Abstract
XPC participates in the initial recognition of DNA damage during the DNA nucleotide excision repair process in global genomic repair. Polymorphisms in XPC gene have been analyzed in case–control studies to assess the cancer risk attributed to these variants, but results are conflicting. To clarify the impact of XPC polymorphisms in cancer risk, we performed a meta-analysis that included 33 published case–control studies. Polymorphisms analyzed were Lys939Gln and Ala499Val. The overall summary odds ratio (OR) for the associations of the 939Gln/Gln genotype with risk of cancer was 1.01 (95% confidence interval (95% CI): 0.94–1.09), but there were statistically significant associations for lung cancer, observed for the recessive genetic model (Lys/Lys+Lys/Gln vs Gln/Gln), (OR 1.30; 95% CI: 1.113–1.53), whereas for breast cancer a reduced but nonsignificant risk was observed for the same model (OR 0.87; 95% CI: 0.74–1.01). The results for Ala499Val showed a significant overall increase in cancer risk (OR 1.15; 95% CI: 1.02–1.31), and for bladder cancer in both the simple genetic model (Ala/Ala vs Val/Val) (OR 1.30; 95% CI: 1.04–1.61) and the recessive genetic model (Ala/Ala+Ala/Val vs Val/Val) (OR 1.32; 95% CI: 1.06–1.63). Our meta-analysis supports that polymorphisms in XPC may represent low-penetrance susceptibility gene variants for breast, bladder, head and neck, and lung cancer. XPC is a good candidate for large-scale epidemiological case–control studies that may lead to improvement in the management of highly prevalent cancers.
Similar content being viewed by others
Introduction
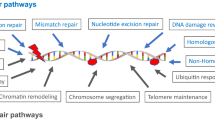
The development and progression of cancer has been attributed to the acquisition of capabilities that would allow for tumor growth, encompassing self-sufficiency for growth signals, tissue invasion and metastasis. The acquisition of these hallmarks are due to mutations in the genome of the cell,1 arising from unrepaired DNA lesions caused by UV exposure, ionizing radiation, environmental chemical agents and some substances produced by cell metabolism. Accumulation of such mutations drives the carcinogenic process. To safeguard the integrity of the genome, eukaryotic cells developed complex DNA repair systems that can recognize the lesions, excise them and restore the DNA, allowing for cell survival and thus preventing cancer.2 At least six distinct repair systems have been identified so far, including the nucleotide excision repair (NER), which has an important function as a versatile system that can eliminate a wide variety of lesions, such as UV-induced lesions, intrastrand cross-links and bulky adducts induced by chemical carcinogens.3
Nucleotide excision repair operates through two subpathways, namely the transcription-coupled repair (TCR) and the global genomic repair (GGR), which differ, among other aspects, in the step of recognition of damaged DNA. While in TCR it is the stalled RNA polII that initiates the repair process, in GGR the recognition of damage and initiation of repair is triggered by the XPC protein. The gene XPC is localized at 3p25 and encodes a protein of 940 amino acids that in vivo form a supramolecular complex including HR23B, homolog to Rad23 in yeast, and centrin 2.4 Regarding its function in DNA repair, the XPC complex binds to various types of lesions by recognizing alterations in the DNA structures rather than the lesions themselves.5 Following recognition, the XPC complex interacts with XPA, which constitutes with RPA the preincision complex. XPC also interacts with TFIIH, recruiting it to the lesion site, where this factor acts as a helicase opening the double helix allowing for the following steps of NER, excision and restoring of DNA.6
The relationship of XPC and cancer is based on the observation of high incidence of skin cancers (about 1000-fold) in patients with mutations in the XPC gene, which characterize the inherited disease xeroderma pigmentosum (XP). In XP patients, defects in NER are caused by mutations in one of the seven genes (XPA–G),7 increasing cancer susceptibility. Moreover, it has been suggested that XPC may play an important role in lung carcinogenesis8 and lymphomagenesis.9 Thus, based on its functions in recognition and initiation of NER, XPC is a key player in the repair of potential carcinogenic lesions.
A growing body of evidence has indicated that several low-penetrance gene variants (polymorphisms) have been considered to be involved in the pathogenesis of cancer, each contributing with smalls effects to the total genetic component.10 On the basis of that, a large number of molecular epidemiologic studies have been performed to evaluate the role of polymorphisms in DNA repair genes, including XPC, on various types of cancer. In XPC gene, three common polymorphisms are the most studied and include (1) a substitution of alanine for valine in codon 499 (Ala499Val), in the interaction domain of XPC with hHRAD23; (2) a substitution of lysine for glutamine in codon 939 (Lys939Gln), located in the interaction domain with TFIIH; and (3) a poly AT region on intron 9.11 Even with a considerable number of reports analyzing XPC polymorphisms, the results remain conflicting rather than conclusive. Studies with relative small sample sizes may have been underpowered to detect the effect of low-penetrance genes and their estimates may lack precision.
Thus, a quantitative synthesis may help to provide clearer evidence on the association of such genetic polymorphisms with cancer, as previously reported,12, 13 and find, even small but, relevant associations. The aim of the present study was to obtain summary risk estimates for the association of specific polymorphisms in XPC and risk of cancer, by conducting a meta-analysis from the available studies.
Materials and methods
Identification and eligibility of relevant studies
To identify all studies that examined the association of XPC polymorphisms with cancer, a search in the PubMed database (last search update in May 2007) was conducted using the keywords ‘XPC,’ ‘polymorphism,’ ‘polymorphisms’ and ‘cancer.’ Additional studies were identified by search of references from original papers and review articles. The inclusion criteria adopted included published studies that used an unrelated case–control design and had genotype frequency available.
Data extraction
Data were collected on the authors, journal, year of publication, country of origin, selection and characteristics of cancer cases and controls, sociodemographic information, ethnicity, genotyping information, genotyping method and interaction between environmental factors. Ethnicity was categorized as Caucasian, African and Asian. If a study did not state the ethnic descendent or if it was not possible to separate participants according to such phenotype, the group reported was termed ‘mixed ethnicity.’
Meta-analysis
Polymorphisms analyzed were the substitution Lys/Gln in codon 939 and Ala/Val in codon 499 of XPC. Another XPC polymorphism reported in the studies identified was an insertion of a 83-bp PAT (poly AT) in intron 9.11 However, this polymorphism is in strong linkage disequilibrium (LD) with Lys939Gln (Lewontin's LD=1.0).14, 15, 16 Thus, studies that just analyzed XPC PAT were considered as the same genotype frequency that XPC 939 due to this LD.
The association between XPC polymorphisms and cancer was estimated by calculating summary odds ratios (ORs). For Lys939Gln, we estimated the OR of cancer associated with Gln/Gln genotype compared with the wild-type Lys/Lys, and then the OR of Gln/Gln with (Lys/Lys+Lys/Gln) in a recessive genetic model and the OR of (Gln/Gln+Lys/Gln) with Lys/Lys in a dominant genetic model. Similar models were analyzed for the Ala499Val polymorphism. In addition to estimates of OR for all subjects in each study, studies were categorized into subgroups according to their sample's ethnicity and tumor type. Tumors sites that were studied in only one article were categorized into the ‘other cancers’ group.
Summary OR were obtained using fixed-effects models using the Mantel–Haenszel method.17 For each meta-analysis performed, a χ2-based Q-statistic test and an I2 was performed to asses the between-study heterogeneity,17 and heterogeneity was considered significant for P<0.05. In the case of heterogeneity, random-effects models with DerSimonian and Laird method were then used to pool the results.18 The significance of the pooled OR was determined by the Z-test. Publication bias was investigated by funnel plots and by the Egger's test.19 Hardy–Weinberg equilibrium (HWE) was tested by the χ2 test for goodness of fit. A sensitivity analysis was performed by calculating summary OR without studies not in HWE, and comparing results with those obtained with all available studies. All analyses were done in GraphPad Prism version 4 (GraphPad, San Diego, CA, USA), Stata version 9 and Review Manager 4.2 (Stata, Oxford, England). All the P-values were for two-sided analysis.
Bioinformatics analysis
Two in silico algorithms, the PolyPhen algorithm (http://tux.embl-heidelberg.de/ramensky/) and the SIFT algorithm (http://blocks.fhcrc.org/sift//SIFT.html), were used to predict the putative impact of the two polymorphisms in XPC on protein function. PolyPhen predicts the functional impact of amino-acid changes by considering evolutionary conservation, the physicochemical differences and the proximity of the substitution to predicted functional domains or structural features. PolyPhen scores (PISC) were designated as probably damaging (2.00), possibly damaging (1.50–1.99), potentially damaging (1.25–1.49), borderline (1.00–1.24) or benign (0.00–0.99) according to the classification proposed.20 SIFT predicts the functional importance of amino-acid substitutions on the basis of the alignment of ortholog or paralog protein sequences. SIFT scores were classified as intolerant (0.00–0.05), potentially intolerant (0.051–0.10), borderline (0.101–0.20) or tolerant (0.201–1.00) according to the classification proposed.20, 21 PolyPhen has been shown to have more than 80% accuracy of predicting deleterious amino-acid changes and the accuracy of SIFT is over 70%.20 The sensitivity of PolyPhen and SIFT for identifying deleterious mutations in XP genes is 85 and 83%, respectively.20, 21
Results
Meta-analysis data
The literature search yielded 37 studies that examined the relationship between XPC polymorphisms and different types of cancer. Four studies were excluded because they had case-only designs22, 23, 24, 25 and another because it investigated a sample already reported.26 Thus, the meta-analyses were based on 33 studies (Table 1). Thirty-one studies analyzed the relationship between XPC Lys939Gln and/or XPC PAT and risk of cancer, and two analyzed only XPC Ala499Val. Eleven studies analyzed both polymorphisms in the same article. Two studies reported results on different racial descendent population35, 58 and each population was treated as a separate comparison in meta-analysis. In general, tumors samples were confirmed by histological analysis. Regarding the choice of genotyping assays, a classic PCR-RFLP was done in 21 studies (64%), whereas the other 12 studies (36%) used a fluorogenic-based real-time PCR method. When the efficiency of both genotyping assays was compared, the PCR-RFLP method showed a mean rate of sample loss <1% (range: from 0 to 7%), whereas in real-time PCR-based method the mean rate of sample loss was 4% (range: from 1 to 14.5%). Table 1 shows the 33 studies included in this analysis and lists the cancer type of the study, country, ethnicity, number of cases and controls genotyped, and the frequency of minor allele in cases and controls.
The distribution of genotype frequencies among the control groups indicated that the gene variants were in HWE in all studies, but one38 (P=0.03). The interaction between XPC polymorphisms and environmental risk factors was investigated in 23 out of 33 studies (70%), 9 of which found some type of positive gene–environment association. In all, 2 of the 10 studies (30%) that did not analyze risk factors found association between XPC variants and the tumor studied.
Quantitative synthesis
XPC Lys939Gln
The studies that examined the relation between XPC Lys939Gln polymorphism and cancer risk analyzed a total of 14 080 cases and 14 011 controls. The Gln allele frequency in the three major ethnicities was 37.5% (95% CI: 35.4–39.6) for Caucasians, 36.1% (95% CI: 29.6–42.6) for Asians and 28% for African (Figure 1a), indicating a significant difference among Africans, as compared with the two other groups (P<0.001).
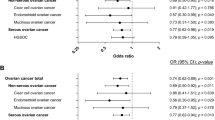
Overall, individuals with XPC Gln/Gln genotype did not have elevated cancer risk compared with individuals with Lys/Lys genotype, as shown by a summary OR of 1.01 (95% CI: 0.94–1.09; Figure 2). There was a trend for heterogeneity between studies, as suggested by a P-value of 0.05 for the χ2 test for heterogeneity, but the amount of variance between studies attributed to heterogeneity was only 30%. When the analyses were performed by cancer type (Figure 2), the homozygous variant of XPC Gln/Gln was associated with a significant increase in risk of lung cancer (OR 1.21; 95% CI: 1.02–1.44), and a borderline risk effect for head and neck (OR 1.37; 95% CI: 0.97–1.93) was observed. On the other hand, a nonsignificant (P=0.07) borderline protective effect was observed for breast cancer (OR 0.86; 95% CI: 0.73–1.01). To assess the importance of the heterozygous genotype, dominant and recessive genetic models were applied (Table 2). The recessive model confirmed the increased risk for lung cancer (OR 1.30; 95% CI: 1.11–1.53) and the nonsignificant borderline protective effect for breast cancer (P=0.05; OR 0.87; 95% CI: 0.74–1.01). No association with cancer risk was found in either tumor site using the dominant model (Table 2). Exclusion of the one study that was not HWE did not change the observed results. The next step was to analyze the studies according to racial descendent (Caucasians, Asians and mixed ethnicity), and to ethnicity and tumor site (Table 3). We did not find any association between XPC Lys939Gln for any genetic model stratified by ethnicity. However, lung cancer in the Asian samples showed a borderline risk associated with the Gln/Gln genotype (OR 1.23; 95% CI: 1.00–1.51) and an increased risk when the recessive genetic model was applied (OR 1.26; 95% CI: 1.04–1.52).
Forest plot showing ORs (log scale) (box) and 95% CI (horizontal line) for each study of cancer associated with XPC codon 939 for Lys/Lys genotype compared with Gln/Gln genotype. Studies are categorized by tumor site. n/N, n=number of Gln/Gln genotype, N=number of Lys/Lys plus Gln/Gln genotype. ♦, pooled OR and its 95% CI.
XPC Ala499Val
The studies included genotyped a total of 7603 cases and 7772 controls. There were significant differences in Val allele frequency between Caucasians and Asians (Caucasians: 24.75%, 95% CI: 21.2–28.27; Asian: 30.5%, 95% CI: 26.7–34.2; P<0.001; Figure 1b). No study analyzed cancer risk attributed to Ala499Val genotype in Africans.
Overall, individuals with Val/Val had an increased risk of cancer, compared with individuals with the Ala/Ala genotype, as shown by the summary OR of 1.15 (95% CI: 1.02–1.31; P=0.03), with no evidence of heterogeneity between studies (Figure 3). This increased cancer risk was also observed using the recessive genetic model (Table 2). When the analysis were performed by tumor site, an increased risk of bladder cancer was found in Val/Val genotype individuals, as compared with Ala/Ala ones (OR 1.30; 95% CI: 1.04–1.61) (Figure 3). This increased risk for bladder cancer was also observed when data were analyzed using the recessive model (Table 2). When summary OR values were examined by racial descendent, no significant association was verified, but Caucasians showed a borderline effect in Val/Val genotype and in the recessive model (Table 3).
Test of heterogeneity
There was marginal statistical heterogeneity between the 33 studies that examined the Lys939Gln polymorphism (P=0.05). When analyzing all cancer sites together, only the recessive model showed significant heterogeneity (χ2=49.21, d.f.=32, P=0.03). Significant heterogeneity was also observed in the ‘other cancers’ subgroup using the recessive model (χ2=8.10, d.f.=3, P=0.04), in the melanoma group using the dominant model (χ2=4.18, d.f.=1, P=0.04), and between studies with Caucasian samples (Lys/Lys vs Gln/Gln; χ2=22.18, d.f.=11, P=0.02; recessive model, χ2=21.40, d.f.=11, P=0.03). No significant heterogeneity was observed between the 15 studies that analyzed XPC Ala499Val, in any genetic model, apart from when the data were stratified by tumor site, the ‘other cancers’ subgroup showing heterogeneity using the dominant model (χ2=8.23, d.f.=2, P=0.02).
Publication bias
The funnel plot and Egger's test for the OR of studies comparing Gln/Gln with Lys/Lys for XPC Lys939Gln provided no evidence of publication bias (t=0.37, P=0.37). Similarly, there was no evidence of publication bias for the comparison Val/Val vs Ala/Ala for XPC Ala499Val (t=0.45, P=0.66).
‘In silico’ analysis of polymorphisms in XPC
With the aim of understanding the possible impact of these two amino-acid substitutions on XPC protein structure, we performed in silico analysis using PolyPhen and SIFT algorithms. Predictions utilizing PolyPhen indicated that the substitution of Lys939Gln could be possibly damaging to the protein, with a PSIC score of 1.618, whereas for the Ala499Val polymorphic variant, this exchange was predicted as benign, based on the score previously established, since the observed PSIC score was 0.346. When the predictions were carried out using the SIFT program, the Lys939Gln variant was classified as intolerant (score 0.00), whereas Ala499Val had a score of 0.20, classifying it as a ‘borderline’ substitution. However, predictions to Lys939Gln using SIFT had a low confidence result due to less than six aligned representative sequences.
Discussion
This meta-analysis of 33 case–control studies examined the association of two well-characterized polymorphisms of the DNA repair gene XPC (Lys939Gln and Ala499Val) with cancer risk. There was no overall effect on cancer risk for the Lys939Gln polymorphism in any genetic model, while a small increase in cancer risk was found for the Ala499Val polymorphism. When the XPC variants were analyzed by tumor site, for dominant or recessive genetic models, and by ethnic group, a few summary OR values were statistically significant. Different scenarios concerning the role of these polymorphisms can be interpreted from these results.
The overall no increase in risk of cancer for Lys939Gln represents the weighted average of the OR obtained from each study. While this suggests that this polymorphism may not play a major role in the risk of any cancer, it is a potential candidate for interaction studies. When examining the role of polymorphisms in DNA repair genes on cancer susceptibility, it is important to consider the importance of ‘gene–environment’ interactions, which are crucial to characterize low-penetrance genes.58 For example, for lung cancer and head and neck cancer, the main risk factor seems to be the exposure to tobacco carcinogens, such as benzo(a)pyrene, which form DNA adducts59 preferentially repaired by the NER pathway, where XPC acts in the initial recognition process of the DNA lesion. Therefore, interactions of the Gln/Gln genotype and tobacco exposure may lead to an increased risk of lung cancer, as previously reported,40 and head and neck cancer45 as well. Intriguingly, although smoking status also seems to be a risk factor for breast carcinogenesis.60 The XPC Gln/Gln variant seems to play a borderline protective role for breast cancer, such protective role was not statistically significant in any genetic model applied. It is still not possible to compare the exposure of airway epithelial cells and breast epithelial cells with the tobacco-derived carcinogens. Future studies using surrogate markers of carcinogen exposure will be useful to clarify this issue. However, as it is widely accepted, tobacco is quite a complex mixture of carcinogens, which may act in distinct phases of the carcinogenesis process. For example, toxicological studies point to the presence of organic solvents and distinct polycyclic aromatic hydrocarbons, which are not only mutagenic but may also be breast cancer promoters, acting as hormone disruptors.61
When applying the recessive genetic model to the different tumor sites, a significant increase in risk for lung cancer was also observed, strengthening the possibility of a role of XPC Gln/Gln genotype in these tissues. For head and neck cancer, however, no association could be seen in the recessive genetic model. Head and neck cancer may be a too general classification including a variety of cancer types in this anatomic region, whereas the role of XPC may be tissue specific.
The polymorphism in the XPC codon 939 replaces the positively charged amino acid lysine by the polar uncharged amino acid glutamine. The substitution does not change radically the hydropathy index of these amino acids (−3.9 to −3.5).62 However, in silico, the results indicated that Lys939Gln could be possibly damaging to the protein. It is important to consider that even though the computational analysis suggests a possible damage, this prediction may be not always correct, since protein–protein interactions may minimize amino-acid substitution-dependent conformation changes. Again, if this is correct, the effects observed will be either cell or tissue specific. Both biological and biochemical evidence also indicate the importance of XPC Lys939Gln polymorphism associated with a differential repair capacity.63, 64, 65, 66 Low repair capacity of benzo(a)pyrene DNA adducts was also observed in lymphocytes of individuals with this XPC variant,67 although the protein levels of XPC were not measured in that study. Moreover, XPC has two other polymorphisms in linkage with the Gln allele – an insertion of an 83-bp ATs (PAT) in intron 9 and an exchange of C to A in splicing site of intron 11 – that might contribute to a decreased expression of XPC, due to abnormalities in RNA stability and splicing process.11, 21 In a very elegant study, Wei et al68 showed that lymphocytes from head and neck cancer patients had an attenuated induction of XPC protein expression when transfected with damaged DNA, as compared with controls. For lung cancers, there is evidence for XPC promoter hypermethylation, leading to decreased transcription levels of the gene in the early events of carcinogenesis.69 Depletion of the XPC gene also led to lung tumors in mice, pointing to the importance of this gene for lung carcinogenesis.8 More recently, the involvement of XPC in photocarcinogenesis was suggested by experiments using XPC/CDKN2a double knockout mice.70
The in silico results for conformational changes in the substitution Ala499Val indicated an effect ranging between benign and borderline. However, in the Val/Val vs Ala/Ala and in the recessive genetic model, the variant genotype showed increased risk for bladder cancer, where the Val allele might contribute to the low efficiency for DNA repair in bladder tissue, contributing then to carcinogenesis. There are no biochemical reports showing the contribution of Val allele in repair or genetic stability assays, but a study analyzing the telomere length in bladder cancer patients demonstrated that individuals with Val/Val genotype had a nonsignificant (P=0.06) telomere shortening.71 For bladder cancer, established risk factors also include smoking habits as well as exposure to industrially related aromatic amines.72 The only two studies that analyzed the influence of risk factors adjusted to XPC Val/Val genotype also found an interaction between genetic and environmental factors for bladder cancer.29, 30
The present study has some limitations. First, the effect of XPC might be best represented by its haplotype. Among the nine studies that analyzed XPC haplotype,30, 33, 34, 37, 39, 42, 47, 50, 54 six found some type of association with cancer risk for the haplotypes analyzed. However, in the present meta-analysis, it was not possible to use XPC haplotypes, due to the small number of studies that had information about haplotype frequencies. Second, multiple testing may explain some or even all statistically significant associations observed in the present study. Given the multiplicity of comparisons for different cancer types, different genetic models and ethnic groups, and the unavoidable flexibility of choosing and defining the correlates, associations might have been detected by chance alone. Third, for some subgroup analyses there were only a very limited number of studies available, and therefore not having enough statistical power to detect association may explain some negative results. Even the total number of studies that analyzed Ala499Val polymorphism (15) was relatively small, in comparison to the number of studies that looked at the Lys939Gln polymorphism. Heterogeneity of ethnic ancestry may have also limited the ability of the meta-analyses in finding estimates of the true associations, since pooling samples with different ethnic backgrounds might produce a population average toward the null. However, summary estimates were obtained using the Mantel–Haenszel method, which calculates first an OR for each study and then an average estimate weighted for the precision of each study, and it is likely that within each study cases and controls had similar ethnic ancestry, thus reducing the likelihood of a summary estimate biased by differential ethnic ancestry between studies. On the other hand, the current meta-analysis has some key advantages, compared with individual studies. First, a substantial number of cases and controls were pooled from several studies, which significantly increased the statistical power of the analysis. Second, the subgroups that showed significant or borderline results did not have heterogeneity, which strengthens the analysis. Third, publication bias was not observed in both polymorphisms, which indicates that the pooled results should be unbiased.
In conclusion, this meta-analysis supports that polymorphisms in XPC gene may represent a low-penetrance gene for cancer risk, especially for breast, lung, head and neck, and bladder cancer. The current meta-analysis also illustrates the need to design epidemiologic case–control studies that include samples sizes with adequate statistical power to provide more conclusive evidence for associations between genotypes and diseases. These larger studies should also include analysis of risk factors, clarifying the interaction of haplotypes, gene–gene and gene–environment to tissue-specific cancer and to ethnicity specific populations. XPC plays a central role as a sensor of DNA distortions, such as those caused by both mutagenic and chemotherapeutic agents, such as cisplatin.73 Therefore, it will be important to complete the analysis of the involvement of specific XPC variants in tumorigenesis, with studies of its role on tumor progression and response to chemotherapy. Moreover, based on what we have addressed in the present study, XPC is a good candidate for the new generation of large-scale epidemiological case–control studies that will ultimately lead to a better diagnosis, treatment and prevention of prevalent cancers.
References
Hanahan D, Weinberg RA : The hallmarks of cancer. Cell 2000; 100: 57–70.
Sancar A, Lindsey-Boltz LA, Unsal-Kacmaz K, Linn S : Molecular mechanisms of mammalian DNA repair and the DNA damage checkpoints. Annu Rev Biochem 2004; 73: 39–85.
Friedberg EC : How nucleotide excision repair protects against cancer. Nat Rev Cancer 2001; 1: 22–33.
Araki M, Masutani C, Takemura M et al: Centrosome protein centrin 2/caltractin 1 is part of the xeroderma pigmentosum group C complex that initiates global genome nucleotide excision repair. J Biol Chem 2001; 276: 18665–18672.
Sugasawa K, Shimizu Y, Iwai S, Hanaoka F : A molecular mechanism for DNA damage recognition by the xeroderma pigmentosum group C protein complex. DNA Repair 2002; 1: 95–107.
Costa RM, Chigancas V, Galhardo Rda S, Carvalho H, Menck CF : The eukaryotic nucleotide excision repair pathway. Biochimie 2003; 85: 1083–1099.
Magnaldo T, Sarasin A : Xeroderma pigmentosum: from symptoms and genetics to gene-based skin therapy. Cells Tissues Organs 2004; 177: 189–198.
Hollander MC, Philburn RT, Patterson AD et al: Deletion of XPC leads to lung tumors in mice and is associated with early events in human lung carcinogenesis. Proc Natl Acad Sci USA 2005; 102: 13200–13205.
Soufir N, Ribojad M, Magnaldo T et al: Germline and somatic mutations of the INK4a-ARF gene in a xeroderma pigmentosum group C patient. J Invest Dermatol 2002; 119: 1355–1360.
Mohrenweiser HW, Wilson 3rd DM, Jones IM : Challenges and complexities in estimating both the functional impact and the disease risk associated with the extensive genetic variation in human DNA repair genes. Mutat Res 2003; 526: 93–125.
Khan SG, Metter EJ, Tarone RE et al: A new xeroderma pigmentosum group C poly(AT) insertion/deletion polymorphism. Carcinogenesis 2000; 21: 1821–1825.
Hu Z, Ma H, Chen F, Wei Q, Shen H : XRCC1 polymorphisms and cancer risk: a meta-analysis of 38 case–control studies. Cancer Epidemiol Biomarkers Prev 2005; 14: 1810–1818.
Han S, Zhang HT, Wang Z et al: DNA repair gene XRCC3 polymorphisms and cancer risk: a meta-analysis of 48 case–control studies. Eur J Hum Genet 2006; 14: 1136–1144.
Khan SG, Muniz-Medina V, Shahlavi T et al: The human XPC DNA repair gene: arrangement, splice site information content and influence of a single nucleotide polymorphism in a splice acceptor site on alternative splicing and function. Nucleic Acids Res 2002; 30: 3624–3631.
Blankenburg S, Konig IR, Moessner R et al: Assessment of 3 xeroderma pigmentosum group C gene polymorphisms and risk of cutaneous melanoma: a case–control study. Carcinogenesis 2005; 26: 1085–1090.
Sak SC, Barrett JH, Paul AB, Bishop DT, Kiltie AE : The polyAT, intronic IVS11-6 and Lys939Gln XPC polymorphisms are not associated with transitional cell carcinoma of the bladder. Br J Cancer 2005; 92: 2262–2265.
Egger M, Smith GD, Altman DG : Systematic Reviews in Health Care: Meta-Analysis in Context. London: BMJ Publishing Group, 2001.
Petitti DB : Meta-Analysis, Decision Analysis, and Cost-Effectiveness Analysis. New York: Oxford University Press, 1994.
Egger M, Davey Smith G, Schneider M, Minder C : Bias in meta-analysis detected by a simple, graphical test. BMJ 1997; 315: 629–634.
Xi T, Jones IM, Mohrenweiser HW : Many amino acid substitution variants identified in DNA repair genes during human population screenings are predicted to impact protein function. Genomics 2004; 83: 970–979.
Ng PC, Henikoff S : Predicting deleterious amino acid substitutions. Genome Res 2001; 11: 863–874.
Gu J, Zhao H, Dinney CP et al: Nucleotide excision repair gene polymorphisms and recurrence after treatment for superficial bladder cancer. Clin Cancer Res 2005; 11: 1408–1415.
Berndt SI, Platz EA, Fallin MD, Thuita LW, Hoffman SC, Helzlsouer KJ : Genetic variation in the nucleotide excision repair pathway and colorectal cancer risk. Cancer Epidemiol Biomarkers Prev 2006; 15: 2263–2269.
Millikan RC, Hummer A, Begg C et al: Polymorphisms in nucleotide excision repair genes and risk of multiple primary melanoma: the Genes Environment and Melanoma Study. Carcinogenesis 2006; 27: 610–618.
Ryk C, Kumar R, Sanyal S et al: Influence of polymorphism in DNA repair and defence genes on p53 mutations in bladder tumours. Cancer Lett 2006; 241: 142–149.
Weiss JM, Weiss NS, Ulrich CM, Doherty JA, Chen C : Nucleotide excision repair genotype and the incidence of endometrial cancer: effect of other risk factors on the association. Gynecol Oncol 2006; 103: 891–896.
Sanyal S, Festa F, Sakano S et al: Polymorphisms in DNA repair and metabolic genes in bladder cancer. Carcinogenesis 2004; 25: 729–734.
Garcia-Closas M, Malats N, Real FX et al: Genetic variation in the nucleotide excision repair pathway and bladder cancer risk. Cancer Epidemiol Biomarkers Prev 2006; 15: 536–542.
Sak SC, Barrett JH, Paul AB, Bishop DT, Kiltie AE : Comprehensive analysis of 22 XPC polymorphisms and bladder cancer risk. Cancer Epidemiol Biomarkers Prev 2006; 15: 2537–2541.
Zhu Y, Lai M, Yang H et al: Genotypes, haplotypes and diplotypes of XPC and risk of bladder cancer. Carcinogenesis 2007; 28: 698–703.
Wu X, Gu J, Grossman HB et al: Bladder cancer predisposition: a multigenic approach to DNA-repair and cell-cycle-control genes. Am J Hum Genet 2006; 78: 464–479.
Forsti A, Angelini S, Festa F et al: Single nucleotide polymorphisms in breast cancer. Oncol Rep 2004; 11: 917–922.
Shen J, Desai M, Agrawal M et al: Polymorphisms in nucleotide excision repair genes and DNA repair capacity phenotype in sisters discordant for breast cancer. Cancer Epidemiol Biomarkers Prev 2006; 15: 1614–1619.
Jorgensen TJ, Visvanathan K, Ruczinski I, Thuita L, Hoffman S, Helzlsouer KJ : Breast cancer risk is not associated with polymorphic forms of xeroderma pigmentosum genes in a cohort of women from Washington County, Maryland. Breast Cancer Res Treat 2007; 101: 65–71.
Mechanic LE, Millikan RC, Player J et al: Polymorphisms in nucleotide excision repair genes, smoking and breast cancer in African Americans and whites: a population-based case–control study. Carcinogenesis 2006; 27: 1377–1385.
Zhang L, Zhang Z, Yan W : Single nucleotide polymorphisms for DNA repair genes in breast cancer patients. Clin Chim Acta 2005; 359: 150–155.
Hu Z, Wang Y, Wang X et al: DNA repair gene XPC genotypes/haplotypes and risk of lung cancer in a Chinese population. Int J Cancer 2005; 115: 478–483.
Shen M, Berndt SI, Rothman N et al: Polymorphisms in the DNA nucleotide excision repair genes and lung cancer risk in Xuan Wei, China. Int J Cancer 2005; 116: 768–773.
Lee GY, Jang JS, Lee SY et al: XPC polymorphisms and lung cancer risk. Int J Cancer 2005; 115: 807–813.
Marin MS, Lopez-Cima MF, Garcia-Castro L, Pascual T, Marron MG, Tardon A : Poly (AT) polymorphism in intron 11 of the XPC DNA repair gene enhances the risk of lung cancer. Cancer Epidemiol Biomarkers Prev 2004; 13: 1788–1793.
Vogel U, Overvad K, Wallin H, Tjonneland A, Nexo BA, Raaschou-Nielsen O : Combinations of polymorphisms in XPD, XPC and XPA in relation to risk of lung cancer. Cancer Lett 2005; 222: 67–74.
Bai Y, Xu L, Yang X et al: Sequence variations in DNA repair gene XPC is associated with lung cancer risk in a Chinese population: a case–control study. BMC Cancer 2007; 7: 81.
Yang M, Kang MJ, Choi Y et al: Associations between XPC expression, genotype, and the risk of head and neck cancer. Environ Mol Mutagen 2005; 45: 374–379.
Sugimura T, Kumimoto H, Tohnai I et al: Gene–environment interaction involved in oral carcinogenesis: molecular epidemiological study for metabolic and DNA repair gene polymorphisms. J Oral Pathol Med 2006; 35: 11–18.
Shen H, Sturgis EM, Khan SG et al: An intronic poly (AT) polymorphism of the DNA repair gene XPC and risk of squamous cell carcinoma of the head and neck: a case–control study. Cancer Res 2001; 61: 3321–3325.
Kietthubthew S, Sriplung H, Au WW, Ishida T : Polymorphism in DNA repair genes and oral squamous cell carcinoma in Thailand. Int J Hyg Environ Health 2006; 209: 21–29.
Li C, Hu Z, Liu Z et al: Polymorphisms in the DNA repair genes XPC, XPD, and XPG and risk of cutaneous melanoma: a case–control analysis. Cancer Epidemiol Biomarkers Prev 2006; 15: 2526–2532.
Blankenburg S, Konig IR, Moessner R et al: No association between three xeroderma pigmentosum group C and one group G gene polymorphisms and risk of cutaneous melanoma. Eur J Hum Genet 2005; 13: 253–255.
Casson AG, Zheng Z, Evans SC, Veugelers PJ, Porter GA, Guernsey DL : Polymorphisms in DNA repair genes in the molecular pathogenesis of esophageal (Barrett) adenocarcinoma. Carcinogenesis 2005; 26: 1536–1541.
Zhou RM, Li Y, Wang N, Zhang XJ, Dong XJ, Guo W : Correlation of XPC Ala499Val and Lys939Gln polymorphisms to risks of esophageal squamous cell carcinoma and gastric cardiac adenocarcinoma. Ai Zheng (Chinese Journal of Cancer) 2006; 25: 1113–1119.
Ye W, Kumar R, Bacova G, Lagergren J, Hemminki K, Nyren O : The XPD 751Gln allele is associated with an increased risk for esophageal adenocarcinoma: a population-based case–control study in Sweden. Carcinogenesis 2006; 27: 1835–1841.
Festa F, Kumar R, Sanyal S et al: Basal cell carcinoma and variants in genes coding for immune response, DNA repair, folate and iron metabolism. Mutat Res 2005; 574: 105–111.
Nelson HH, Christensen B, Karagas MR : The XPC poly-AT polymorphism in non-melanoma skin cancer. Cancer Lett 2005; 222: 205–209.
Weiss JM, Weiss NS, Ulrich CM, Doherty JA, Voigt LF, Chen C : Interindividual variation in nucleotide excision repair genes and risk of endometrial cancer. Cancer Epidemiol Biomarkers Prev 2005; 14: 2524–2530.
Huang WY, Berndt SI, Kang D et al: Nucleotide excision repair gene polymorphisms and risk of advanced colorectal adenoma: XPC polymorphisms modify smoking-related risk. Cancer Epidemiol Biomarkers Prev 2006; 15: 306–311.
Hirata H, Hinoda Y, Matsuyama H et al: Polymorphisms of DNA repair genes are associated with renal cell carcinoma. Biochem Biophys Res Commun 2006; 342: 1058–1062.
Hirata H, Hinoda Y, Tanaka Y et al: Polymorphisms of DNA repair genes are risk factors for prostate cancer. Eur J Cancer 2007; 43: 231–237.
Lohmueller KE, Pearce CL, Pike M, Lander ES, Hirschhorn JN : Meta-analysis of genetic association studies supports a contribution of common variants to susceptibility to common disease. Nat Genet 2003; 33: 177–182.
Hecht SS : DNA adduct formation from tobacco-specific N-nitrosamines. Mutat Res 1999; 424: 127–142.
Terry PD, Rohan TE : Cigarette smoking and the risk of breast cancer in women: a review of the literature. Cancer Epidemiol Biomarkers Prev 2002; 11: 953–971.
Brody JG, Rudel RA : Environmental pollutants and breast cancer. Environ Health Perspect 2003; 111: 1007–1019.
Kyte J, Doolittle RF : A simple method for displaying the hydropathic character of a protein. J Mol Biol 1982; 157: 105–132.
Qiao Y, Spitz MR, Shen H et al: Modulation of repair of ultraviolet damage in the host–cell reactivation assay by polymorphic XPC and XPD/ERCC2 genotypes. Carcinogenesis 2002; 23: 295–299.
Qiao Y, Spitz MR, Guo Z et al: Rapid assessment of repair of ultraviolet DNA damage with a modified host–cell reactivation assay using a luciferase reporter gene and correlation with polymorphisms of DNA repair genes in normal human lymphocytes. Mutat Res 2002; 509: 165–174.
Cornetta T, Festa F, Testa A, Cozzi R : DNA damage repair and genetic polymorphisms: assessment of individual sensitivity and repair capacity. Int J Radiat Oncol Biol Phys 2006; 66: 537–545.
Naccarati A, Soucek P, Stetina R et al: Genetic polymorphisms and possible gene–gene interactions in metabolic and DNA repair genes: effects on DNA damage. Mutat Res 2006; 593: 22–31.
Pavanello S, Pulliero A, Siwinska E, Mielzynska D, Clonfero E : Reduced nucleotide excision repair and GSTM1-null genotypes influence anti-B[a]PDE-DNA adduct levels in mononuclear white blood cells of highly PAH-exposed coke oven workers. Carcinogenesis 2005; 26: 169–175.
Wei Q, Wang LE, Sturgis EM, Mao L : Expression of nucleotide excision repair proteins in lymphocytes as a marker of susceptibility to squamous cell carcinomas of the head and neck. Cancer Epidemiol Biomarkers Prev 2005; 14: 1961–1966.
Wu YH, Tsai Chang JH, Cheng YW, Wu TC, Chen CY, Lee H : Xeroderma pigmentosum group C gene expression is predominantly regulated by promoter hypermethylation and contributes to p53 mutation in lung cancers. Oncogene 2007; 26: 4761–4773.
Yang G, Curley D, Bosenberg MW, Tsao H : Loss of xeroderma pigmentosum C (Xpc) enhances melanoma photocarcinogenesis in Ink4a-Arf-deficient mice. Cancer Res 2007; 67: 5649–5657.
Broberg K, Bjork J, Paulsson K, Hoglund M, Albin M : Constitutional short telomeres are strong genetic susceptibility markers for bladder cancer. Carcinogenesis 2005; 26: 1263–1271.
Cohen SM, Shirai T, Steineck G : Epidemiology and etiology of premalignant and malignant urothelial changes. Scand J Urol Nephrol Suppl 2000; 205: 105–115.
Chen Z, Yang J, Wang G, Song B, Li J, Xu Z : Attenuated expression of xeroderma pigmentosum group C is associated with critical events in human bladder cancer carcinogenesis and progression. Cancer Res 2007; 67: 4578–4585.
Acknowledgements
PRM, JEN and RC were partly funded by the CNPq-Brazil. GF was a fellow from Fundação de Amparo a Pesquisa do Estado de São Paulo (FAPESP).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Francisco, G., Menezes, P., Eluf-Neto, J. et al. XPC polymorphisms play a role in tissue-specific carcinogenesis: a meta-analysis. Eur J Hum Genet 16, 724–734 (2008). https://doi.org/10.1038/ejhg.2008.6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2008.6
Keywords
This article is cited by
-
Smoking and XPC Gene Polymorphism Interact to Modulate the Risk of Oral Cancer
Journal of Maxillofacial and Oral Surgery (2021)
-
Polymorphisms in the XPC gene affect urinary bladder cancer risk: a case-control study, meta-analyses and trial sequential analyses
Scientific Reports (2016)
-
Potential risk of esophageal squamous cell carcinoma due to nucleotide excision repair XPA and XPC gene variants and their interaction among themselves and with environmental factors
Tumor Biology (2016)
-
Oxidative and Energy Metabolism as Potential Clues for Clinical Heterogeneity in Nucleotide Excision Repair Disorders
Journal of Investigative Dermatology (2015)
-
XPC gene polymorphisms contribute to bladder cancer susceptibility: a meta-analysis
Tumor Biology (2014)