Abstract
Up to 25% of colorectal cancer (CRC) may be caused by inherited genetic variants that have yet to be identified. Previous genome-wide linkage studies (GWLSs) have identified a new loci postulated to contain novel CRC risk genes amongst affected families carrying no identifiable mutations in any of the known susceptibility genes for familial CRC syndromes. To undertake a new GWLS, we recruited members from 54 non-syndromic families from Australia and Spain where at least two first-degree relatives were affected by CRC. We used single-nucleotide polymorphism arrays to genotype 98 concordant affected relative pairs that were informative for linkage analyses. We tested for genome-wide significance (GWS) for linkage to CRC using a quantile statistic method, and we found that GWS was achieved at the 5% level. Independently, using the PSEUDO gene-dropping algorithm, we also found that GWS for linkage to CRC was achieved (P=0.02). Merlin non-parametric linkage analysis revealed significant linkage to CRC for chromosomal region 10p15.3–p15.1 and suggestive linkage to CRC for regions on 14q and 9q. The 10p15.3–p15.1 has not been reported to be linked to hereditary CRC in previous linkage studies, but this region does harbour the Kruppel-like factor 6 (KLF6) gene that is known to be altered in common CRC. Further studies aimed at localising the responsible genes, and characterising their function will give insight into the factors responsible for susceptibility in such families, and perhaps shed further light on the mechanisms of CRC development.
Similar content being viewed by others
Introduction
There is a good understanding of the molecular genetics underlying the major familial colorectal cancer (CRC) syndromes: Lynch syndrome or hereditary non-polyposis CRC (HNPCC), familial adenomatous polyposis (FAP) and MYH-associated polyposis (MAP). However, these represent only ∼5% of CRC cases in the community.1, 2 Of the remainder, there is a clear indication of inheritance suggesting that other predisposing genes are involved in up to 25% of all cases.3 The Amsterdam criteria4 are a set of diagnostic criteria used to help identify families that are likely to have Lynch syndrome. Families meeting these criteria, but without evidence of mismatch repair (MMR) deficiency (eg, normal immunohistochemical staining for the MMR genes and/or microsatellite stable cancers) and in the absence of any other identifiable mutations in any of the known familial CRC risk genes, have been termed familial CRC type X5 or syndrome X6 families or more recently7 as hereditary non-syndromic CRC families. Various genome-wide linkage studies (GWLSs) have been undertaken to identify the underlying causative gene variants in these and similar families. They have resulted in a number of different regions of linkage being reported,7, 8, 9, 10, 11, 12, 13, 14, 15 with regions on 3q7, 10, 12 and 9q8, 11, 15 being independently identified by different laboratories.
It was the aim of our study to undertake a new genome-wide linkage study (GWLS) to investigate what region of the genome was likely to contain genes conferring increased risk of CRC in a new set of families from Australia and Spain.
Materials and methods
Ascertainment of families and exclusion of known syndromes
We restricted our study to non-syndromic, high-risk CRC families, defined as those containing at least one affected person who has one or more first-degree affected relative(s) and where the known causal mutations had been excluded. This study group was enriched for familial CRC type X families but also included other high-risk CRC families. Volunteers from 54 such families were enrolled in our study by ascertaining the families with the following characteristics in order of priority:
-
Preference 1: Patients from families containing at least one affected person who have had two or more first-degree affected relatives, and at least one affected person diagnosed before reaching 50 years of age (28 families).
-
Preference 2: Patients from families containing at least one affected person who have had one or more first-degree affected relatives (26 families).
Note that Preference 1 is identical to the Amsterdam I or familial CRC type X criteria, except for the requirement of multiple generations in the latter.
Affected status was defined as diagnosis with either colorectal carcinoma (CA) or advanced adenoma (AA), where AA was defined as three or more synchronous or metachronous adenomas and/or adenoma(s) with villous morphology, and/or with severe dysplasia, and/or diameter ≥10 mm. Diagnoses were confirmed by pathology reports. The study was reviewed and approved by the Human Research Ethics Committees of the three participating centres: Flinders Medical Centre, Adelaide (25 families), The Royal Melbourne Hospital, Melbourne (21 families), and Institut Català d’Oncologia, Barcelona (8 families), and informed consent was obtained from all participants. Families with known colon cancer syndromes including HNPCC or Lynch syndrome, FAP, hereditary mixed polyposis, juvenile polyposis, Peutz–Jeghers, Cowden's syndrome or MYH-associated polyposis were excluded by review of medical records.
Persons for whom tumours were available underwent microsatellite instability (MSI) testing of BAT-25 and BAT-26, and/or immunohistochemistry (IHC) of hMLHI, hMSH2, hMSH6 and hPMS2, and, if positive, their family was excluded from the study. In 48 families, one tumour was available and in four families two tumours were available. All tested negative, and therefore these 52 families were included in the study. Two additional families satisfied the requirements for Preference 2, but did not have MSI or IHC data available. These were also judged to be unlikely to be carrying any of the known mismatch repair gene mutations, as (1) no family member was clinically diagnosed with CRC earlier than 60 years, and (2) identity-by-descent (IBD) sharing analysis, conducted on derived single-nucleotide polymorphism (SNP) data for each of the 2 Mb regions centred on hMLHI, hMSH2, hMSH6 and hPMS2, indicated no inherited contribution. These two families were therefore included in the study to give a total of 54 families.
Demographics
Summary statistics of families that met our selection criteria were:
-
27 familial CRC type X families;
-
1 family meeting Preference 1, but failing to meet familial CRC type X requirements, as all affected individuals were in a single generation;
-
14 meeting Preference 1, except that all cases were diagnosed at an age ≥50 years;
-
6 meeting Preference 2 and one reported family individual with an age of diagnosis <50 years;
-
6 meeting Preference 2 with minimum age of diagnosis ≥50 years.
Genotypes of 132 individuals from these 54 families were used in the analyses. Of these, 98 were from individuals with a diagnosis of CA with or without adenoma and 34 from individuals with AA but without CA. The number of available affected persons per family ranged from 2 to 11 and the number of affected persons per family with DNA available ranged from 2 to 4. Of the 54 families, 16 were reported to contain affected persons in three or more generations, whereas 38 were reported with affected family members in only one or two generations.
As summarised in Table 1, the median age at diagnosis of CRC in the families was 59 years, significantly less than the median ages of diagnosis of 70 years for men and 71 years for women observed in both the general Australian16 and Spanish populations17 (χ2=25.8, for females; χ2=21.4 for males; both on 1 d.f. and having P<10−6). The median age at diagnosis of colorectal adenomas was 56 years.
Families used in our linkage analyses
The Merlin non-parametric linkage (NPL) analyses used complete pedigree information from all 54 families, as the Sall statistic weights pedigrees containing more than two affected members. Concordant affected pairs used in the analyses are shown in Table 1.
Genotyping with the 50K mapping array
Blood samples obtained from consenting family members and DNAs were extracted and genotyped using the GeneChip Human Mapping 50K Array Xba 240 assay (the 50K Array) (Affymetrix Inc., Santa Clara, CA, USA) as described in Supplementary data, to deliver non-redundant genotype data from 305 individuals. Annotation of SNPs was conducted as described in Supplementary data. Also, as described in Supplementary data, pedigree checking was undertaken to account for the relatedness of the siblings and adjustments were made to the data to take account of genotyping errors and linkage disequilibrium between SNPs.
Genome-wide significance
Genome-wide significance (GWS) of the linkage with CRC was tested by two independent methods: the quantile statistic (QS) described in Saunders et al18 and the gene-dropping algorithm in the PSEUDO package.19 The QS method is based on a summary statistic of the set of likelihood ratio (LR) statistics computed for each SNP for sibling pairs.18 The 5% summary statistic was used here. Its GWS was determined from the results of Saunders et al.18 IBD sharing was estimated for use in the QS method using an implementation in the R package of the forward/backward algorithm20 with correction for 0.2% genotyping error rate (as found in Saunders et al21).
Merlin22 was used to generate NPL scores using the Kong and Cox23 linear model with the Whittemore and Halpern24 Sall sharing statistic. Empirical GWS levels for the NPL LOD scores were calculated using PSEUDO 0.3.5 (PSEUDO)19 generated from a pool of 100 random gene-dropping replicates.
The mode of action of the gene is unknown so methods not heavily reliant on a specific genetic model were used. As shown in Saunders et al,18 the alternative model used to calculate the LR statistic for the GWLS test does not have a large effect on the power of the QS statistics. The calculations here used a dominant model with allele frequency 0.3 and penetrance 0.18 for carriers and 0.017 for non-carriers, which was found to perform well across a range of alternative models.18 For locating the gene, a NPL approach was used, which is again insensitive to the exact inheritance model.
Early onset families
The GWS of the statistic was determined for data obtained from 42 families that contained suitable sibling pairs informative for linkage. It would be expected that an individual carrying an inherited susceptibility allele would have an earlier age of onset than usual for the population. Therefore, additional analyses were carried out on three restricted data sets that included only families reporting a case of CA or AA diagnosed either before age 55 years or 50 years or 45 years. However, each of these age restrictions reduced the number of individuals available for analyses compared to the full data set and GWS was not achieved for any of these three additional data sets (data not shown).
Results
Establishing GWS
The GWS of linkage with CRC was determined for data (Table 1) obtained from 54 families that contained suitable affected pairs, informative for linkage. Two independent methods were used: the QS,18 that is specific for families containing sibling pairs and the gene-dropping algorithm in the PSEUDO package,19 that uses all relative pair data. For the subset of 42 families that contain concordant affected sibling pairs, the 5% QS18 reached the 5% level of GWS. Supplementary Figure 1 shows the set of LR statistics for all SNPs used in the analysis. We also assessed GWS for subsets of earlier onset families (see below). However, these groups proved to be too small to allow detection of linkage. For the full set of 54 families, GWS of linkage was also assessed by the Merlin method22 of PSEUDO gene dropping and empirical P-values were computed. As summarised in Table 2, a genome-wide threshold of significance (P=0.02) was achieved for the highest non-parametric linkage (NPL) score of 3.45 which occurred at cytoband 10p15.3–10p15.1.
Locating chromosomal regions linked to increased CRC risk
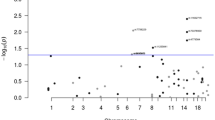
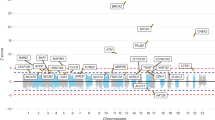
For the genome-wide non-parametric analysis executed in the Merlin package, the results for all 54 families containing two or more genotyped affected members are shown in Figure 1. As summarised in Table 2, the strongest signal was located on 10p15.3–p15.1 (maxNPL=3.45) for the full set of families. Although this did not quite satisfy a proposed generalised definition for significant linkage (maximal Sall linear model nonparametric LOD scores (maxNPL)>3.62),25 we found significant linkage using an empirical gene-dropping approach. Using a 1-LOD support interval approach, we estimated the size of this region of linkage as 2 Mb and to be bounded by SNPs rs10489254 to rs10494827. Using the same approach, regions of suggestive linkage ((maxNPL)>2.2)25 were observed for a 6.5 Mb region of chromosome 14q24.3–q31.1 (maxNPL=2.28) and for a 12 Mb region of 9q33.3–q34.3 (maxNPL=2.24). These three regions represent the most promising loci for further study. Regions of weaker linkage were located across a 19 Mb region on 10q22.3–q24.1 (maxNPL=2.14), across 8 Mb on 2p25.3–p25.1 (maxNPL=2.04) and across 21 Mb on 1q25.2–q32.1 (maxNPL=1.71).
Merlin linkage analysis using data from all families containing two or more affected members: plots of the likelihood of SNPs on chromosomes being linked to CRC. Locations for genes known to be causative for known syndromes are shown as vertical lines. The dotted vertical lines indicate the locations of genes known or suspected to be causative for established familial CRC syndromes or from other GWLS or GWAS: (1) EPHB2,34 (2) MUTYH,35 (3) MSH2,36 (4) MSH6,36 (5) MLH1,36 (6) APC,37 (7) PMS2,36 (8) rs7014346/TCF4-binding site,38, 39 (9) TGFBR1,40, 41 (10) BMPR1A,42 (11) PTEN,43 (12) CRAC1,44 (13) HIC1,9 (14) TP53,45, 46, 47 (15) SMAD7,48 (16) SMAD449 and (17) STK11.50
Sensitivity analysis
To determine whether geographical differences in founder effect may be contributing to our analyses, we undertook a sensitivity analysis by separately analysing the samples from Spain and Australia. It was found that the signals in the data from the two countries were consistent and jointly contributed to the overall significance of the linkage results (data not shown).
Discussion
In the present GWLS, using 54 Australian and Spanish non-syndromic CRC families, we identified promising regions for further study on chromosomes 1q, 2p, 9q, 10p, 10q and 14q. We also observe a minor linkage peak at 9q22 (Figure 1).
The chromosomal region of linkage most strongly implicated to be harbouring a novel susceptibility gene for CRC was 10p15.3–p15.1. Although this region has not previously been implicated as a region of increased risk for hereditary CRC, an examination of current databases provides some evidence for such a likelihood. This signal on 10p15 does lie in a region that has previously been associated26 with common or sporadic CRC. Interrogation of the RefSeq database27 revealed three protein coding genes map to the 2 Mb region of 10p15.3–p15.1; PFKP, PITRM1 and Kruppel-like factor 6 (KLF6). Of these, KLF6 is a plausible candidate gene for increased risk of familial CRC. KLFs are key transcription factors,28 and while there is debate26, 29 about the frequency of mutation activation in common CRC, there is good evidence that LOH of KLF6 is a feature of common CRC.29 Interestingly, these inactivation events are rarely observed in HNPCC or FAP,26, 29 two hereditary syndromes that we excluded from our study.
There is also biological plausibility for the 14q and 9q regions to be harbouring new risk genes for hereditary CRC. In particular, it is interesting that the MMR gene, MLH3, is among the 84 protein coding genes that map to the 6.5 Mb region of 14q24.3–q31.1 (Supplementary Table 1) and is among the nine genes in this region that have been annotated in the Genecards database30 as being altered in CRC (Supplementary Table 2). We also note that of the 152 protein coding genes that map to the 12 Mb region of 9q32–q34.13 (Supplementary Table 3), 22 have been annotated in the Genecards database,30 as being altered in CRC (Supplementary Table 4) and that this includes three key enzymes in the prostaglandin biosynthesis pathway, COX-1, PTGES and PTGES2. Recently, increased CRC risk was reported to be associated with a silent mutation in COX-1.31 Increased expression of PTGES2 and PTGES, that function downstream of COX-2, has also been correlated with prognosis in CRC patients.32
It is likely that large scale DNA sequencing of these regions will be needed to determine the identity of the underlying causal variants for CRC in our non-syndromic CRC families.
Importantly, in this particular group of patients where we excluded families that fitted into the known syndromes, we found no linkage peaks near the genes known to cause familial CRC syndromes including FAP, Lynch syndrome or HNPCC, hereditary mixed polyposis, juvenile polyposis, Peutz–Jeghers, Cowden's syndrome or MAP.
Those genes and SNPs, either known to be causative of established familial CRC syndromes or implicated in other GWLS or GWAS as being pre-disposing for CRC, are shown in Figure 1: EPHB2, MUTYH, MSH2, MSH6, MLH1, APC, PMS2, rs7014346/TCF4-binding site, TGFBR1, BMPR1A, PTEN, CRAC1, SMAD7, SMAD4, STK11, TP53 and HIC1. Interestingly, none of these genes is located within the novel regions of linkage described in this report, except for BMPR1A and PTEN, which reside in the minor region of linkage on 10q22.3–q24.1.
However, we also did not find any linkage to those regions of hereditary but non-syndromic CRC that have been recently reported on 3q,7, 10, 12 7q14 or 9q22,8, 11, 15 and it is interesting to ask why. Similar to ours, these studies7, 8, 9, 10, 11, 12, 13, 14, 15 used similar sample size and also excluded families with mutations in known predisposition genes for CRC to maximise the likelihood of discovering novel CRC risk genes. However, their experimental designs showed differences with each other and with ours. These included the degree to which families were stratified by clinical phenotypes,9 whether unaffected relatives were included or excluded from the linkage analyses and whether subjects with AA but no adenocarcinoma were classified as affected.7, 8, 9, 10, 11, 12, 13, 14, 15 A feature of all these studies, including ours, is that while promising candidate regions have been identified, there is limited concordance in the regions identified. Independent confirmation of common regions between studies has only been achieved for regions on 3q7, 10, 12 and 9q22.8, 11, 15 Such observations may be explained in a number of ways. One developing view is that outside of the mutations in the known syndromic genes, there are very few, if any, further high penetrance gene variants that predispose carriers to CRC. The bulk of familial CRC may result from the co-inheritance of relatively common variants in multiple other genes. These may individually impart a small but finite risk of CRC but in combination are responsible for the observed elevated risk in non-syndromic families.
An alternative explanation is that, in addition to the well-characterised familial CRC genes, mutations in a number of different genes may still be causative for familial CRC but that the frequency in the population of carriers of mutations in each of these genes is much lower. This could arise through the different target sizes of the genes or the need to retain certain functions. Clustering of mutations in a single causative gene within any one study, leading to detection of significant linkage, may then arise randomly through some hidden founder effect in a population group or through interaction of the causative mutation with other common genetic or environmental factors in that population group. A recent paper33 identifying a causative gene for familial pancreatic cancer is illustrative. Originally identified through whole-genome exon sequencing of an individual with familial pancreatic cancer, mutations in the same gene were subsequently identified as being causative in 3 of 90 families studied. If there are multiple different genes each contributing to a low percentage of familial CRC cases, the current study's designs and sizes will continue to have difficulty consistently identifying equivalent genes or loci.
We also considered whether different founder mutations existed in the Spanish and Australian population and that by combining them, there had been a dilution of the linkage signal. In fact a sensitivity analysis showed that the signals in the data from the two countries were consistent and jointly contributed to the overall significance of the results. Although this does not rule out different founder effects contributing to the observed differences between studies, it does suggest that it is not likely to be major factor. This should, however, be further investigated by larger inter-population comparisons.
Combining data from across a number of similarly performed individual studies may improve the overall power particularly where accumulated numbers are sufficient to stratify the data more rigorously. This could be based on any of a number of criteria but could include disease phenotype (either clinical or molecular), familial phenotype or cancer genotype.
To summarise, our data point to the likelihood that a mutation occurring in chromosomal region 10p15.3–p15.1 can cause an increased risk of CRC in these families. Next steps include localising the responsible genes and characterising their function. This is likely to provide insights into the factors responsible for susceptibility in non-syndromic CRC families and perhaps shed further light on the mechanisms of CRC development.
References
Johns LE, Houlston RS : A systematic review and meta-analysis of familial colorectal cancer risk. Am J Gastroenterol 2001; 96: 2992–3003.
Lipton LR, Johnson V, Cummings C et al: Refining the Amsterdam Criteria and Bethesda Guidelines: testing algorithms for the prediction of mismatch repair mutation status in the familial cancer clinic. J Clin Oncol 2004; 22: 4934–4943.
Terdiman JP, Conrad PG, Sleisenger MH : Genetic testing in hereditary colorectal cancer: indications and procedures. Am J Gastroenterol 1999; 94: 2344–2356.
Vasen HF, Mecklin JP, Khan PM, Lynch HT : The International Collaborative Group on Hereditary Non-Polyposis Colorectal Cancer (ICG-HNPCC). Dis Colon Rectum 1991; 34: 424–425.
Lindor NM, Rabe K, Petersen GM et al: Lower cancer incidence in Amsterdam-I criteria families without mismatch repair deficiency: familial colorectal cancer type X. JAMA 2005; 293: 1979–1985.
Lipkin SM, Afrasiabi K : Familial colorectal cancer syndrome X. Semin Oncol 2007; 34: 425–427.
Papaemmanuil E, Carvajal-Carmona L, Sellick GS et al: Deciphering the genetics of hereditary non-syndromic colorectal cancer. Eur J Hum Genet 2008; 16: 1477–1486.
Wiesner GL, Daley D, Lewis S et al: A subset of familial colorectal neoplasia kindreds linked to chromosome 9q22.2-31.2. Proc Natl Acad Sci U S A 2003; 100: 12961–12965.
Daley D, Lewis S, Platzer P et al: Identification of susceptibility genes for cancer in a genome-wide scan: results from the colon neoplasia sibling study. Am J Hum Genet 2008; 82: 723–736.
Kemp Z, Carvajal-Carmona L, Spain S et al: Evidence for a colorectal cancer susceptibility locus on chromosome 3q21-q24 from a high-density SNP genome-wide linkage scan. Hum Mol Genet 2006; 15: 2903–2910.
Kemp ZE, Carvajal-Carmona LG, Barclay E et al: Evidence of linkage to chromosome 9q22.33 in colorectal cancer kindreds from the United Kingdom. Cancer Res 2006; 66: 5003–5006.
Picelli S, Vandrovcova J, Jones S et al: Genome-wide linkage scan for colorectal cancer susceptibility genes supports linkage to chromosome 3q. BMC Cancer 2008; 8: 87.
Djureinovic T, Skoglund J, Vandrovcova J et al: A genome wide linkage analysis in Swedish families with hereditary non-familial adenomatous polyposis/non-hereditary non-polyposis colorectal cancer. Gut 2006; 55: 362–366.
Neklason DW, Kerber RA, Nilson DB et al: Common familial colorectal cancer linked to chromosome 7q31: a genome-wide analysis. Cancer Res 2008; 68: 8993–8997.
Gray-McGuire C, Guda K, Adrianto I et al: Confirmation of linkage to and localization of familial colon cancer risk haplotype on chromosome 9q22. Cancer Res 2010; 70: 5409–5418.
Weber MF, Banks E, Ward R, Sitas F : Population characteristics related to colorectal cancer testing in New South Wales, Australia: results from the 45 and Up Study cohort. J Med Screen 2008; 15: 137–142.
Ribes J, Navarro M, Cleries R et al: Colorectal cancer mortality in Spain: trends and projections for 1985–2019. Eur J Gastroenterol Hepatol 2009; 21: 92–100.
Saunders IW, Hannan GN, Brohede J et al: A range of simple summary genome-wide statistics for detecting genetic linkage using high density marker data. Genet Epidemiol 2007; 31: 565–576.
Wigginton JE, Abecasis GR : An evaluation of the replicate pool method: quick estimation of genome-wide linkage peak p-values. Genet Epidemiol 2006; 30: 320–332.
Baum L, Petrie T, Soules G, Weiss N : A maximization technique occuring in the statistical analysis of probabilistic functions of Markov Chains. Ann Math Statist 1970; 41: 164–171.
Saunders IW, Brohede J, Hannan GN : Estimating genotyping error rates from Mendelian errors in SNP array genotypes and their impact on inference. Genomics 2007; 90: 291–296.
Abecasis GR, Cherny SS, Cookson WO, Cardon LR : Merlin – rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 2002; 30: 97–101.
Kong A, Cox NJ : Allele-sharing models: LOD scores and accurate linkage tests. Am J Hum Genet 1997; 61: 1179–1188.
Whittemore AS, Halpern J : A class of tests for linkage using affected pedigree members. Biometrics 1994; 50: 118–127.
Lander E, Kruglyak L : Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet 1995; 11: 241–247.
Miyaki M, Yamaguchi T, Iijima T, Funata N, Mori T : Difference in the role of loss of heterozygosity at 10p15 (KLF6 locus) in colorectal carcinogenesis between sporadic and familial adenomatous polyposis and hereditary nonpolyposis colorectal cancer patients. Oncology 2006; 71: 131–135.
Pruitt KD, Tatusova T, Maglott DR : NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res 2007; 35: D61–D65.
Narla G, Heath KE, Reeves HL et al: KLF6, a candidate tumor suppressor gene mutated in prostate cancer. Science 2001; 294: 2563–2566.
Reeves HL, Narla G, Ogunbiyi O et al: Kruppel-like factor 6 (KLF6) is a tumor-suppressor gene frequently inactivated in colorectal cancer. Gastroenterology 2004; 126: 1090–1103.
Safran M, Chalifa-Caspi V, Shmueli O et al: Human gene-centric databases at the Weizmann institute of science: GeneCards, UDB, CroW 21 and HORDE. Nucleic Acids Res 2003; 31: 142–146.
Kury S, Buecher B, Robiou-du-Pont S et al: Low-penetrance alleles predisposing to sporadic colorectal cancers: a French case-controlled genetic association study. BMC Cancer 2008; 8: 326.
Seo T, Tatsuguchi A, Shinji S et al: Microsomal prostaglandin E synthase protein levels correlate with prognosis in colorectal cancer patients. Virchows Arch 2009; 454: 667–676.
Jones S, Hruban RH, Kamiyama M et al: Exomic sequencing identifies PALB2 as a pancreatic cancer susceptibility gene. Science 2009; 324: 217.
Kokko A, Laiho P, Lehtonen R et al: EPHB2 germline variants in patients with colorectal cancer or hyperplastic polyposis. BMC Cancer 2006; 6: 145.
Tenesa A, Campbell H, Barnetson R, Porteous M, Dunlop M, Farrington SM : Association of MUTYH and colorectal cancer. Br J Cancer 2006; 95: 239–242.
Vasen HF, Hendriks Y, de Jong AE et al: Identification of HNPCC by molecular analysis of colorectal and endometrial tumors. Dis Markers 2004; 20: 207–213.
Bodmer W : Familial adenomatous polyposis (FAP) and its gene, APC. Cytogenet Cell Genet 1999; 86: 99–104.
Tenesa A, Farrington SM, Prendergast JG et al: Genome-wide association scan identifies a colorectal cancer susceptibility locus on 11q23 and replicates risk loci at 8q24 and 18q21. Nat Genet 2008; 40: 631–637.
Tuupanen S, Turunen M, Lehtonen R et al: The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling. Nat Genet 2009; 41: 885–890.
Skoglund Lundin J, Vandrovcova J, Song B et al: TGFBR1 variants TGFBR1(*)6A and Int7G24A are not associated with an increased familial colorectal cancer risk. Br J Cancer 2009; 100: 1674–1679.
Daley D, Morgan W, Lewis S et al: Is TGFBR1*6A a susceptibility allele for nonsyndromic familial colorectal neoplasia? Cancer Epidemiol Biomarkers Prev 2007; 16: 892–894.
O’Riordan J, O’Donoghue D, Green A et al: Hereditary mixed polyposis syndrome due to a Bmpr1a mutation. Colorectal Dis 2010; 12: 570–573.
Nassif NT, Lobo GP, Wu X et al: PTEN mutations are common in sporadic microsatellite stable colorectal cancer. Oncogene 2004; 23: 617–628.
Jaeger E, Webb E, Howarth K et al: Common genetic variants at the CRAC1 (HMPS) locus on chromosome 15q13.3 influence colorectal cancer risk. Nat Genet 2008; 40: 26–28.
Gemignani F, Moreno V, Landi S et al: A TP53 polymorphism is associated with increased risk of colorectal cancer and with reduced levels of TP53 mRNA. Oncogene 2004; 23: 1954–1956.
Russo A, Bazan V, Iacopetta B, Kerr D, Soussi T, Gebbia N : The TP53 colorectal cancer international collaborative study on the prognostic and predictive significance of p53 mutation: influence of tumor site, type of mutation, and adjuvant treatment. J Clin Oncol 2005; 23: 7518–7528.
Iacopetta B : TP53 mutation in colorectal cancer. Hum Mutat 2003; 21: 271–276.
Pittman AM, Naranjo S, Webb E et al: The colorectal cancer risk at 18q21 is caused by a novel variant altering SMAD7 expression. Genome Res 2009; 19: 987–993.
Royce SG, Alsop K, Haydon A et al: The role of SMAD4 in early onset colorectal cancer. Colorectal Dis 2010; 12: 213–219.
Resta N, Simone C, Mareni C et al: STK11 mutations in Peutz-Jeghers syndrome and sporadic colon cancer. Cancer Res 1998; 58: 4799–4801.
Acknowledgements
We thank the patients and their families for their participation in the study, Bernadette Viney and Kerry Phillips for clinical research nurse support and Drs Konsta Duesing and Mike Buckley for critically reviewing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Saunders, I., Ross, J., Macrae, F. et al. Evidence of linkage to chromosomes 10p15.3–p15.1, 14q24.3–q31.1 and 9q33.3–q34.3 in non-syndromic colorectal cancer families. Eur J Hum Genet 20, 91–96 (2012). https://doi.org/10.1038/ejhg.2011.149
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2011.149
Keywords
This article is cited by
-
Linkage analysis revealed risk loci on 6p21 and 18p11.2-q11.2 in familial colon and rectal cancer, respectively
European Journal of Human Genetics (2019)
-
Exome sequencing in one family with gastric- and rectal cancer
BMC Genetics (2016)
-
Little evidence for association between the TGFBR1*6A variant and colorectal cancer: a family-based association study on non-syndromic family members from Australia and Spain
BMC Cancer (2014)
-
An association between the PTGS2 rs5275 polymorphism and colorectal cancer risk in families with inherited non-syndromic predisposition
European Journal of Human Genetics (2013)
-
Effects of Kruppel-like factor 6 on osteosarcoma cell biological behavior
Tumor Biology (2013)