Abstract
The dissection of S116 in more than 1500 individuals from Atlantic Europe and the Iberian Peninsula has provided important clues about the controversial evolutionary history of M269. First, the results do not point to an origin of M269 in the Franco–Cantabrian refuge, owing to the lack of sublineage diversity within M269, which supports the new theories proposing its origin in Eastern Europe. Second, S116 shows frequency peaks and spatial distribution that differ from those previously proposed, indicating an origin farther west, and it also shows a high frequency in the Atlantic coastline. Third, an outstanding frequency of the DF27 sublineage has been found in Iberia, with a restricted distribution pattern inside this peninsula and a frequency maximum in the area of the Franco–Cantabrian refuge. This entire panorama indicates an old arrival of M269 into Western Europe, because it has generated at least two episodes of expansion in the Franco–Cantabrian area. This study demonstrates the importance of continuing the dissection of the M269 lineage in different European populations because the discovery and study of new sublineages can adjust or even completely revise the theories about European peopling, as has been the case for the place of origin of M269.
Similar content being viewed by others
Introduction
The current genetic makeup of Europe is the result of many population migrations and settlements influenced principally by climate, cultural progress and the historical conquests of territory.1, 2 The genetic evidence provided by the analysis of the Y chromosome (Ychr), which is a valuable tool for the study of the evolution of the paternal lineages because of its uniparental mode of inheritance, has revealed that a large majority of the individuals currently in Central and Western Europe (40–90%) belong to a single lineage, R-M269.2, 3 The lineage M269 has its maximum frequency in the Franco–Cantabrian area, and it shows a cline of decreasing frequency with distance. This has led to numerous theories about the role of the Franco–Cantabrian region in European genetic history.
To date, the most widely accepted theories have argued that this pattern of frequencies may be the result of origin in, and subsequent postglacial expansion from, the Franco–Cantabrian refuge.2, 3, 4 Another theory, based on the variance of Y-STR haplotypes within M269, also supports its postglacial expansion but argues that M269 could have had a parallel expansion from a refuge in Eastern Europe (Anatolia).5 The new theory proposed by Balaresque et al,6 based on the higher diversity of Y-STR haplotypes in Eastern European M269 individuals than in Western European ones, concludes that there is a single origin for haplogroup M269 in Eastern Europe. In addition, the Balaresque et al6 theory shifts the origin from the glacial period to the Neolithic, because they apply germinal mutation rates rather than evolutionary, generating younger coalescence times.
The arrival of M269 from Eastern Europe proposed by Balaresque et al6 has been strongly refuted by Busby et al.7 Busby et al recalculated the diversity of Y-STRs haplotypes within M269 in a larger and geographically broader sample, indicating not higher diversity in Eastern Europe but a homogeneous background of microsatellite variation in the whole European sample.7
The dissection of haplogroup M269 has shown a wide range of European areas possessing geographically located subhaplogroup expansions,8, 9 which provides useful information for reconstructing the phylogeographic history of this lineage. However, the study of these sublineages, far from helping to find a consensus about the origin, growth and history of this great lineage, has increased the controversy. Myres et al9 analysed M269 and the sublineages M412, L11, U106, S116, U152 and M529. The obtained coalescence times and frequency distribution patterns led them to conclude that the current distribution of M269 sublineages is owing to allele surfing at the periphery of the westwards expansion of M269. Therefore, Myres et al9 proposed the origin of M269 in Eastern Europe, similar to Balaresque et al,6 but earlier during the Mesolithic period.
Finally, a different theory, not supported to date, argues that M269 entered the Iberian Peninsula in the late Neolithic and that its subhaplogroups S116 and M529 would appear during the expansion of the Bell Beakers northwards.10
This multi-sided debate affects not only European paternal lineages but also maternal lineages. In principle, the task of inferring the evolutionary histories of paternal lineages is actually more complicated than that of the maternal lineages, because the increased size and complexity of the Ychr makes the development of comprehensive and complete time-scaled phylogenetic trees more arduous than for mitochondrial DNA (mtDNA). In addition, mtDNA has more information in aDNA.
However, despite this, there is currently a major controversy about the origin and expansion of maternal haplogroup H, which shares a similar pattern of frequencies with paternal haplogroup R, and for which a similar and contemporaneous history has been suggested3 (Supplementary Box 1).
The controversy cannot be more interesting. Efforts to unravel the evolutionary history of the most frequent haplogroups in Europe have generated a cordial and productive discussion about new calculation methods and new approaches for the study of these haplogroups and sub-haplogroups.
Our study goes deeply into the study of the M269 sublineages of the European Atlantic coast and the Iberian Peninsula. This territory has a high frequency of the still-unresolved paragroup S116* (× U152, × M529) (data from7, 9). Therefore, this study offers the deepest analysis of haplogroup S116 made to date in Europe. These new data, as well as their comparison when possible with previous Ychr and mtDNA data, resolve important questions and offer novel clues about the evolutionary history of M269, in addition to finding new sublineages with important and restricted geographic locations.
Materials and methods
A total of 1560 healthy, unrelated males from the Iberian Peninsula (Galicia, Asturias, Cantabria, Basque Country, Barcelona, Alicante, Andalucía, Madrid, Portugal) and Atlantic Europe (Brittany (Brest), Ireland, Denmark) were studied (Supplementary Table S1). The Y-SNPs M269, L11, U106, S116, U152, M529, DF27, DF19 and L238 were analysed by TaqMan assays (Applied Biosystems, Carlsbad, CA, USA) or by High Resolution Melting Technology (for further details see Supplementary Box 2 and Supplementary Table S2). Individuals from Basque Country were also genotyped for a set of 17 Y-STR loci using the AmpFlSTR YfilerTM kit (Applied Biosystems).
Maps of haplogroup frequency distribution were constructed using the Surfer Golden software v 10.0.500 (Golden Software, Golden, CO, USA) by the kriging method. The spatial genetic patterns were studied through spatial principal component analyses (sPCAs) using the R software package adegenet (R Foundation for Statistical Computing, Vienna, Austria; http://adegenet.r-forge.r-project.org/). Genetic distances (Fst) between populations based on haplogroup frequencies were calculated using the Arlequin v 3.1 (University of Bern, Bern, Switzerland) software and plotted in Multidimensional Scaling graphs using the PAST software (University of Oslo, Oslo, Norway). The phylogenetic relationships of Y-STR haplotypes were estimated by median joining networks using NETWORK v 4.5.1.6 (Fluxus Technology Ltd., Kiel, Germany). Higher phylogenetic weight was allocated to the loci with lower mutation rate,11, 12 lower variance (VL, Kayser et al13) and higher linearity (D, Busby et al7; calculated with the actual range published in YHRD, Willuweit et al14; Supplementary Box 4). Coalescent times were estimated using the Network software and the evolutionary STR mutation rate 6.9 × 10−4/locus/25 years, established by Zhivotovsky et al15 and confirmed by Shi et al16 for the set of YSTRs analysed here. Further details about statistical treatment can be found in the Supplementary Box 2.
Data generated in this study can be accessed in Supplementary Tables S1. The Basque Y-STR–Y-SNP haplotype data have been uploaded to the public database YHRD under accession numbers YA003672-77, YA003718 and YA004063.14
Results and discussion
The Y-SNPs M269, L11, U106, S116, U152, M529, L238, DF19 and DF27 were analysed in 1560 individuals from 12 different populations from the Atlantic Coast and the Iberian Peninsula (Supplementary Tables S1).
Surprisingly, the inclusion of new populations from the Atlantic Coast and Iberia in this study has identified a frequency distribution of haplogroup S116 that differs from the previously proposed distribution. Myres et al9 proposed a frequency peak in the Upper Danube Basin and Paris, with declining frequency towards Italy, Iberia, southern France and British Isles. By contrast, these new data show maximum frequencies in northern Iberia, the western coast of France and the British Isles, raising questions about the possible expansion of this lineage during the early Neolithic LBK culture (Linearbandkeramik or Linear Pottery culture), as proposed by Myres et al.9
Supplementary Figure S1 shows distribution maps that compile all of the frequency data for M269 sublineages published to date (more than 16 000 male individuals;7, 9, 17 present study) but at a lower level of resolution than that achieved in the current study. From the maps, it can be appreciated that M269 sublineages show distinct areas of distribution in Europe: U106 is distributed in the countries of Central-Northern Europe, and S116 occurs in Western and South-western Europe. With regard to the sublineages of S116, U152 is more common in northern Italy and the Alpine region, whereas M529 is more common in the British Isles and Brittany. However, there is a large percentage of S116 individuals unassigned to any of these sublineages, described here as paragroup S116* (× U152, × M529). The frequency of this paragroup reaches approximately 50% in the Iberian Peninsula and exceeds 80% in the Basque region. It has also been observed in the area of Brittany and the British Isles, but the frequencies there do not exceed 20%.
The dissection analysis of S116 has provided very informative results for further completing the history of M269. The paragroup S116* (× U152, × M529) has been largely resolved owing to the discovery of the highly frequent sublineage DF27 in the Iberian Peninsula. DF27 has a frequency of 40–48% in Iberia but reaches frequencies over 60% in the Franco–Cantabrian region, particularly in the Basque population. However, outside the Iberian Peninsula, the frequency is below 20% (Supplementary Figure S2 and Supplementary Table S1). Thus, the sublineage S116-DF27 is located in a different geographic area than that occupied by the other S116 sublineages M529 and U152 (Supplementary Figure S1).
The DF19 and L238 sublineages show very low frequencies in Western Europe. The DF19 sublineage was not detected in any individuals, and L238 was detected only in one individual from Brest (Brittany) (Supplementary Table S1).
The new population data highlight the high frequencies of M529 found in Brest (>50%) (Supplementary Figure S1), outside the British Isles, which may raise doubts about whether it originated in the European continent or in the British Isles.
The sublineage U152 shows a striking distribution in the Iberian Peninsula (Supplementary Figure S1), where frequency peaks appear in the coastal corners in the SW (southern Portugal, 13%), NW (Galicia, Asturias, 8%) and NE (Barcelona, Alicante, 6%), and the minimum lies in the Basque region (2%). In Europe, haplogroup U152 has its maximum in the Alpine region, and thus perhaps its frequency pattern could be explained by a migration from the Alpine region of origin to the Iberian Peninsula, along the coast, avoiding areas historically known to have remained more isolated, as is the case with Basque Country.
After analysing all five sublineages so far known for S116 (U152, M529, L238, DF19 and DF27), some individuals remained who did not belong to any of these five sublineages, and they were assigned as belonging to the new, more concise S116* paragroup (× U152, × M529, × L238, × DF19, × DF27) (hereafter called S116*). The maximum frequency of S116* has been found in Irish (17%) and Basque (12%) populations. In both populations, the vast majority of individuals belonging to the S116 haplogroup belong to their respective M529 or DF27 sublineage, and those who do not belong to either of these sublineages belong almost entirely to paragroup S116* (Supplementary Table S1). Only the discovery of new Y-SNPs will determine whether these individuals can be assigned to new sublineages, which may be identical or different between Ireland and Basque country, providing more clues about the genetic relationship and evolution between the two populations.
One of the main reasons leading to the proposal of the hypothesis of origin and/or expansion of M269 from the Franco–Cantabrian refuge is its maximum frequency and pattern of decreasing frequency with increasing distance from this area. The Basque population is located in the heart of the refuge area, and our results indicate that almost all of their M269 lineages belong to sublineage S116 (Basque Country; M269–82%; S116-80%, Supplementary Table S1). If M269 had originated in this area, it would seem logical to find higher variability of M269 sublineages, such as M269xL11, L11 or U106*. Thus, the dissection of M269 in the refuge area raises questions about its origin in this region. Unfortunately, the homogeneity in the variability of Y-STRs within M269 makes it impossible to pinpoint a more likely origin,7 but the frequency distribution of M269 sublineages in the European continent suggests an origin in the East with a subsequent migration westwards, with the appearance of its sublineages during the advance of the migration wave.9
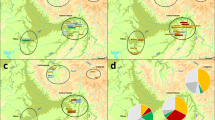
However, the Basque region has maximum frequencies of S116 and its sublineages S116* and DF27, the latter showing a decreasing gradient with distance. Meanwhile, M529 and U152 frequencies are extremely low. This may indicate that this region is a source for S116 and its sublineage DF27. Myres et al9 proposed the Upper Danube basin and Paris area as the geographic sources of S116. The patterns of frequencies obtained here also suggest that S116 emerged on the crest of the wave of migration but somewhere closer to the Franco–Cantabrian region. Thus, a possible evolutionary scenario of these lineages may be chronologically as shown in Figure 1: (1) origin of M269 in Eastern Europe; (2) origin of L11 on the wave of the westward advance of M269;9 and (3) colonization of the entire continent by L11, as evidenced by the high frequency of L11* in different parts of the Atlantic coast, from the Baltic to the southern coast of Portugal (data from7, 9) (L11 origin has been hypothesized in the map in Northern Europe); (4) origin of U106 from L11 individuals who inhabited the southern coast of the North Sea; (5) origin of S116 from L11 individuals inhabiting the Eastern Cantabrian coast, that is, the area of the Franco–Cantabrian refuge; and (6) origin of the DF27 sublineage from S116 individuals inhabiting the refuge area, while other S116 individuals spread to the rest of Iberia and Europe along the Atlantic and Mediterranean coasts, originating M529 and U152, respectively. Subsequently, (7) the U152, M529 and DF27 subtypes spread and came to occupy their present territories, with U152 and M529 re-entering the Iberian Peninsula (Figure 1). U152 and M529 may have re-entered the Iberian Peninsula during one of the numerous subsequent migrations to Iberia, during either Neolithic or historical times, that is, with the arrival of Phoenicians, Carthaginians, Romans, Goths or Vikings.18
To delve into the phylogenetic structure of S116 and DF27 haplogroups, a median joining network was performed with 15 Y-STR haplotypes of only M269 Basque native individuals (Supplementary Figure S3 and Supplementary Table S3). Thus, the study of the potentially ancient lineages that have inhabited the Franco–Cantabrian region until today is intended (Supplementary Box 3).
The phylogeny was constructed following carefully selected settings (Supplementary Box 4). The network showed a bipartite structure with two main groups corresponding to the individuals belonging to the S116* and DF27 haplogroups (Supplementary Figure S3). In addition, haplogroup DF27 appears to be split into two parts owing to the presence of two different haplotypes in the Y-STRs, DYS437/DYS448 (Supplementary Figure S3). Both Y-STRs have low mutation rates, and they are therefore more robust in distinguishing Y-chr haplogroups or established phylogenetic splits within haplogroups. DYS448, aside from being the Y-STR with lower mutation rate11, 12 and a small variance VL19 in the Basque population, has a long hexanucleotide repeat unit, which gives even higher phylogenetic weight.19 This may indicate the presence of different sub-haplogroups within DF27 in the Basque population, indicating that continuing the dissection of DF27 may contribute new information regarding the evolutionary history of this region.
Finally, a proper mutation rate was carefully selected for calculating TMRCAs, although the authors are aware of the lack of a definitive time scale for the Ychr; therefore, these calculations remain merely indicative. The classical mutation rate 6.9 × 10–4/locus/25 years, established initially by Zhivotovsky et al,15 was finally selected for being calibrated based on well-dated historical events and because its proper operation has been re-evaluated afterwards for the set of YSTRs analysed in this study.16 Concretely, Shi et al16 compared, in a very comprehensive study including a large panel of worldwide samples, the human male demographic inferences obtained with three different mutational rates: an observed mutation rate from the mutations counts in father–son pairs, the classical evolutionary mutation rate15 and a recalibrated evolutionary mutation rate (rEMR) corrected for the differences in variance of different sets of YSTRs. For the set of YSTRs analysed here, the evolutionary mutation rate and the rEMR were equivalent. Shi et al16 concluded that the rEMR provided the most comprehensive demographic inferences according to previous studies and actual geographical distributions.
The obtained coalescence times date the origin of haplogroup S116 in the native Basque region 11 673±1962 ybp, and the origin of DF27 soon after, 10 468±1831 ybp, which would place their origins after the last cold period of the Younger Dryas, that is, the early Holocene warm period, when weather conditions reached the current temperatures during the course of a few decades, encouraging population growth and expansion.3
These phylogenies and dates were confirmed also including non-native individuals in the network analysis, which allowed reaching identical conclusions (Supplementary Figure S4).
The spatial genetic patterns of the different haplogroups were deeply studied through sPCAs. Supplementary Figure S5 shows the sPCAs including the population data analysed here and data compiled from the literature (and therefore at the same low level of resolution as the distribution maps of Supplementary Figure S1). The analysis detected four spatial patterns that explain most of the variance related to M529, S116*, U106 and U152 (Supplementary Figure S6). By increasing the level of resolution of the sPCAs, including only the more resolved Western European data from this study, a new spatial pattern was detected for DF27 in Iberia (Supplementary Figures S7–S9). Interestingly, the analysis finds strong affinity among all Iberian populations, with the exception of the Basque population, which shows little affinity with the populations both outside and inside the peninsula but appears to participate in the distribution patterns affecting both of those populations. This may indicate that the Basque country has been involved in the history of the different haplogroups that principally characterize both Western European regions (M529 and DF27), and this would support the previously proposed scenario (Figure 1), in which, first, S116 expands outside the refuge and, second, U152 and M529 originate outside the peninsula and DF27 inside.
MDS representation of Fst genetic distances between populations, calculated based on haplogroup frequencies, shows results consistent with those obtained in the sPCAs (Supplementary Figure S10,Supplementary Table S4).
In summary, this study provides new genetic evidence indicating the absence of diversity of M269 lineages over S116 in the current population of what once was the refuge, the maximum frequencies of S116, S116* and DF27 in the refuge area and their spatial distributions in Iberia and Western European coast. This is in addition to the evidence from previous studies: the homogeneity in Y-STR diversity within M269 in Europe7 and the emergence of new sublineages such as L11 on the wave of the advance of M269 into Western Europe9 consistent with the scenario proposed in Figure 1.
This scenario proposes an origin in the East for M269, in contrast to the classical theories.2, 3 The controversy in calculating TMRCAs makes it impossible to reliably date these evolutionary episodes, at least until the more complete Ychr allows more accurate time scales and/or until genotyped and firmly dated archaeological remains become available.
However, the authors believe that it is unlikely that an arrival to Europe of M269 during the Neolithic period has generated such a complex scenario of expansions for its sublineages, especially when genetic evidence of cultural diffusion has been found for Ychr in Anatolia20, 21 and for mtDNA in the refuge.22 Thus, the spread of Neolithic culture would mean a lower demic movement. The theories that argue for an origin in the East and during the Neolithic period assume a rapid expansion of M269 throughout Europe, replacing most of the previously settled haplogroups, which would be compatible with a main scenario of demic diffusion.
The scenario proposed here would be most compatible with an arrival of M269 from the East occurring in Palaeolithic times. The Wurm glaciation had numerous ups and downs in temperature that would have led to the existence of multiple glacial refugia, which has been proposed both for mtDNA and Ychr.5, 23 Improved weather conditions would allow colonization of more northern territories from all refuges simultaneously. Similarly, the mtDNA-H and Ychr-R lineages that evolved in the East from Palaeolithic times, could have expanded westwards during the Neolithic period, thereby mixing with other H and R lineages that arrived to Western Europe in Paleolithic times and evolved independently in these western territories. This may be one reason for the complexity of interpreting the results, in addition to the assumption that post-Neolithic movements may be masking and confounding the oldest traces.
In this context, the genetic evidence found for the sister haplogroup of M269 in the maternal line, haplogroup H, has been helpful for complementing and giving clues about M269 history (Supplementary Box 5).
In sum, this study demonstrates the importance of continuing the dissection of the M269 lineage in different European populations because the discovery and study of new sublineages can adjust or even completely rewrite the theories about European peopling, as has been the case with the place of origin of M269. Similarly, the future availability of complete sequences of the Ychr and of desirable Palaeolithic aDNA data may definitively reveal the complete and true history of this major lineage.
References
Pinhasi R, Thomas MG, Hofreiter M, Currat M, Burger J : The genetic history of Europeans. Trends Genet 2012; 28: 496–505.
Semino O, Passarino G, Oefner PJ et al: The genetic legacy of Paleolithic Homo sapiens sapiens in extant Europeans: a Y chromosome perspective. Science 2000; 290: 1155–1159.
Soares P, Achilli A, Semino O et al: The archaeogenetics of Europe. Curr Biol 2010; 4: R174–R183.
Cardoso S, Valverde L, Alfonso-Sánchez MA et al: The expanded mtDNA phylogeny of the Franco-Cantabrian region upholds the pre-neolithic genetic substrate of Basques. PLoS One 2013; 8: e67835.
Cinnioğlu C, King R, Kivisild T et al: Excavating Y chromosome haplotype strata in Anatolia. Hum Genet 2004; 114: 127–148.
Balaresque P, Bowden GR, Adams SM et al: A predominantly neolithic origin for European paternal lineages. PLoS Biol 2010; 8: e1000285.
Busby GB, Brisighelli F, Sánchez-Diz P et al: The peopling of Europe and the cautionary tale of Y chromosome lineage R-M269. Proc Biol Sci 2012; 279: 884–892.
Cruciani F, Trombetta B, Antonelli C et al: Strong intra- and inter-continental differentiation revealed by Y chromosome SNPs M269, U106 and U152. Forensic Sci Int Genet 2011; 5: e49–e52.
Myres NM, Rootsi S, Lin AA et al: A major Y-chromosome haplogroup R1b Holocene era founder effect in Central and Western Europe. Eur J Hum Genet 2011; 19: 95–101.
Klyosov A : Ancient history of the Arbins, bearers of haplogroup R1b, from central Asia to Europe, 16,000 to 1500 years before present. Advances in Anthropology 2012; 2: 87–105.
Goedbloed M, Vermeulen M, Fang RN et al: Comprehensive mutation analysis of 17 Y-chromosomal short tandem repeat polymorphisms included in the AmpFlSTR Yfiler PCR amplification kit. Int J Legal Med 2009; 123: 471–482.
Ballantyne KN, Goedbloed M, Fang O et al: Mutability of Y-chromosomal microsatellites: rates, characteristics, molecular bases, and forensic implications. Am J Hum Genet 2010; 87: 341–353.
Kayser M, Krawczak M, Excoffier L et al: An extensive analysis of Y-chromosomal microsatellite haplotypes in globally dispersed human populations. Am J Hum Genet 2001; 68: 990–1018.
Willuweit S, Roewer L : International Forensic Y Chromosome User Group: Y chromosome haplotype reference database (YHRD): update. Forensic Sci Int Genet 2007; 1: 83–87.
Zhivotovsky LA, Underhill PA, Cinnioğlu C et al: The effective mutation rate at Y chromosome short tandem repeats, with application to human population-divergence time. Am J Hum Genet 2004; 74: 50–61.
Shi W, Ayub Q, Vermeulen M et al: A worldwide survey of human male demographic history based on Y-SNP and Y-STR data from the HGDP-CEPH populations. Mol Biol Evol 2010; 27: 385–393.
Larmuseau MH, Vanderheyden N, Jacobs M, Coomans M, Larno L, Decorte R : Micro-geographic distribution of Y-chromosomal variation in the Central-Western European region Brabant. Forensic Sci Int Genet 2011; 5: 95–99.
Encarta Online Encyclopedia. Spain 2007, http://www.webcitation.org/5kwqnGivb.
Järve M, Zhivotovsky LA, Rootsi S et al: Decreased rate of evolution in Y chromosome STR loci of increased size of the repeat unit. PLoS One 2009; 4: e7276.
Mirabal S, Varljen T, Gayden T et al: Human Y-chromosome short tandem repeats: a tale of acculturation and migrations as mechanisms for the diffusion of agriculture in the Balkan Peninsula. Am J Phys Anthropol 2010; 142: 380–390.
Morelli L, Contu D, Santoni F, Whalen MB, Francalacci P, Cucca F : A comparison of Y-chromosome variation in Sardinia and Anatolia is more consistent with cultural rather than demic diffusion of agriculture. PLoS One 2010; 5: e10419.
Hervella M, Izagirre N, Alonso S et al: Ancient DNA from hunter-gatherer and farmer groups from northern Spain supports a random dispersion model for the Neolithic expansion into Europe. PLoS One 2012; 7: e34417.
Pala M, Olivieri A, Achilli A et al: Mitochondrial DNA signals of late glacial recolonization of Europe from near eastern refugia. Am J Hum Genet 2012; 90: 915–924.
Acknowledgements
Funds were provided by the Basque Government (IT-424-07). LV received a postdoctoral grant from the Basque Government (DKR-2012-439), applied through the Ikerbasque Foundation. The authors are deeply indebted to Emmanuelle Génin, Dan Bradley, Kevin Kenna, the Basque Foundation of Science (BIOEF), the Spanish National DNA Bank, SGIker (UPV/EHU, MICINN, GV/EJ, ERDF and ESF) and to all the people who voluntarily participated in this study.
Author contributions
LV and MMP conceived the study. JA, SC, KR, CF, OH, SJ, MFP, BMJ, JO, NM and MMP contributed samples/materials. LV, MJI, PV, AMG, AG and JO analysed the samples. LV analysed the data. LV and MMP wrote the manuscript. All co-authors reviewed the manuscript before submission.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
JA and SC are employed by the commercial company Progenika Biopharma SA (Grifols, Derio, Spain). This does not alter the authors’ adherence to all the policies on sharing data and materials. The remaining authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Valverde, L., Illescas, M., Villaescusa, P. et al. New clues to the evolutionary history of the main European paternal lineage M269: dissection of the Y-SNP S116 in Atlantic Europe and Iberia. Eur J Hum Genet 24, 437–441 (2016). https://doi.org/10.1038/ejhg.2015.114
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2015.114
This article is cited by
-
Y-chromosome target enrichment reveals rapid expansion of haplogroup R1b-DF27 in Iberia during the Bronze Age transition
Scientific Reports (2022)
-
The Y chromosome of autochthonous Basque populations and the Bronze Age replacement
Scientific Reports (2021)
-
Phylogeographic review of Y chromosome haplogroups in Europe
International Journal of Legal Medicine (2021)
-
The Marquesans at the fringes of the Austronesian expansion
European Journal of Human Genetics (2019)
-
The Beaker phenomenon and the genomic transformation of northwest Europe
Nature (2018)