Abstract
Age-related macular degeneration (AMD) is an inflammatory disease, which causes visual impairment and blindness in older people. The proteins of the complement system are central to the development of this disease. Local and systemic inflammation in AMD are mediated by the deregulated action of the alternative pathway of the complement system. Variants in complement system genes alter an individual's risk of developing AMD. Recent studies have shown how some risk-associated genetic variants alter the function of the complement system. In this review, we describe the evolution of the complement system and bring together recent research to form a picture of how changes in complement system genes and proteins affect the function of the complement cascade, and how this affects the development of AMD. We discuss the application of this knowledge to prevention and possible future treatments of AMD.
Similar content being viewed by others
Introduction
Age-related macular degeneration (AMD) is a major cause of blindness in people of European descent.1 This disease is closely linked to the complement system by its aetiology and pathology. The complement system is an ancient defence mechanism against infectious microbes.2, 3 It has many other important physiological functions, including clearance of apoptotic cells and immune complexes, and other roles related to tissue homeostasis.4 Because of the multitude of interactions between complement and non-complement proteins, the complement ‘cascade’ may be best represented as a protein network among several other interacting pathways.5
For the complement system to cause AMD, there must be initiation (a ‘trigger’), amplification, and local retinal damage caused by the process. The alternative pathway of the complement system is spontaneously activating and self-perpetuating. The complement system therefore acts not only as a messenger network and the instrument by which damage is caused, but it may also be important in the initiation of the disease process. This may explain why individuals with different variants of diverse complement system genes may respond to the same environmental triggers with more or less harmful inflammation.
In this review, we will discuss recent progress in understanding how the complement system is related to the initiation, development, and treatment of AMD. We will focus particularly on recent genetic associations and studies of function that shed light on pathophysiological mechanisms and therefore offer insights that may provide opportunities for the development of new treatments.
Age-related macular degeneration
AMD is experienced by patients as progressive distortions and diminutions of vision, particularly of central vision. Episodes of declining vision are often sudden, leaving scotomas that do not subsequently improve.6 The clinician observes AMD in its earliest stages as drusen, which are deposits of cellular debris and inflammatory material seen between the retinal pigment epithelium and Bruch's membrane.7 The word ‘drusen’ is from the German word meaning geode, a spherical collection of crystals sometimes found in rock.8 Drusen do not themselves affect vision and many older people develop drusen without progressing to AMD. However, the presence of drusen is a sign of increased risk of development of the disease, and is in part determined by some of the same genes that affect AMD risk.9 More severe AMD is characterised by the development of either geographic atrophy (‘dry’ AMD) or neovascular AMD (exudative, ‘wet’ AMD). Geographic atrophy is more common and appears as hypopigmented patches on the retina. Neovascular AMD is rarer, but is responsible for more severe visual impairment. Fragile new blood vessels form (choroidal neovascularisation) and may break, resulting in subretinal haemorrhages and scarring.8
The evolution of complement
The proteins of the complement system are responsible for a major part of the human immune response against microbes and can cause damage to self-tissues. The evolution and relatedness of complement proteins explain the overlapping—and sometimes competing—functions of several components.
Infection with microbes has been the major evolutionary selection pressure driving the development of the complement system.2, 10 Tight coevolution of hosts (including humans) and infectious microbes has occurred throughout evolutionary history and resulted in the evolution of microbes that evade host immune recognition. Likely all successful infectious microbes have developed mechanisms to subvert or evade host complement attack by binding or mimicking complement proteins.11 For example, Neisseria meningitidis specifically survive in human hosts and express proteins that bind to human negative regulators of the complement system such as complement factor H (CFH) in order to protect themselves from complement attack.12 A selection pressure is therefore exerted on the microbe, which evolves quickly due to its short generation time, to successfully deceive the host immune system. A countervailing selection pressure is also exerted on the host to detect infectious microbes, to change its own proteins to prevent them either being bound to microbes with high affinity or to have a configuration different from any mimicked proteins on the microbe's exterior. Changes in the complement system will often be deleterious, however, given the importance of its many functions.
The evolutionary history of the complement system has been traced by searching for complement genes in distantly related animals. The results of this search suggest that the oldest parts of the system are complement component 3 (C3) and complement factor B (CFB), which were identified in sea anemones of the phylum Cnidaria.10 Cnidaria diverged from Bilateralia (which include vertebrates and therefore humans) ∼1300 million years ago, indicating that the basic complement system is at least of this age.10 Given the central role of C3 and CFB in the complement pathways, it is not surprising that they are the oldest elements of the system. The alternative pathway is the oldest of the complement activation pathways. The lectin pathway evolved subsequently and was followed by the classical pathway.10
Many of the complement genes contain small exons, which encode short peptides called short consensus repeats (SCRs) or complement control proteins, many of which have a specific enzymatic or binding function.13 These exons have allowed rapid evolution of complement proteins by exon shuffling.13 In this evolutionary process, an exon coding for a peptide with a particular property (eg an enzymatic action), may be duplicated, inserted or deleted, either within or between genes. This is the basis for generating modular proteins, which include not only complement proteins, but many other proteins such as those of the clotting cascade. Examples of this include the SCR-containing complement regulators and C1r, C1s, C2, and C4 proteins, which possess serine protease modules, allowing them to enzymatically cleave serine peptide bonds such as those found in C3 and C5.14
During evolution, the complement genes have increased in number from 2 to >40 by several means. Current evidence suggests that the entire human genome has gone through two tetraploidisation events. The whole genome was twice duplicated, resulting in four copies of most chromosome pairs.15 Genes duplicated by this means are called ohnologues. Some whole regions have been duplicated on other chromosomes. For example, Lundin has mapped similarities across chromosomes 1, 11, 12, 15, and 19.16 Furthermore, several genes have undergone tandem duplications. Examples of this include CFB and C2, C4A and C4B, and CFH and the CFHR3, CFHR1, CFHR4, CFHR2, and CFHR5 genes. Genes duplicated in this way are called paralogues. Duplication of whole genomes or single genes allows for some redundancy of function by the presence of an unnecessary extra copy, which may undergo mutation and selection without adversely affecting the function of the other copy. Genes created by duplication are often found clustered together, particularly in cases of more recent duplication events. Close positional association may also be conserved because of shared genetic control elements.14
The shared sequences of duplicated, juxtaposed complement genes are responsible for the development at least one of the polymorphisms associated with disease risk by way of a process known as non-allelic homologous recombination. A sufficiently similar nearby sequence may allow a deletion to result if a new complementary strand of DNA becomes separated from the original strand and rejoins at the paralogous area, missing out part of the original sequence. For example, the terminal three exons of CFH share almost 100% sequence similarity with the terminal three exons in CFHR1. Deletion of CFHR3 and CFHR1, which are located between these segmental duplications, seems to have occurred by this mechanism.17 Deletion of CFHR1 and CFHR4 is also known, with the exclusion of a segment between two other highly similar regions.18 These deletions are not usually found de novo, but are inherited, having occurred in one or more ancestors. The question of what, if any, selective advantage deletions of these genes may confer is not yet answered. Altered risk of AMD is very unlikely to account for the evolutionary persistence of the deletion of CFHR3 and CFHR1, as the onset of AMD is usually long after child-bearing. The answer is more likely to be found in susceptibility to one or more infections.
Knowledge of how the complement system genes evolved is useful in the context of understanding the similar functions of many of the proteins. The regulators of complement activation (RCA) cluster on chromosome 1, for example, contains genes encoding related inhibitory proteins, including CFH, membrane cofactor protein (MCP), decay-accelerating factor, and C4-binding protein (C4BP).
The complement cascades
The basic structure of all the complement pathways can be summarised as follows: initiation, formation of a C3 convertase, cleavage of C3, formation of a C5 convertase, cleavage of C5, and formation of the membrane attack complex.19 On the simplest interpretation, each of the three complement pathways (alternative, lectin, and classical) is a unique method of activating the C3 molecule, initiating pro-inflammatory reactions, and activating the terminal complement pathway. Key to the amplification of the pathway is the fact that C3 is proteolytically activated to C3b due to C3 cleavage, and forms a new C3 convertase molecule that will cleave more C3. A positive feedback loop therefore follows.3 In some contexts, the activation of the terminal pathway is a vital end result (this may be the case for Neisserial infection, where host genetic deficiencies of the terminal pathway proteins increase susceptibility greatly).20 However, in many situations, the release of the anaphylatoxins C3a, C4a, and C5a or formation of the opsonins C3b and iC3b may be more important. At almost every step of all three pathways, there is cross-talk with proteins from other physiological pathways.5, 21
The alternative pathway is continuously activated by spontaneous hydrolysis of the internal thioester bond in C3 to form C3(H20) (Figure 1). This molecule, although not cleaved, can fulfil the same role as C3b in a C3 convertase.22 This process is known as ‘tick-over’ and occurs constantly at a low rate in biological fluids (0.005% per hour at 37 °C in vitro).22 C3(H20) binds to CFB, and this complex is cleaved by the complement serine protease factor D (CFD), resulting in splitting of the CFB protein into Ba and Bb fragments.23 C3(H20)Bb is a C3 convertase, which splits C3 into C3a and C3b. Once produced by these means, C3b perpetuates the positive feedback loop described above, joining CFB to form more convertase enzyme. Newly cleaved C3b is also deposited on nearby surfaces. C3(H20)Bb is a less efficient enzyme than C3bBb, but is less easily inactivated by CFH and its cofactor complement factor I (CFI).24 Because of the continuous activation of the alternative pathway in this manner, it requires inhibitory regulators to prevent inappropriate overactivation and damage to self-tissues.
The alternative pathway of the complement system and its regulators. The alternative pathway is central to the pathogenesis of AMD. Variants in complement component and regulator genes result in altered protein function and therefore altered dynamics of the system. The symbol  indicates decay-accelerating function (of convertase) or inactivation (of C3b to iC3b), + indicates enzymatic action, and – indicates inhibitory action.
indicates decay-accelerating function (of convertase) or inactivation (of C3b to iC3b), + indicates enzymatic action, and – indicates inhibitory action.
Initiation of the lectin pathway occurs when pattern recognition molecules (any of mannose-binding lectin (MBL), L-ficolin, H-ficolin, or M-ficolin) bind to the exterior surfaces of bacteria.25 These pattern recognition molecules bind to MBL-associated serine proteases (MASP-1, MASP-2, and MASP-3),5 which are closely related to C1r and C1s. The MBL–MASP-2 complex activates C4 and C2 by cleavage of thioester bonds, from which complement activation continues in the manner of the classical pathway, described below.
The classical pathway is initiated when C1q (a pattern recognition molecule) binds to immunoglobulin in immune complexes, to CRP on self or microbial surfaces or directly to molecules expressed on microbial membranes.5 This complex activates C1r, which becomes enzymatically active and cleaves C1s. C1s then activates C2 and C4, releasing C2a, C2b, C4a, and C4b. C2b and C4b become joined to form a C3 convertase, C4b2b. C2 is similar (and related) to CFB, while C4 is similar (and related) to C3. The C4b2b molecule is functionally similar to C3bBb.26
All three pathways result in production of C3b, and therefore regulators that act at this step of the cascade (such as CFH) are not only important in the alternative pathway, but influence all three pathways.
The activated pathways cause an increased plasma concentration of C3b molecules, leading to the incorporation of an extra C3b molecule into the C3 convertase. This results in production of C3bBb3b in all pathways in addition to C4b2b3b in the classical and lectin pathways, molecules that act as C5 convertases.27 The convertase cleaves C5 into C5a (an anaphylatoxin) and C5b, which forms part of the membrane attack complex. Unlike in the case of C3, the cleavage of C5 is not self-perpetuating.
C5b binds to C6, C7, C8, and C9 in sequence.28 C8 binds the incomplete membrane attack complex to cell surfaces, and up to 16 C9 molecules are then assembled on the membrane to generate a circular polymer, forming a pore that can perforate the membranes of non-encapsulated microbes. The membrane attack complex can also bind to self-membranes, stimulating release of growth factors from vascular endothelium.29
C3a, C4a, and C5a anaphylatoxins are released during these reactions. They are chemoattractants for phagocytic cells.30 C3a and C4a also exert anti-microbial action, and can damage bacterial and fungal cell membranes.31 The anaphylatoxins can bind to seven transmembrane spanning G-protein-coupled receptors, affecting smooth muscle contraction, degranulation of mast cells and neutrophils, and antigen presentation of dendritic cells.5 As well as playing a central role in the system, C3b, iC3b (inactivated C3b) and C4b are important opsonins. Phagocytes possess receptors (complement receptors CR1 to CR4 and CR1g) that bind to opsonins they encounter, attaching them to target cells.5
There are several receptors and regulatory proteins associated with the pathways (Figure 1). CFH is the main inhibitor of the alternative pathway, and has a particularly important role because of the constant activation of that pathway.32 The gene encoding CFH also encodes an alternatively transcribed truncated protein, factor H-like 1 (FHL1), which shares the first seven SCRs with CFH, and therefore some of the same enzymatic functions, but does not share a heparin-binding domain in SCRs 18–20, which binds CFH to cell membranes.33, 34 CFH, with CFI, inactivates C3b (forming iC3b), reducing formation of the C3 and C5 convertases in the fluid phase and on self surfaces. CFH-related 1 (CFHR1) is a complement inhibitor that inhibits C5 convertase activity and membrane attack complex assembly.35 MCP (or CD46) is smaller than CFH, containing only four SCRs in comparison to CFH's 20, but is closely related and fulfils the function of inactivation of C3b and C4b on self-surface membranes.36 C4BP fulfils the same function as CFH for the lectin and classical pathway convertases.19 These related genes are found clustered together in the RCA region at chromosome 1q31–32.
Evidence for complement activation in AMD
The drusen in AMD contain almost all alternative complement pathway proteins, including CFH, C3, and the products of its activation and degradation, and the terminal pathway proteins C5, C6, C7, C8, C9, separately and in combination as the membrane attack complex.37, 38 This indicates that there is local, complement-mediated inflammation in the retina in AMD. The products of complement activation in AMD can also be detected in blood. Scholl et al39 found that C3d, the Ba fragment of CFB and CFD were much increased in patients with AMD. The presence of activation products in circulating blood indicates that the inflammation found in AMD is not limited to the retina, but is systemic. To what extent systemic complement activation reflects pathological events in the eye, and to what extent systemic inflammation may be manifest in the eye is not yet clear.
Environmental exposure and complement
The effect of environmental exposures and behaviours on AMD risk may be at least partly mediated through the complement cascade. Smoking tobacco has consistently been found to increase risk of AMD. The Age-Related Eye Disease Study (AREDS) reported an odds ratio of 1.6 for neovascular AMD and 1.8 for geographic atrophy when comparing people who had smoked >10 pack-years to those who had not.40 The Beaver Dam Eye Study found that women who were current heavy smokers had a relative risk of 2.4 compared with female never-smokers.41 The Blue Mountains Eye Study found that current smokers had a relative risk of 3.9 of late AMD compared with non-smokers.42 Cigarette smoke has been shown to activate C3 in vitro and makes it less susceptible to inactivation by CFH and CFI.43 In vivo, tobacco activates the classical pathway.44
AREDS reported that high dietary intake of omega-3 long chain polyunsaturated fatty acids was associated with lower risk of neovascular AMD.45 The Blue Mountains Study supported these findings and furthermore observed that weekly consumption of fish and weekly consumption of nuts decreased risk of AMD, while high LDL cholesterol raised risk.46 HDL, which is protective against AMD, contains complement proteins, including clusterin, vitronectin, C3, C4A, C4B, and C9.47 HDL has been reported to inhibit binding of the terminal complement complex to cell surfaces, interfering with either the insertion of C9 into the membrane or its polymerisation.48 Halkes et al49 have shown that C3 levels increase in humans after eating fat.
Exercise was shown to reduce AMD risk in the Beaver Dam Study50 and by Williams in a cohort study of 33 000 people.51 Moderate exercise has anti-inflammatory effects mediated via the complement system: CFB is decreased, while protective CFH is increased.52
Genetic variants of complement genes associated with AMD and their functional effects
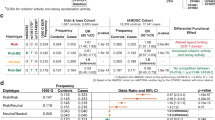
The CFH gene was the first of the complement genes found to be associated with AMD risk.53, 54, 55 The association was originally detected in small studies and has been replicated numerous times. CFH is mainly produced in the liver (though some is produced by endothelial cells) and is a major circulating protein.56 It binds to self-surfaces in order to protect them from activated complement. A coding single-nucleotide polymorphism (SNP), rs1061170 (Y402H), is responsible for an amino-acid change resulting in either a tyrosine (Y) or a histidine (H) molecule in the final protein. The variant is found in the ninth exon of the CFH gene, which encodes the seventh SCR module of the protein. This SCR is responsible for attaching CFH to glycosaminoglycan molecules on self-surfaces.33 Using donor eyes, Clark et al57 recently showed that the 402H (AMD high risk) variant of CFH found fewer binding sites in Bruch's membrane and choroidal blood vessels than the 402Y (low risk) variant. Interestingly, they showed that there was no difference in binding of the two variants to surfaces of the retinal pigment epithelium. They also found that CFH bound to heparan sulphate and dermatan sulphate on cell membranes: when these two molecules were removed, CFH did not bind to the cell surface. The authors suggested that tissue specificity of these CFH-binding molecules may be important in the pathology of the disease, and in separating the mechanism of AMD from other CFH-related diseases, such as renal dense-deposit disease.58 This study did not examine binding at the other GAG binding part of CFH, SCRs 18–20, which are duplicated in CFHR1. The risk variant 402H binds less well to monomeric CRP, an opsonin, resulting in less protection by CFH on necrotic or apoptotic self-tissues in the retina.59
A second coding variant in CFH, rs800292 (I62V), is also associated with AMD. This SNP is in the second exon of CFH and results in an isoleucine (I) amino acid instead of (more commonly) a valine (V) in the first SCR of the protein. The isoleucine variant is protective against AMD.60 The first four SCRs of CFH are the site for C3 convertase decay acceleration and for the inactivation of C3b (along with CFI).33 Tortajada et al61 reported that the I62 variant of CFH binds more strongly to C3b than the V62 variant, with the measured consequences that the C3 convertase, C3bBb, is formed less efficiently and C3b is decayed to iC3b more readily. The decay-accelerating function was not affected by the polymorphism.
A third polymorphism in this region with important functional effects is a deletion of the CFHR3 and CFHR1 genes which follow CFH and which are closely related to it. The deletion is associated with lower risk of AMD.17 CFHR3 and CFHR1 are expressed genes whose proteins have functions in the complement system that are independent of those of CFH, as shown in studies carried out in CFH-depleted serum.35, 62 CFHR1 protein inhibits the production of the C5 convertase (C3bBb3b).35 CFHR1 contains five SCRs, the final three of which share 98% identity to the final three SCRs of CFH, which form a heparin-binding domain.33 The function of CFHR3 has recently been characterised in an elegant series of experiments by Fritsche et al.62 The terminal two SCRs of CFHR3 are 96% similar to SCRs six and seven of CFH, which are functionally important, as described above.33 CFHR3 was found to act with CFI to degrade C3b in the absence of CFH.62 Like CFHR1, CFHR3 also inhibits the C5 convertase, a function that CFH does not share.62 The sequence identity between both of these CFHR proteins and CFH adds an additional level of complexity. Both CFHR1 and CFHR3 compete with CFH for binding on C3b, and the presence of either reduces CFH binding.62 The CFHR1 and CFHR3 molecules do not share with CFH the function of accelerating the decay of the C3 convertase. When they bind to a C3 convertase, at the expense of CFH, they prevent the decay of the C3 convertase. Additionally, even in the functions that they share with CFH, their efficiency as enzymes is usually less. Their concentration therefore affects the level of complement activation in a complex way.62 Deletion of the genes and therefore reduction (in heterozygosity) or absence (in homozygosity) of the CFHR3 and CFHR1 proteins means that there is a deficiency of these two anti-inflammatory molecules, but an increase in the activity of CFH, resulting in a lower inflammatory state, at least in the case of the retina in AMD. Fritsche et al62 showed that CFHR1 was more important in this regard than CFHR3. CFHR1 shares a heparin-binding site with CFH, and so in the case of reduction or absence of CFHR1, CFH can be expected to bind more abundantly to self-surfaces, as well as carrying out its inhibitory functions with less competition from the CFHR proteins.
Homozygous deletion of CFHR3 and CFHR1, protective against AMD, is an important risk factor for atypical haemolytic uraemic syndrome (aHUS).63 In aHUS, auto-antibodies to the terminal SCRs of CFH form and impair its binding and function, thereby leaving self-tissues underprotected against spontaneous complement activation.34, 64 Homozygous deletion of any gene of the classical complement system pathway is an important genetic cause of systemic lupus erythematosus, which is usually associated with the development of auto-antibodies to the missing protein.65 Some of those antibodies cross-react with other proteins, causing clinical effects. The presence of auto-antibodies to the C-terminus of CFH is very likely to be an example of the same phenomenon, with the antibodies to the absent CFHR1 molecule cross-reacting with the homologous region of CFH and interfering with its normal function. The deletion of CFHR3 and CFHR1, therefore, increases the efficiency of CFH protection against complement attack on host membranes, unless auto-antibodies form to the C-terminus. In this event, CFH function is disrupted, resulting in damage to host membranes, as is seen in aHUS. It seems, however, that auto-antibodies to CFH are not a significant cause of AMD. A recent study found that auto-antibodies to fragments of CFH were less common in AMD cases than in age-matched controls.66 This reflects the much lower frequency of the CFHR3 and CFHR1 deletion among AMD cases than controls, because of its protective effect. However, the paper further demonstrates the complexity of the interactions of antibodies with CFH, as it shows the presence auto-antibodies to parts of CFH other than the C-terminus, and that the incidence of these auto-antibodies increases with age.
Functional studies have uncovered the probable disease mechanisms underlying these associations and may lead to future treatments aimed at modifying the complement system in people affected by AMD and other diseases. Genetic studies suggest that there may be further variants in the region associated with altered risk that have not yet been uncovered.
CFB forms part of the alternative pathway C3 and C5 convertase enzymes. The minor allele of a CFB polymorphism, rs641153 (R32Q), with a glutamine (Q) substituted for an arginine (R) is associated with protection from AMD.67 The same codon is also affected by a SNP of the next base pair (rs12614, R32W), which results in a tryptophan (W) substitution. This polymorphic residue is in the Ba fragment of the CFB protein, which does not serve any known function after it is split from CFB by CFD.23 The polymorphisms are adjacent to the site of enzymatic cleavage and occur in a binding site for C3b during the formation of the proenzyme C3bB. Montes et al, using both human and then recombinant CFB, showed that the 32R variant of CFB binds to C3b with four times stronger affinity than the 32Q variant and formed proenzyme and enzyme more efficiently than the 32Q variant. The 32W variant formed enzyme at a rate intermediate between the 32R and 32Q variants.23 The protection against AMD associated with the 32Q variant of CFB is due to less efficient formation of the C3 convertase, a less efficient complement system being advantageous in this context. The findings of the Montes et al study would suggest that the 32W variant should also be protective, though with a less strong effect than 32Q. We have investigated this hypothesis in a case–control study and found that the 32W variant is protective, with an odds ratio intermediate between 32R and 32Q.68
Another variant of CFB, L9H, is independent of R32Q and was found to be associated with protection against AMD.67 The variant is in the signal peptide of the protein, which affects its travel through cytoplasm and is removed before export from the cell. Gold et al suggested that the peptide variant may alter secretion of CFB. There have been no functional studies of the effect this polymorphism published to date.
CFB is paralogous with complement component 2 (C2). These genes are located in tandem in the major histocompatibility complex class III region, a cluster of immune-related genes on chromosome 6p. Linkage disequilibrium spans several genes, which has made elucidation of the causative variants more difficult.69 In the first paper to report the association, the authors speculated that variants in CFB were the likely source of the association with AMD risk.67 The role of CFB in the alternative pathway and subsequent evidence of the functional effect of R32Q support this idea.23
CFI is a cofactor with CFH for the inactivation of C3b. Non-coding polymorphisms adjacent to the gene and in an intron of CFI are associated with altered risk of AMD.70 The associations have been independently replicated,71, 72 but no functional cause of the association has been found. Whether variants in the promoter region affect expression or whether undiscovered coding changes in the gene are responsible remains unknown.
Complement C3 is central to the complement cascade. The most strongly associated genetic variant in C3, rs2230199 (R80G) is a coding change resulting in the substitution of a glycine for an arginine.73 This polymorphism causes a change in the electrical charge of the C3 protein observed in the fast and slow electrophoretic variants of the protein. The amino-acid position is in the C3b part of the molecule, not in the anaphylatoxin C3a. The same variant appears to affect the outcome of renal transplantation.74 A 1972 study of the fast and slow variants found no difference in haemolytic activity.75 There have been no recent published functional studies of the effect of this polymorphism and so the mechanism of its effect in AMD is not yet known.
A candidate gene study of complement genes by Ennis et al76 showed an association between (mainly neovascular) AMD with SNPs in the promoter and introns of serpin peptidase inhibitor, clade G, member 1 (SERPING1), in both a UK and a USA cohort of patients. The authors showed that the gene, which is an inhibitor of complement component 1, is expressed in the retina.77 Subsequent studies using seven independent cohorts of AMD patients and controls failed to replicate the association with the gene variants.78 However, a further independent study of neovascular AMD and disease-free controls79 has recently replicated the original Ennis et al76 findings, noting that the previous replication groups were of mixed phenotype. The SERPING1 association remains controversial, but if it becomes established, this is the first indication that the classical complement pathway may be involved in AMD pathogenesis.
Several groups have searched for associations between other complement-related genes and risk of AMD, with some apparent associations being found, including C7 and mannose-binding protein.80 An association was found with a coding polymorphism in C5 but it failed to replicate and may have represented a type I error.81 At the present time, these cannot be considered to be established risk factors for AMD.
The major genetic risk factor for AMD is found on chromosome 10q, in the region of ARMS2 and HTRA1.82 The causative variant affecting risk is thought to be a polymorphism in the promoter of HTRA1 (high temperature requirement A-1 serine peptidase 1). The risk variant is associated with a twofold increase in expressed HTRA1 protein.83 This gene encodes a serine peptidase (like C1r, C1s, C2, and C4), which in vitro cleaves fibromodulin, clusterin, ADAM9, vitronectin, and α-2-macroglobulin.83 Fibromodulin is a structural protein associated with collagen and contains keratan sulphate chains.84 It binds to C1q and CFH on different binding sites (ie, without competition between the two molecules) and activates both the alternative and classical complement pathways, but usually does not result in membrane attack complex formation because of the inhibition by CFH.84 Clusterin and vitronectin are inhibitory complement proteins that prevent formation of the membrane attack complex.5 ADAM9 is a metallopeptidase, which acts as an enzyme to release a growth factor, pro-HB-EGF, which interacts with heparan sulphate, the CFH ligand.85 α-2-Macroglobulin is closely related to C3, C4, and C5, but interacts with the coagulation cascade as a protease inhibitor.86 The reason for the HTRA1 association has been elusive, but it seems now that progress is being made towards a better understanding of its involvement, and it may indeed be related to the complement system.
Treatments
No current treatments for AMD directly act on the complement system. There is no effective treatment for geographic atrophy, and treatment of neovascular AMD is currently based on monoclonal antibodies against vascular endothelial growth factor (anti-VEGF).87 This treatment aims to prevent growth of fragile new blood vessels.
The uncovering of the complement pathway as the major contributing pathway to AMD pathogenesis has resulted in interest in this system as a potential therapeutic target. Compstatin has been developed as a potential treatment that targets the complement system.21 It is a peptide that is administered by intra-vitreal injection, and which binds to C3 protein and inhibits binding of the C3 molecule to the C3b in the C3 convertase. The injected material forms a gel that slowly releases the molecule over many months, therefore dosing intervals could be relatively long. The drug has completed phase I trials. A small study in a primate model of AMD found improvement of drusen in four subjects.88
CFH is the major inhibitor of the complement system and has potential as a therapeutic agent if it could be supplemented or its production induced. A group in Germany has successfully transfected a moss with human CFH and produced a culture from which biologically active recombinant human CFH can be extracted.89
CFH and C3 are both central to the complement cascades, and successful enhancement of CFH function or inhibition of C3 activation will reduce downstream events, including anaphylatoxin, opsonin, and membrane attack complex production. Treatments intended to disrupt the complement cascade at the level of C5 are also being developed and may have useful applications in the treatment of AMD.90
The future of complement in AMD
AMD is a complex genetic disorder with important environmental risk factors.91 The risks associated with some variants of complement genes, however, are high, particularly in combination. Montes et al23 suggested the term ‘complotype’ to describe the combination of complement variants that may alter systemic complement activation and affect disease specificity. This is an idea that deserves further exploration, as it may be that the cumulative risk of combinations of risk variants is more complex than a simple combination of probabilities would suggest.
It is also important to note that although some variants are associated with reduced risk of AMD, where this is brought about by a reduction in the efficiency of the complement system, there may be an expensive trade-off. The 32Q variant of CFB, which decreases risk of AMD, increases susceptibility to some infections and to the complications of infections, for example, cardiomyopathy in Chagas disease.23 The CFH Y402H polymorphism associated with high AMD risk has recently been found to be protective against systemic inflammatory response syndrome in children.92 Deletion of CFHR3 and CFHR1 is protective against AMD,17 but homozygous deletion predisposes to the very serious systemic disease, aHUS, because of the formation of auto-antibodies that cross-react with CFH protein.63, 64 Given two alternative configurations of a protein, each may predispose to different pathologies, and with different effects in the heterozygous and homozygous states. We should recognise, therefore, that there often is no ‘model’ protein that functions perfectly and without any possible adverse consequences. Rather, there may be several variants that each carry different risks, the effects of which may be dependent on the context of other genetic, environmental or microbial exposures. Knowledge of these dichotomies will be important in the development of new treatments for AMD aimed at influencing the complement system.
Genome-wide association studies first revealed associations between AMD and variants in the complement genes. The nature of these studies is that they do not often directly reveal the causative variants of the genes, but may implicate markers in linkage disequilibrium with the true causative polymorphisms. Since the discovery of the gene associations, detailed genetic studies and careful functional studies have tried to unravel the means by which gene variants affect the protein interactions that lead to the disease. It is most likely that future treatments for AMD will depend on developing ways of enhancing or disrupting the complement system at appropriate stages to lessen harmful complement activation in the retina. The observation that lifestyle factors such as diet, exercise, and smoking alter complement activation provides an explanation for the efficacy of prevention measures, and an impetus to facilitate and encourage health improving behaviour.
The discovery of the genes involved in AMD was one of the first successes of the genome-wide association studies that followed the completion of the Human Genome Project. It seems likely that AMD will also be among the first diseases to benefit from treatments that have grown directly from that knowledge.
References
Klein R, Klein BE, Knudtson MD, Wong TY, Cotch MF, Liu K et al. Prevalence of age-related macular degeneration in 4 racial/ethnic groups in the multi-ethnic study of atherosclerosis. Ophthalmology 2006; 113: 373–380.
Pinto MR, Melillo D, Giacomelli S, Sfyroera G, Lambris JD . Ancient origin of the complement system: emerging invertebrate models. Adv Exp Med Biol 2007; 598: 372–388.
Zipfel PF . Complement and immune defense: from innate immunity to human diseases. Immunol Lett 2009; 126: 1–7.
Flierman R, Daha MR . The clearance of apoptotic cells by complement. Immunobiology 2007; 212: 363–370.
Ricklin D, Hajishengallis G, Yang K, Lambris JD . Complement: a key system for immune surveillance and homeostasis. Nat Immunol 2010; 11: 785–797.
Tolentino MJ, Miller S, Gaudio AR, Sandberg MA . Visual field deficits in early age-related macular degeneration. Vision Res 1994; 34: 409–413.
Ferris FL, Davis MD, Clemons TE, Lee LY, Chew EY, Lindblad AS et al. A simplified severity scale for age-related macular degeneration: AREDS Report No. 18. Arch Ophthalmol 2005; 123: 1570–1574.
de Jong PT . Age-related macular degeneration. N Engl J Med 2006; 355: 1474–1485.
Magnusson KP, Duan S, Sigurdsson H, Petursson H, Yang Z, Zhao Y et al. CFH Y402H confers similar risk of soft drusen and both forms of advanced AMD. PLoS Med 2006; 3: e5.
Nonaka M, Kimura A . Genomic view of the evolution of the complement system. Immunogenetics 2006; 58: 701–713.
Zipfel PF, Wurzner R, Skerka C . Complement evasion of pathogens: common strategies are shared by diverse organisms. Mol Immunol 2007; 44: 3850–3857.
Schneider MC, Prosser BE, Caesar JJ, Kugelberg E, Li S, Zhang Q et al. Neisseria meningitidis recruits factor H using protein mimicry of host carbohydrates. Nature 2009; 458: 890–893.
Jackson M . Human Genome Evolution. Routledge: New York, 2004.
Cooper D . Human Gene Evolution. BIOS Scientific Publishers: Oxford, 1999.
Larhammar D, Sundstrom G, Dreborg S, Daza DO, Larsson TA . Major genomic events and their consequences for vertebrate evolution and endocrinology. Ann N Y Acad Sci 2009; 1163: 201–208.
Lundin LG . Evolution of the vertebrate genome as reflected in paralogous chromosomal regions in man and the house mouse. Genomics 1993; 16: 1–19.
Hughes AE, Orr N, Esfandiary H, Diaz-Torres M, Goodship T, Chakravarthy U . A common CFH haplotype, with deletion of CFHR1 and CFHR3, is associated with lower risk of age-related macular degeneration. Nat Genet 2006; 38: 1173–1177.
Abarrategui-Garrido C, Martinez-Barricarte R, Lopez-Trascasa M, de Cordoba SR, Sanchez-Corral P . Characterization of complement factor H-related (CFHR) proteins in plasma reveals novel genetic variations of CFHR1 associated with atypical hemolytic uremic syndrome. Blood 2009; 114: 4261–4271.
Zipfel PF, Skerka C . Complement regulators and inhibitory proteins. Nat Rev Immunol 2009; 9: 729–740.
Brouwer MC, de Gans J, Heckenberg SG, Zwinderman AH, van der Poll T, van de Beek D . Host genetic susceptibility to pneumococcal and meningococcal disease: a systematic review and meta-analysis. Lancet Infect Dis 2009; 9: 31–44.
Ricklin D, Lambris JD . Compstatin: a complement inhibitor on its way to clinical application. Adv Exp Med Biol 2008; 632: 273–292.
Pangburn MK, Schreiber RD, Muller-Eberhard HJ . Formation of the initial C3 convertase of the alternative complement pathway. Acquisition of C3b-like activities by spontaneous hydrolysis of the putative thioester in native C3. J Exp Med 1981; 154: 856–867.
Montes T, Tortajada A, Morgan BP, Rodriguez de Cordoba S, Harris CL . Functional basis of protection against age-related macular degeneration conferred by a common polymorphism in complement factor B. Proc Natl Acad Sci USA 2009; 106: 4366–4371.
Bexborn F, Andersson PO, Chen H, Nilsson B, Ekdahl KN . The tick-over theory revisited: formation and regulation of the soluble alternative complement C3 convertase (C3(H2O)Bb). Mol Immunol 2008; 45: 2370–2379.
Thiel S . Complement activating soluble pattern recognition molecules with collagen-like regions, mannan-binding lectin, ficolins and associated proteins. Mol Immunol 2007; 44: 3875–3888.
Walport MJ . Complement. First of two parts. N Engl J Med 2001; 344: 1058–1066.
Pangburn MK, Rawal N . Structure and function of complement C5 convertase enzymes. Biochem Soc Trans 2002; 30: 1006–1010.
Morgan BP . Regulation of the complement membrane attack pathway. Crit Rev Immunol 1999; 19: 173–198.
Benzaquen LR, Nicholson-Weller A, Halperin JA . Terminal complement proteins C5b-9 release basic fibroblast growth factor and platelet-derived growth factor from endothelial cells. J Exp Med 1994; 179: 985–992.
Haas PJ, van Strijp J . Anaphylatoxins: their role in bacterial infection and inflammation. Immunol Res 2007; 37: 161–175.
Nordahl EA, Rydengard V, Nyberg P, Nitsche DP, Morgelin M, Malmsten M et al. Activation of the complement system generates antibacterial peptides. Proc Natl Acad Sci USA 2004; 101: 16879–16884.
Zipfel PF . Complement factor H: physiology and pathophysiology. Semin Thromb Hemost 2001; 27: 191–199.
Jozsi M, Zipfel PF . Factor H family proteins and human diseases. Trends Immunol 2008; 29: 380–387.
Jozsi M, Oppermann M, Lambris JD, Zipfel PF . The C-terminus of complement factor H is essential for host cell protection. Mol Immunol 2007; 44: 2697–2706.
Heinen S, Hartmann A, Lauer N, Wiehl U, Dahse HM, Schirmer S et al. Factor H-related protein 1 (CFHR-1) inhibits complement C5 convertase activity and terminal complex formation. Blood 2009; 114: 2439–2447.
Persson BD, Schmitz NB, Santiago C, Zocher G, Larvie M, Scheu U et al. Structure of the extracellular portion of CD46 provides insights into its interactions with complement proteins and pathogens. PLoS Pathog 2010; 6: e1001122.
Hageman GS, Luthert PJ, Victor Chong NH, Johnson LV, Anderson DH, Mullins RF . An integrated hypothesis that considers drusen as biomarkers of immune-mediated processes at the RPE-Bruch's membrane interface in aging and age-related macular degeneration. Prog Retin Eye Res 2001; 20: 705–732.
Anderson DH, Radeke MJ, Gallo NB, Chapin EA, Johnson PT, Curletti CR et al. The pivotal role of the complement system in aging and age-related macular degeneration: hypothesis re-visited. Prog Retin Eye Res 2010; 29: 95–112.
Scholl HP, Charbel Issa P, Walier M, Janzer S, Pollok-Kopp B, Borncke F et al. Systemic complement activation in age-related macular degeneration. PLoS One 2008; 3: e2593.
Clemons TE, Milton RC, Klein R, Seddon JM, Ferris III FL, Age-Related Eye Disease Study Research Group. Risk factors for the incidence of advanced age-related macular degeneration in the age-related eye disease study (AREDS) AREDS report no. 19. Ophthalmology 2005; 112: 533–539.
Klein R, Klein BE, Moss SE . Relation of smoking to the incidence of age-related maculopathy. The Beaver Dam Eye Study. Am J Epidemiol 1998; 147: 103–110.
Tan JS, Mitchell P, Kifley A, Flood V, Smith W, Wang JJ . Smoking and the long-term incidence of age-related macular degeneration: the Blue Mountains Eye Study. Arch Ophthalmol 2007; 125: 1089–1095.
Kew RR, Ghebrehiwet B, Janoff A . Cigarette smoke can activate the alternative pathway of complement in vitro by modifying the third component of complement. J Clin Invest 1985; 75: 1000–1007.
Koethe SM, Nelson KE, Becker CG . Activation of the classical pathway of complement by tobacco glycoprotein (TGP). J Immunol 1995; 155: 826–835.
SanGiovanni JP, Chew EY, Clemons TE, Davis MD, Ferris FL, Gensler GR et al. The relationship of dietary lipid intake and age-related macular degeneration in a case-control study: AREDS Report No. 20. Arch Ophthalmol 2007; 125: 671–679.
Tan JS, Wang JJ, Flood V, Mitchell P . Dietary fatty acids and the 10-year incidence of age-related macular degeneration: the Blue Mountains Eye Study. Arch Ophthalmol 2009; 127: 656–665.
Vaisar T, Pennathur S, Green PS, Gharib SA, Hoofnagle AN, Cheung MC et al. Shotgun proteomics implicates protease inhibition and complement activation in the antiinflammatory properties of HDL. J Clin Invest 2007; 117: 746–756.
Rosenfeld SI, Packman CH, Leddy JP . Inhibition of the lytic action of cell-bound terminal complement components by human high density lipoproteins and apoproteins. J Clin Invest 1983; 71: 795–808.
Halkes CJ, van Dijk H, de Jaegere PP, Plokker HW, van Der Helm Y, Erkelens DW et al. Postprandial increase of complement component 3 in normolipidemic patients with coronary artery disease: effects of expanded-dose simvastatin. Arterioscler Thromb Vasc Biol 2001; 21: 1526–1530.
Knudtson MD, Klein R, Klein BE . Physical activity and the 15-year cumulative incidence of age-related macular degeneration: the Beaver Dam Eye Study. Br J Ophthalmol 2006; 90: 1461–1463.
Williams PT . Prospective study of incident age-related macular degeneration in relation to vigorous physical activity during a 7-year follow-up. Invest Ophthalmol Vis Sci 2009; 50: 101–106.
Yang KD, Chang WC, Chuang H, Wang PW, Liu RT, Yeh SH . Increased complement factor H with decreased factor B determined by proteomic differential displays as a biomarker of tai chi chuan exercise. Clin Chem 2010; 56: 127–131.
Klein RJ, Zeiss C, Chew EY, Tsai JY, Sackler RS, Haynes C et al. Complement factor H polymorphism in age-related macular degeneration. Science 2005; 308: 385–389.
Haines JL, Hauser MA, Schmidt S, Scott WK, Olson LM, Gallins P et al. Complement factor H variant increases the risk of age-related macular degeneration. Science 2005; 308: 419–421.
Edwards AO, Ritter R, Abel KJ, Manning A, Panhuysen C, Farrer LA . Complement factor H polymorphism and age-related macular degeneration. Science 2005; 308: 421–424.
Friese MA, Hellwage J, Jokiranta TS, Meri S, Muller-Quernheim HJ, Peter HH et al. Different regulation of factor H and FHL-1/reconectin by inflammatory mediators and expression of the two proteins in rheumatoid arthritis (RA). Clin Exp Immunol 2000; 121: 406–415.
Clark SJ, Perveen R, Hakobyan S, Morgan BP, Sim RB, Bishop PN et al. Impaired binding of the age-related macular degeneration-associated complement factor H 402H allotype to Bruch's membrane in human retina. J Biol Chem 2010; 285: 30192–30202.
Clark SJ, Bishop PN, Day AJ . Complement factor H and age-related macular degeneration: the role of glycosaminoglycan recognition in disease pathology. Biochem Soc Trans 2010; 38: 1342–1348.
Skerka C, Lauer N, Weinberger AA, Keilhauer CN, Suhnel J, Smith R et al. Defective complement control of factor H (Y402H) and FHL-1 in age-related macular degeneration. Mol Immunol 2007; 44: 3398–3406.
Hageman GS, Anderson DH, Johnson LV, Hancox LS, Taiber AJ, Hardisty LI et al. A common haplotype in the complement regulatory gene factor H (HF1/CFH) predisposes individuals to age-related macular degeneration. Proc Natl Acad Sci USA 2005; 102: 7227–7232.
Tortajada A, Montes T, Martinez-Barricarte R, Morgan BP, Harris CL, de Cordoba SR . The disease-protective complement factor H allotypic variant Ile62 shows increased binding affinity for C3b and enhanced cofactor activity. Hum Mol Genet 2009; 18: 3452–3461.
Fritsche LG, Lauer N, Hartmann A, Stippa S, Keilhauer CN, Oppermann M et al. An imbalance of human complement regulatory proteins CFHR1, CFHR3 and factor H influences risk for age-related macular degeneration (AMD). Hum Mol Genet 2010; 19 (23): 4694–4704.
Zipfel PF, Edey M, Heinen S, Jozsi M, Richter H, Misselwitz J et al. Deletion of complement factor H-related genes CFHR1 and CFHR3 is associated with atypical hemolytic uremic syndrome. PLoS Genet 2007; 3: e41.
Jozsi M, Strobel S, Dahse HM, Liu WS, Hoyer PF, Oppermann M et al. Anti factor H autoantibodies block C-terminal recognition function of factor H in hemolytic uremic syndrome. Blood 2007; 110: 1516–1518.
Walport MJ . Complement and systemic lupus erythematosus. Arthritis Res 2002; 4: S279–S293.
Dhillon B, Wright AF, Tufail A, Pappworth I, Hayward C, Moore I et al. Complement factor H autoantibodies and age-related macular degeneration. Invest Ophthalmol Vis Sci 2010; 51: 5858–5863.
Gold B, Merriam JE, Zernant J, Hancox LS, Taiber AJ, Gehrs K et al. Variation in factor B (BF) and complement component 2 (C2) genes is associated with age-related macular degeneration. Nat Genet 2006; 38: 458–462.
Hughes AE, Mullan GM, Bradley DT . Complement factor B polymorphism 32W is protective against age related macular degenaration. Mol Vis (in press).
McKay GJ, Silvestri G, Patterson CC, Hogg RE, Chakravarthy U, Hughes AE . Further assessment of the complement component 2 and factor B region associated with age-related macular degeneration. Invest Ophthalmol Vis Sci 2009; 50: 533–539.
Fagerness JA, Maller JB, Neale BM, Reynolds RC, Daly MJ, Seddon JM . Variation near complement factor I is associated with risk of advanced AMD. Eur J Hum Genet 2009; 17: 100–104.
Ennis S, Gibson J, Cree AJ, Collins A, Lotery AJ . Support for the involvement of complement factor I in age-related macular degeneration. Eur J Hum Genet 2010; 18: 15–16.
Kondo N, Bessho H, Honda S, Negi A . Additional evidence to support the role of a common variant near the complement factor I gene in susceptibility to age-related macular degeneration. Eur J Hum Genet 2010; 18: 634–635.
Yates JR, Sepp T, Matharu BK, Khan JC, Thurlby DA, Shahid H et al. Complement C3 variant and the risk of age-related macular degeneration. N Engl J Med 2007; 357: 553–561.
Brown KM, Kondeatis E, Vaughan RW, Kon SP, Farmer CK, Taylor JD et al. Influence of donor C3 allotype on late renal-transplantation outcome. N Engl J Med 2006; 354: 2014–2023.
Colten HR, Alper CA . Hemolytic efficiencies of genetic variants of human C3. J Immunol 1972; 108: 1184–1187.
Ennis S, Jomary C, Mullins R, Cree A, Chen X, Macleod A et al. Association between the SERPING1 gene and age-related macular degeneration: a two-stage case-control study. Lancet 2008; 372: 1828–1834.
Mullins RF, Faidley EA, Daggett HT, Jomary C, Lotery AJ, Stone EM . Localization of complement 1 inhibitor (C1INH/SERPING1) in human eyes with age-related macular degeneration. Exp Eye Res 2009; 89: 767–773.
Allikmets R, Dean M, Hageman GS, Baird PN, Klaver CC, Bergen AA et al. The SERPING1 gene and age-related macular degeneration. Lancet 2009; 374: 875–876;author reply 876–877.
Lee AY, Kulkarni M, Fang AM, Edelstein S, Osborn MP, Brantley MA . The effect of genetic variants in SERPING1 on the risk of neovascular age-related macular degeneration. Br J Ophthalmol 2010; 94: 915–917.
Dinu V, Miller PL, Zhao H . Evidence for association between multiple complement pathway genes and AMD. Genet Epidemiol 2007; 31: 224–237.
Baas DC, Ho L, Ennis S, Merriam JE, Tanck MW, Uitterlinden AG et al. The complement component 5 gene and age-related macular degeneration. Ophthalmology 2010; 117: 500–511.
Jakobsdottir J, Conley YP, Weeks DE, Mah TS, Ferrell RE, Gorin MB . Susceptibility genes for age-related maculopathy on chromosome 10q26. Am J Hum Genet 2005; 77: 389–407.
An E, Sen S, Park SK, Gordish-Dressman H, Hathout Y . Identification of novel substrates for the serine protease HTRA1 in the human RPE secretome. Invest Ophthalmol Vis Sci 2010; 51: 3379–3386.
Sjoberg A, Onnerfjord P, Morgelin M, Heinegard D, Blom AM . The extracellular matrix and inflammation: fibromodulin activates the classical pathway of complement by directly binding C1q. J Biol Chem 2005; 280: 32301–32308.
White JM . ADAMs: modulators of cell-cell and cell-matrix interactions. Curr Opin Cell Biol 2003; 15: 598–606.
Cvirn G, Gallistl S, Koestenberger M, Kutschera J, Leschnik B, Muntean W . Alpha 2-macroglobulin enhances prothrombin activation and thrombin potential by inhibiting the anticoagulant protein C/protein S system in cord and adult plasma. Thromb Res 2002; 105: 433–439.
Bressler NM . Antiangiogenic approaches to age-related macular degeneration today. Ophthalmology 2009; 116: S15–S23.
Chi ZL, Yoshida T, Lambris JD, Iwata T . Suppression of drusen formation by compstatin, a peptide inhibitor of complement C3 activation, on cynomolgus monkey with early-onset macular degeneration. Adv Exp Med Biol 2010; 703: 127–135.
Buttner-Mainik A, Parsons J, Jerome H, Hartmann A, Lamer S, Schaaf A et al. Production of biologically active recombinant human factor H in Physcomitrella. Plant Biotechnol J 2010; e-pub ahead of print 17 August 2010; doi:10.1111/j.1467-7652.2010.00552.x.
Manthey HD, Woodruff TM, Taylor SM, Monk PN . Complement component 5a (C5a). Int J Biochem Cell Biol 2009; 41: 2114–2117.
Hughes AE, Orr N, Patterson C, Esfandiary H, Hogg R, McConnell V et al. Neovascular age-related macular degeneration risk based on CFH, LOC387715/HTRA1, and smoking. PLoS Med 2007; 4: e355.
Agbeko RS, Fidler KJ, Allen ML, Wilson P, Klein NJ, Peters MJ . Genetic variability in complement activation modulates the systemic inflammatory response syndrome in children. Pediatr Crit Care Med 2010; 11: 561–567.
Acknowledgements
DTB is funded by a research and development doctoral fellowship from the Public Health Agency (Northern Ireland).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Bradley, D., Zipfel, P. & Hughes, A. Complement in age-related macular degeneration: a focus on function. Eye 25, 683–693 (2011). https://doi.org/10.1038/eye.2011.37
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/eye.2011.37
Keywords
This article is cited by
-
Transcriptome profiling of Staphylococci-infected cow mammary gland parenchyma
BMC Veterinary Research (2017)
-
Human stem cell-derived retinal epithelial cells activate complement via collectin 11 in response to stress
Scientific Reports (2017)
-
Abrogation of immune complex glomerulonephritis by native carboxypeptidase and pharmacological antagonism of the C5a receptor
Cellular & Molecular Immunology (2016)
-
Prevalence of anti-retinal autoantibodies in different stages of Age-related macular degeneration
BMC Ophthalmology (2014)
-
Gelsolin and Ceruloplasmin as Potential Predictive Biomarkers for Cervical Cancer by 2D-DIGE Proteomics Analysis
Pathology & Oncology Research (2014)