Abstract
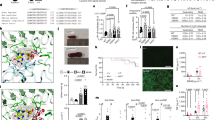
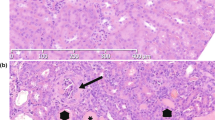
Epistatic interactions between the non-autoimmune strains 129 and C57BL/6 (B6), used for generating gene-targeted animals, can induce a lupus-like disease. Genome-wide scan analyses of testcross progeny between these two strains have identified several lupus susceptibility loci, with the strongest linkage to the production of autoantibodies (auto-Abs) displayed by an interval on chromosome 1 of 129 origin (Sle16). However, the contribution of B6 loci to the lupus phenotype remained unknown. We used a congenic approach to deduce the contribution to the autoimmune traits of the B6 genomic interval on chromosome 3 (Sle18), previously shown to be linked to antinuclear Ab production. This interval, when transferred on a 129 background (a strain termed 129.B6–Sle18), promoted auto-Ab production targeting a broad spectrum of autoantigens, expansion of activated CD4+T and B cells and mild glomerulonephritis. Surprisingly, these immunological and serological defects were accompanied by a significant increase in the percentage of regulatory T cells (Tregs; CD4+ Foxp3+). However, these cells, that expressed lower levels of Foxp3, had no impaired regulatory function when tested in vitro. These findings illustrate further the efficacy of congenic dissection for functional characterisation of individual lupus susceptibility loci and highlight the contribution of loci derived from non-autoimmune strains to the disease pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Rhodes B, Vyse TJ . General aspects of the genetics of SLE. Autoimmunity 2007; 40: 550–559.
Harley JB, Alarcon-Riquelme ME, Criswell LA, Jacob CO, Kimberly RP, Moser KL et al. Genome-wide association scan in women with systemic lupus erythematosus identifies susceptibility variants in ITGAM, PXK, KIAA1542 and other loci. Nat Genet 2008; 40: 204–210.
Kozyrev SV, Abelson AK, Wojcik J, Zaghlool A, Linga Reddy MV, Sanchez E et al. Functional variants in the B-cell gene BANK1 are associated with systemic lupus erythematosus. Nat Genet 2008; 40: 211–216.
Hom G, Graham RR, Modrek B, Taylor KE, Ortmann W, Garnier S et al. Association of systemic lupus erythematosus with C8orf13-BLK and ITGAM-ITGAX. N Engl J Med 2008; 358: 900–909.
Nath SK, Han S, Kim-Howard X, Kelly JA, Viswanathan P, Gilkeson GS et al. A nonsynonymous functional variant in integrin-alpha(M) (encoded by ITGAM) is associated with systemic lupus erythematosus. Nat Genet 2008; 40: 152–154.
Graham DS, Graham RR, Manku H, Wong AK, Whittaker JC, Gaffney PM et al. Polymorphism at the TNF superfamily gene TNFSF4 confers susceptibility to systemic lupus erythematosus. Nat Genet 2008; 40: 83–89.
Wakeland EK, Liu K, Graham RR, Behrens TW . Delineating the genetic basis of systemic lupus erythematosus. Immunity 2001; 15: 397–408.
Morel L, Blenman KR, Croker BP, Wakeland EK . The major murine systemic lupus erythematosus susceptibility locus, Sle1, is a cluster of functionally related genes. Proc Natl Acad Sci USA 2001; 98: 1787–1792.
Bygrave AE, Rose KL, Cortes-Hernandez J, Warren J, Rigby RJ, Cook HT et al. Spontaneous autoimmunity in 129 and C57BL/6 mice-implications for autoimmunity described in gene-targeted mice. PLoS Biol 2004; 2: E243.
Drake CG, Rozzo SJ, Hirschfeld HF, Smarnworawong NP, Palmer E, Kotzin BL . Analysis of the New Zealand Black contribution to lupus-like renal disease. Multiple genes that operate in a threshold manner. J Immunol 1995; 154: 2441–2447.
Vyse TJ, Rozzo SJ, Drake CG, Izui S, Kotzin BL . Control of multiple autoantibodies linked with a lupus nephritis susceptibility locus in New Zealand black mice. J Immunol 1997; 158: 5566–5574.
Wandstrat AE, Nguyen C, Limaye N, Chan AY, Subramanian S, Tian XH et al. Association of extensive polymorphisms in the SLAM/CD2 gene cluster with murine lupus. Immunity 2004; 21: 769–780.
Carlucci F, Cortes-Hernandez J, Fossati-Jimack L, Bygrave AE, Walport MJ, Vyse TJ et al. Genetic dissection of spontaneous autoimmunity driven by 129-derived chromosome 1 Loci when expressed on C57BL/6 mice. J Immunol 2007; 178: 2352–2360.
Bickerstaff MC, Botto M, Hutchinson WL, Herbert J, Tennent GA, Bybee A et al. Serum amyloid P component controls chromatin degradation and prevents antinuclear autoimmunity. Nat Med 1999; 5: 694–697.
Bolland S, Ravetch JV . Spontaneous autoimmune disease in Fc(gamma)RIIB-deficient mice results from strain-specific epistasis. Immunity 2000; 13: 277–285.
Wu X, Jiang N, Deppong C, Singh J, Dolecki G, Mao D et al. A role for the Cr2 gene in modifying autoantibody production in systemic lupus erythematosus. J Immunol 2002; 169: 1587–1592.
Miwa T, Maldonado MA, Zhou L, Sun X, Luo HY, Cai D et al. Deletion of decay-accelerating factor (CD55) exacerbates autoimmune disease development in MRL/lpr mice. Am J Pathol 2002; 161: 1077–1086.
Xue D, Shi H, Smith JD, Chen X, Noe DA, Cedervall T et al. A lupus-like syndrome develops in mice lacking the Ro 60-kDa protein, a major lupus autoantigen. Proc Natl Acad Sci USA 2003; 100: 7503–7508.
Heidari Y, Bygrave AE, Rigby RJ, Rose KL, Walport MJ, Cook HT et al. Identification of chromosome intervals from 129 and C57BL/6 mouse strains linked to the development of systemic lupus erythematosus. Genes Immun 2006; 7: 592–599.
Shi X, Xie C, Kreska D, Richardson JA, Mohan C . Genetic dissection of SLE: SLE1 and FAS impact alternate pathways leading to lymphoproliferative autoimmunity. J Exp Med 2002; 196: 281–292.
Chu EB, Ernst DN, Hobbs MV, Weigle WO . Maturational changes in CD4+ cell subsets and lymphokine production in BXSB mice. J Immunol 1994; 152: 4129–4138.
Chu EB, Hobbs MV, Wilson CB, Romball CG, Linsley PS, Weigle WO . Intervention of CD4+ cell subset shifts and autoimmunity in the BXSB mouse by murine CTLA4Ig. J Immunol 1996; 156: 1262–1268.
Croker BP, Gilkeson G, Morel L . Genetic interactions between susceptibility loci reveal epistatic pathogenic networks in murine lupus. Genes Immun 2003; 4: 575–585.
Lawson BR, Baccala R, Song J, Croft M, Kono DH, Theofilopoulos AN . Deficiency of the cyclin kinase inhibitor p21(WAF-1/CIP-1) promotes apoptosis of activated/memory T cells and inhibits spontaneous systemic autoimmunity. J Exp Med 2004; 199: 547–557.
Trendelenburg M, Manderson AP, Fossati-Jimack L, Walport MJ, Botto M . Monocytosis and accelerated activation of lymphocytes in C1q-deficient autoimmune-prone mice. Immunology 2004; 113: 80–88.
Samardzic T, Marinkovic D, Danzer CP, Gerlach J, Nitschke L, Wirth T . Reduction of marginal zone B cells in CD22-deficient mice. Eur J Immunol 2002; 32: 561–567.
Amano H, Amano E, Moll T, Marinkovic D, Ibnou-Zekri N, Martinez-Soria E et al. The Yaa mutation promoting murine lupus causes defective development of marginal zone B cells. J Immunol 2003; 170: 2293–2301.
Mohan C, Morel L, Yang P, Wakeland EK . Accumulation of splenic B1a cells with potent antigen-presenting capability in NZM2410 lupus-prone mice. Arthritis Rheum 1998; 41: 1652–1662.
Xu Z, Butfiloski EJ, Sobel ES, Morel L . Mechanisms of peritoneal B-1a cells accumulation induced by murine lupus susceptibility locus Sle2. J Immunol 2004; 173: 6050–6058.
Monk CR, Spachidou M, Rovis F, Leung E, Botto M, Lechler RI et al. MRL/Mp CD4+, CD25− T cells show reduced sensitivity to suppression by CD4+, CD25+ regulatory T cells in vitro: a novel defect of T cell regulation in systemic lupus erythematosus. Arthritis Rheum 2005; 52: 1180–1184.
Gaffney PM, Kearns GM, Shark KB, Ortmann WA, Selby SA, Malmgren ML et al. A genome-wide search for susceptibility genes in human systemic lupus erythematosus sib-pair families. Proc Natl Acad Sci USA 1998; 95: 14875–14879.
Haywood ME, Hogarth MB, Slingsby JH, Rose SJ, Allen PJ, Thompson EM et al. Identification of intervals on chromosomes 1, 3, and 13 linked to the development of lupus in BXSB mice. Arthritis Rheum 2000; 43: 349–355.
Mohan C, Alas E, Morel L, Yang P, Wakeland EK . Genetic dissection of SLE pathogenesis. Sle1 on murine chromosome 1 leads to a selective loss of tolerance to H2A/H2B/DNA subnucleosomes. J Clin Invest 1998; 101: 1362–1372.
Vratsanos GS, Jung S, Park YM, Craft J . CD4(+) T cells from lupus-prone mice are hyperresponsive to T cell receptor engagement with low and high affinity peptide antigens: a model to explain spontaneous T cell activation in lupus. J Exp Med 2001; 193: 329–337.
Sakaguchi S, Ono M, Setoguchi R, Yagi H, Hori S, Fehervari Z et al. Foxp3+ CD25+ CD4+ natural regulatory T cells in dominant self-tolerance and autoimmune disease. Immunol Rev 2006; 212: 8–27.
Miyara M, Amoura Z, Parizot C, Badoual C, Dorgham K, Trad S et al. Global natural regulatory T cell depletion in active systemic lupus erythematosus. J Immunol 2005; 175: 8392–8400.
Lee JH, Wang LC, Lin YT, Yang YH, Lin DT, Chiang BL . Inverse correlation between CD4+ regulatory T-cell population and autoantibody levels in paediatric patients with systemic lupus erythematosus. Immunology 2006; 117: 280–286.
Valencia X, Yarboro C, Illei G, Lipsky PE . Deficient CD4+CD25high T regulatory cell function in patients with active systemic lupus erythematosus. J Immunol 2007; 178: 2579–2588.
Chen Y, Cuda C, Morel L . Genetic determination of T-cell help in loss of tolerance to nuclear antigens. J Immunol 2005; 174: 7692–7702.
Cuda CM, Wan S, Sobel ES, Croker BP, Morel L . Murine lupus susceptibility locus Sle1a controls regulatory T cell number and function through multiple mechanisms. J Immunol 2007; 179: 7439–7447.
Atencio S, Amano H, Izui S, Kotzin BL . Separation of the New Zealand Black genetic contribution to lupus from New Zealand Black determined expansions of marginal zone B and B1a cells. J Immunol 2004; 172: 4159–4166.
Fossati L, Takahashi S, Merino R, Iwamoto M, Aubry JP, Nose M et al. An MRL/MpJ-lpr/lpr substrain with a limited expansion of lpr double-negative T cells and a reduced autoimmune syndrome. Int Immunol 1993; 5: 525–532.
Wolfowicz CB, Sakorafas P, Rothstein TL, Marshak-Rothstein A . Oligoclonality of rheumatoid factors arising spontaneously in lpr/lpr mice. Clin Immunol Immunopathol 1988; 46: 382–395.
Acknowledgements
We thank Mrs Margarita Lewis for technical assistance with the processing of tissue for histological studies and the staff of the Biological Services Unit at our institution for the care of the animals involved in this study. We are grateful to Dr Dumitriu for her technical help with the T-suppressor assays. This study was supported by the Wellcome Trust (Grant no. 071467).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Heidari, Y., Fossati-Jimack, L., Carlucci, F. et al. A lupus-susceptibility C57BL/6 locus on chromosome 3 (Sle18) contributes to autoantibody production in 129 mice. Genes Immun 10, 47–55 (2009). https://doi.org/10.1038/gene.2008.78
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2008.78
Keywords
This article is cited by
-
Quantitative PPARγ expression affects the balance between tolerance and immunity
Scientific Reports (2016)
-
Genetics of SLE: evidence from mouse models
Nature Reviews Rheumatology (2010)