HTs are rare. As yet unknown features of TEs may be required to achieve a complete HT, and the transfer of an active copy with a high transposition rate into a new host is insufficient. Moreover, HT has not been analyzed for all the elements. For instance, Sánchez-Gracia et al. (2005), using a comparison of the observed and expected nucleotide variation in a population genetic model, suggest that HT could be frequent in Drosophila. In this respect, among the 14 TEs common to D. melanogaster, D. simulans and D. yakuba, 70% appear to have undergone HT.
HT frequency is probably not constant in terms of space and time. Some species are probably more HT-prone. This may be the case for invasive species. In Drosophila, a detailed analysis of such species is quite possible since several of them are known to be cosmopolitan and more or less recently invasive. Along with D. melanogaster and D. simulans, there are, for instance, D. ananassae, D. malerkotliana, D. subobscura and more recently Zaprionus indianus. Thanks to the data available, including their phylogenies, genetics, biogeography and history, these species are probably highly useful for testing this hypothesis (Nardon et al., 2005).
Several methods have been proposed to identify HT. Since each of them has its own limitations, several independent methods should ideally be used before conclusions are drawn. In this respect, the re-analysis of putative cases, which are mainly based on a single approach, will probably show that additional methods reinforce some putative HTs and refute others.
Since none of the investigations purporting to demonstrate the mechanism of HT have been successful, several alternatives can be considered. Retrotransposons are able to produce infective particles that could be transferred directly, without an intermediary vector. However, even if it is possible, no direct evidence has been reported until now. For TEs that require a vector for HT, the best-rated candidates are probably among bacteria and viruses. In Drosophila, endosymbionts such as Wolbachia and Spiroplasma have been shown to be able to perform HT themselves (Montenegro et al., 2005). In theory, bacteria could transport the TE when infecting a new species. The large nuclear and cytoplasmic DNA viruses with their large genomes, such as mimiviruses, may have the ability to transport exogenous DNA (Raoult et al., 2004; Iyer et al., 2006). Moreover, parasites or parasitoids could be alternative intermediates, able to transfer bacteria and/or viruses, themselves containing TEs. In this respect, it may be worthwhile to apply the metagenomic approach to organisms between which HTs have already being detected. This should allow us to obtain a more relevant view of the ‘guilde’ of species hosted in a eukaryote. The comparison of the ‘guilde’ found in different organisms should help us identify the putative vector(s) or shuttle(s) involved in the transfer of genetic material among species.
References
Alberola TM, de Frutos R (1996). Molecular structure of a gypsy element of Drosophila subobscura (gypsyDs) constituting a degenerate form of insect retroviruses. Nucleic Acids Res 24: 914–923.
Almeida LM, Carareto CMA (2005). Multiple events of horizontal transfer of the Minos transposable element between Drosophila species. Mol Phyl Evol 35: 583–594.
Almeida LM, Carareto CMA (2006). Sequence heterogeneity and phylogenetic relationships between the copia retrotransposon in Drosophila species of the repleta and melanogaster groups. Genet Sel Evol 38: 535–550.
Arca B, Savakis C (2000). Distribution of the transposable element Minos in the genus Drosophila. Genetica 108: 263–267.
Arnault C, Dufournel I (1994). Genome and stresses: reactions against aggressions, behavior of transposable elements. Genetica 93: 149–160.
Bicudo HEMC (1973). Reproductive isolation in the saltans group of Drosophila. I. The saltans subgroup. Genetica 44: 313–329.
Bicudo HEMC (1979). Reproductive isolation in the saltans group of Drosophila. IV. The sturtevanti subgroup. Rev Bras Genet 2: 247–258.
Bicudo HEMC, Prioli AJ (1978). Reproductive isolation in the saltans group of Drosophila. II. The parasaltans subgroup. Genetica 48: 17–22.
Biémont C, Vieira C, Borie N, Lepetit D (1999). Transposable elements and genome evolution: the case of Drosophila simulans. Genetica 107: 113–120.
Bock IR (1984). Interspecific hybridization in the genus Drosophila. In: Hecht MK, Wallace B, Prance GT (eds). Evolutionary Biology, vol. 18. Plenum Press: New York, pp 41–70.
Brégliano JC, Kidwell MG (1983). Hybrid dysgenesis determinants. In: Shapiro J (ed). Mobile Genetic Elements. Academic Press, New York, pp 363–410.
Brunet F, Godin F, Bazin C, Capy P (1999). Phylogenetic analysis of Mos1-like transposable elements in the Drosophilidae. J Mol Evol 49: 760–768.
Brunet F, Godin F, David JR, Capy P (1994). The mariner transposable element in the Drosophilidae family. Heredity 73: 377–385.
Bucheton A, Paro R, Sang HM, Pelisson A, Finnegan D (1984). The molecular basis of I-R hybrid dysgenesis in Drosophila melanogaster: identification, cloning, and properties of the I factor. Cell 38: 153–163.
Capy P, Anxolabéhère D, Langin T (1994). The strange phylogenies of transposable elements: are horizontal transfers the only explanation? Trends Genet 10: 7–12.
Capy P, Pla E, David JR (1993). Phenotypic and geographical variability of morphometrical traits in natural populations of Drosophila melanogaster and D. simulans. I. Geographical variations. Genet Sel Evol 11: 517–536.
Casola C, Lawing AM, Betrán E, Feschotte C (2007). PIF-like transposons are common in Drosophila and have been repeatedly domesticated to generate new host genes. Mol Biol Evol 24: 1872–1888.
Castro JP, Carareto CMA (2004). Canonical P elements are transcriptionally active in the saltans group of Drosophila. J Mol Evol 59: 31–40.
Chalvet F, Teysset L, Terzian C, Prud'homme N, Santamaria P, Bucheton A et al. (1999). Proviral amplification of the gypsy endogenous retrovirus of Drosophila melanogaster involves env-independent invasion of the female germline. EMBO J 18: 2659–2669.
Clark JB, Maddison WP, Kidwell MG (1994). Phylogenetic analysis supports horizontal transfer of P transposable elements. Mol Biol Evol 11: 40–50.
Clark JB, Altheide TK, Scholosser MJ, Kidwell MG (1995). Molecular evolution of P transposable elements in genus Drosophila. I. The saltans and willistoni species groups. Mol Biol Evol 12: 902–913.
Clark JB, Kidwell MG (1997). A phylogenetic perspective on P transposable element evolution in Drosophila. Proc Natl Acad Sci USA 94: 11428–11433.
Clark JB, Silva JC, Kidwell MG (2002). Evidence for horizontal transfer of P transposable elements. In: Syvanen M, Kado CI (eds). Horizontal Gene Transfer. Academic Press: London, pp 161–171.
Cummings MP (1994). Transmission patterns of eukaryotic transposable elements: arguments for and against horizontal transfer. Trends Ecol Evol 9: 141–145.
Daniels SB, Chovnic A, Boussy IS (1990a). Distribution of hobo transposable elements in the genus Drosophila. Mol Biol Evol 7: 589–606.
Daniels SB, Petterson KR, Strausbaugh LD, Kidwell MG, Chovnick AC (1990b). Evidence for horizontal transmission of the P transposable elements between Drosophila species. Genetics 124: 339–355.
Daniels SB, Strausbaugh LD, Ehrman L, Armstrong R (1984). Sequences homologous to P elements occur in Drosophila paulistorum. Proc Natl Acad Sci USA 81: 6794–6797.
Dowsett AP, Young MW (1982). Differing levels of dispersed repetitive DNA among closely related species of Drosophila. Proc Natl Acad Sci USA 79: 4570–4574.
Evgen′ev M, Zelentsova H, Mnjoian L, Poluectova H, Kidwell MG (2000). Invasion of Drosophila virilis by the Penelope transposable element. Chromosoma 109: 350–357.
Fablet M, Souames S, Biémont C, Vieira C (2007). Evolutionary pathways of the tirant LTR retrotransposon in the Drosophila melanogaster subgroup of species. J Mol Evol 64: 438–447.
Fraser MJ (2001). The TTAA-specific family of transposable elements: identification, functional characterization, and utility for transformation of insects. In: Handler AM, James AA (eds). Insect Transgenesis: Methods and Applications. CRC Press: Boca Raton, FL, pp 249–268.
Fraser MJ, Brusca JS, Smith GE, Summers MD (1985). Transposon-mediated mutagenesis of a baculovirus. Virology 145: 356–361.
García-Planells J, Paricio N, Clark JB, Frutos R, Kidwell MG (1998). Molecular evolution of P transposable elements in the genus Drosophila. II. The obscura species group. J Mol Evol 47: 282–291.
Gavotte L, Henri H, Stouthamer R, Charif D, Charlat S, Boulétreau M et al. (2007). A survey of the bacteriophage WO in the endosymbiotic bacteria Wolbachia. Mol Bio Evol 24: 427–435.
Goodman MJ, Czlusniak J, Moore GW, Romero-Herrera AE (1979). Fitting the gene lineage to its species lineage: a parsimony strategy illustrated by cladograms constructed from globin sequences. Syst Zool 28: 132–168.
Hagemann S, Haring E, Pinsker W (1996). Repeated horizontal transfer of P transposons between Scaptomyza pallida and Drosophila bifasciata. Genetica 98: 43–51.
Hagemann S, Haring E, Pinsker W (1998). Horizontal transmission vs vertical inheritance of P elements in Drosophila and Scaptomyza: has the M-type subfamily spread from East Asia? J Zool Syst Evol Res 36: 75–83.
Hagemann S, Miller WJ, Pinsker W (1992). Identification of a complete P-element in the genome of Drosophila bifasciata. Nucleic Acids Res 20: 409–413.
Haring E, Hagemann S, Pinsker W (1995). Different evolutionary behaviour of P element subfamilies: M-type and O-type elements in Drosophila bifasciata and D. imaii. Gene 163: 197–202.
Haring E, Hagemann S, Pinsker W (2000). Ancient and recent horizontal invasions of Drosophilids by P elements. J Mol Evol 51: 577–586.
Hartl DL, Lohe AR, Lozovskaya ER (1997). Modern thoughts on an ancient mariner: function, evolution, regulation. Annu Rev Genet 31: 337–358.
Heredia F, Loreto ELS, Valente VLS (2004). Complex evolution of gypsy in Drosophilid species. Mol Biol Evol 21: 1831–1842.
Hochstenbach R, Harhangi H, Schouren K, Bindels P, Suijkerbuijk R, Hennig W (1996). Transcription of gypsy elements in a Y-chromosome male fertility gene of Drosophila hydei. Genetics 142: 437–446.
Hotopp JCD, Clark ME, Oliveira DCSG, Foster JM, Fischer P, Munoz Torres MC et al. (2007). Widespread lateral gene transfer from intracellular bacteria to multicellular eukaryotes. Science 317: 1753–1756.
Houck MA, Clark JB, Peterson KR, Kidwell MG (1991). Possible horizontal transfer of Drosophila genes by the mite Proctolaelaps regalis. Science 253: 1125–1129.
Iyer LM, Balaji S, Koonin EV, Aravind L (2006). Evolutionary genomics of nucleo-cytoplasmic large DNA viruses. Virus Res 117: 156–184.
Jackson AP, Charleston MA (2004). A cophylogenetic perspective of RNA-virus evolution. Mol Biol Evol 21: 45–57.
Jaenike J, Polak M, Fiskin A, Helou M, Minhas M (2007). Interspecific transmission of endosymbiotic Spiroplasma by mites. Bio Lett 3: 23–25.
Jordan IK, Matyunina LV, McDonald JF (1999). Evidence for the recent horizontal transfer of long terminal repeat retrotransposon. Proc Natl Acad Sci USA 96: 12621–12625.
Jordan IK, McDonald JF (1998). Evolution of the copia retrotransposon in the Drosophila melanogaster species subgroup. Mol Biol Evol 15: 1160–1171.
Kidwell MG (1983). Evolution of hybrid dysgenesis determinants in Drosophila melanogaster. Proc Natl Acad Sci USA 80: 1655–1659.
Kim A, Terzian C, Santamaria P, Pélisson A, Prud'homme N, Bucheton A (1994). Retroviruses in invertebrates: the gypsy retrotransposon is apparently an infectious retrovirus of Drosophila melanogaster. Proc Natl Acad Sci USA 91: 1285–1289.
King JL, Jukes TH (1969). Non-Darwinian evolution. Science 164: 788–798.
Kondo N, Nikoh N, Ijichi N, Shimada M, Fukatsu T (2002). Genome fragment of Wolbachia endosymbiont transferred to X chromosome of host insect. Proc Natl Acad Sci USA 99: 14280–14285.
Kordis D, Gubensek F (1998). Unusual horizontal transfer of a long interspersed nuclear element between distant vertebrate classes. Proc Natl Acad Sci USA 95: 10704–10709.
Kotnova AP, Glukhov IA, Karpova NN, Salenko VB, Lyubomirskaya NV, Ilyin YV (2007). Evidence for recent horizontal transfer of gypsy-homologous LTR-retrotransposon gtwin into Drosophila erecta followed by its amplification with multiple aberrations. Gene 396: 39–45.
Lachaise D, Cariou ML, David JR, Lemeunier F, Tsacas L, Ashburner M (1988). Historical biogeography of the Drosophila melanogaster species subgroup. Evol Biol 22: 159–225.
Lecher P, Bucheton A, Pelisson A (1997). Expression of the Drosophila retrovirus gypsy as ultrastructurally detectable particles in the ovaries of flies carrying a permissive flamenco allele. J Gen Virol 78: 2379–2388.
Lerat E, Biémont C, Capy P (2000). Codon usage and the origin of P elements. Mol Biol Evol 17: 467–468.
Lerat E, Capy P, Biémont C (2002). Codon usage by transposable elements and their host genes among five species. J Mol Evol 54: 625–637.
Le Rouzic A, Capy P (2005). The first steps of transposable elements invasion: parasitic strategy versus genetic drift. Genetics 169: 1033–1043.
Lohe AR, Moriyama EN, Lidholm D-A, Hartl DL (1995). Horizontal transmission, vertical inactivation, and stochastic loss of mariner-like transposable elements. Mol Biol Evol 12: 62–72.
Loreto ELS, Valente VLS, Zaha A, Silva JC, Kidwell MG (2001). Drosophila mediopunctata P elements: a new example of horizontal transfer. J Hered 92: 375–381.
Luan DD, Korman MH, Jakubczak JL, Eickbush TH (1993). Reverse transcription of R2Bm RNA is primed by a nick at the chromosomal target site: a mechanism for non-LTR retrotransposition. Cell 72: 595–605.
Ludwig A, Loreto ELS (2007). Evolutionary pattern of the gtwin retrotransposon in the Drosophila melanogaster subgroup. Genetica 130: 161–168.
Lynch M, Conery JS (2003). The origins of genome complexity. Science 302: 1401–1404.
Lyozin GT, Makarova KS, Velikodvorskaja VV, Zelentsova HS, Khechumian RR, Kidwell MG et al. (2001). The structure and evolution of Penelope in the virilis species group of Drosophila: an ancient lineage of retroelements. J Mol Evol 52: 445–456.
Madi-Ravazzi L, Bicudo HEM, Manzato JA (1997). Reproductive compatibility and chromosome pairing in the Drosophila buzzatii complex. Cytobios 89: 21–30.
Malik HS, Burke WD, Eickbush TH (1999). The age and evolution of non-LTR retrotransposable elements. Mol Biol Evol 16: 793–805.
Martínez-Sebastián MJ, Hernandez M, Mejias B, Gas ME, Perez A, Pascual L et al. (2002). Evolutionary patterns of the gypsy and bilbo retrotransposon families in the Drosophila species of the obscura group. Mol Phylogenet Evol 25: 254–266.
Maruyama K, Hartl DL (1991). Evidence for interspecific transfer of the transposable element mariner between Drosophila and Zaprionus. J Mol Evol 33: 514–524.
Maside X, Bartolomé C, Charlesworth B (2003). Inferences on the evolutionary history of the S-element family of Drosophila melanogaster. Mol Biol Evol 20: 1183–1187.
Miller DW, Miller LK (1982). A virus mutant with an insertion of a copia-like transposable element. Nature 299: 562–564.
Miyake T, Mae N, Shiba T, Kondo S (1987). Production of virus-like particles by the transposable genetic element, copia, of Drosophila melanogaster. Mol Gen Genet 207: 29–37.
Mizrokhi LJ, Mazo AM (1990). Evidence for horizontal transmission of the mobile element jockey between distant Drosophila species. Proc Natl Acad SciUSA 87: 9216–9220.
Montenegro H, Solferini VN, Klaczko LB, Hurst GDD (2005). Male-killing Spiroplasma naturally infecting Drosophila melanogaster. Insect Mol Biol 14: 281–287.
Morales-Hojas R, Vieira CP, Vieira J (2006). The evolutionary history of the transposable element Penelope in the Drosophila virilis group of species. J Mol Evol 63: 262–273.
Nardon C, Deceliere G, Loevenbruck C, Weiss M, Vieira C, Biemont C (2005). Is genome size influenced by colonization of new environments in dipteran species? Mol Ecol 14: 869–878.
Nei M (2005). Selectionism and neutralism in molecular evolution. Mol Biol Evol 22: 2318–2342.
Page RDM (1988). Quantitative cladistic biogeography: constructing and comparing area cladograms. Syst Zool 37: 254–270.
Page RDM, Charleston MA (1997). From gene to organismal phylogeny: reconciled trees and the gene tree species tree problem. Mol Phylogenet Evol 7: 231–240.
Page RDM, Charleston MA (1998). Trees within trees: phylogeny and historical associations. Trends Ecol Evol 13: 356–359.
Parmley JL, Chamary JV, Hurst LD (2006). Evidence for purifying selection against synonymous mutations in mammalian exonic splicing enhancers. Mol Biol Evol 23: 301–309.
Pascual L, Periquet G (1991). Distribution of hobo transposable elements in natural populations of Drosophila melanogaster. Mol Biol Evol 8: 282–296.
Paterson AM, Poulin R (1999). Have chondracanthid copepods co-speciated with their teleost hosts? Syst Parasitol 44: 79–85.
Raoult D, Audic S, Robert C, Abergel C, Renesto P, Ogata H et al. (2004). The 1.2-megabase genome sequence of mimivirus. Science 306: 1344–1350.
Robertson HM, Lampe DJ (1995). Recent horizontal transfer of a mariner transposable element among and between Diptera and Neuroptera. Mol Biol Evol 12: 850–862.
Robertson HM, MacLeod EG (1993). Five major subfamilies of mariner transposable elements in insects, including the Mediterranean fruit fly, and related arthropods. Insect Mol Biol 2: 125–139.
Sánchez-Gracia A, Maside X, Charlesworth B (2005). High rate of horizontal transfer of transposable elements in Drosophila. Trends Genet 21: 200–203.
Silva JC, Kidwell MG (2000). Horizontal transfer and selection in the evolution of P elements. Mol Biol Evol 17: 1542–1557.
Silva JC, Loreto EL, Clark JB (2004). Factors that affect the horizontal transfer of transposable elements. Curr Issues Mol Biol 6: 57–72.
Simmons G (1992). Horizontal transfer of hobo transposable elements within the Drosophila melanogaster species complex: evidence from DNA sequencing. Mol Biol Evol 9: 1050–1060.
Simpson AJ, Reinac FC, Arruda P, Abreu FA, Acencio M, Alvarenga R et al. (2000). The sequence of the plant pathogen Xylella fastidiosa. Nature 406: 151–157.
Skerikova A, Hypsa V, Scholz T (2001). Phylogenetic of european species of Proteocephalus (Cestoda:Proteocephalidea): compatibility of molecular and morphological data, and parasite–host coevolution. Int J Parasitol 31: 1121–1128.
Song SU, Gerasimova T, Kurkulos M, Boeke JD, Corces VG (1994). An env-like protein encoded by a Drosophila retroelement: evidence that gypsy is an infectious retrovirus. Genes Dev 8: 2046–2057.
Syomin BV, Kandror KV, Semakin AB, Tsuprun VL, Stepanov AS (1993). Presence of the gypsy (MDG4) retrotransposon in extracellular virus-like particles. FEBS Lett 323: 285–288.
Terzian C, Ferraz C, Demaille J, Bucheton A (2000). Evolution of the gypsy endogenous retrovirus in the Drosophila melanogaster subgroup. Mol Biol Evol 17: 908–914.
Teysset L, Burns JC, Shike H, Sullivan BL, Bucheton A, Terzian C (1998). A Moloney murine leukemia virus-based retroviral vector pseudotyped by the insect retroviral gypsy envelope can infect Drosophila cells. J Virol 72: 853–856.
Torti C, Gomulski LM, Bonizzoni M, Murelli V, Moralli D, Guglielmino CR et al. (2000). Evolution of gypsy endogenous retrovirus in the Drosophila obscura species group. Mol Biol Evol 17: 1185–1193.
Vázquez-Manrique RP, Hernández M, Martınéz-Sebastian MJ, de Frutos R (2000). Evolution of gypsy endogenous retrovirus in the Drosophila obscura species group. Mol Biol Evol 17: 1185–1193.
Wu M, Sun LV, Vamathevan J, Riegler M, Deboy R, Brownlie JC et al. (2004). Phylogenomics of the reproductive parasite Wolbachia pipientis wMel: a streamlined genome overrun by mobile genetic elements. PLoS Biol 2: 327–341.
Xing Y, Lee C (2006). Can RNA selection pressure distort the measurement of Ka/Ks? Gene 370: 1–5.
Yoshiyama M, Tu Z, Kainoh Y, Honda H, Shono T, Kimura K (2001). Possible horizontal transfer of a transposable element from host to parasitoid. Mol Biol Evol 18: 1952–1958.
Acknowledgements
We gratefully acknowledge funding from the CAPES-COFECUB International Cooperation Program (to CMAC, ELSL and PC), FAPESP and CNPq (to CMAC and ELSL).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Heredity website (http://www.nature.com/hdy)
Supplementary information
Rights and permissions
About this article
Cite this article
Loreto, E., Carareto, C. & Capy, P. Revisiting horizontal transfer of transposable elements in Drosophila. Heredity 100, 545–554 (2008). https://doi.org/10.1038/sj.hdy.6801094
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6801094
Keywords
This article is cited by
-
Modeling early germline immunization after horizontal transfer of transposable elements reveals internal piRNA cluster heterogeneity
BMC Biology (2023)
-
Multiple horizontal transfers of a Helitron transposon associated with a parasitoid wasp
Mobile DNA (2022)
-
Interrogating the 5’UTR tandem repeats of retrotransposon roo of Drosophila about horizontal transfer
Genetica (2021)
-
Horizontal transfer and evolution of transposable elements in vertebrates
Nature Communications (2020)
-
Transcriptionally promiscuous “blurry” promoters in Tc1/mariner transposons allow transcription in distantly related genomes
Mobile DNA (2019)