Abstract
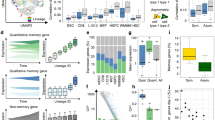
Pre-B lymphoblastic leukemia cells carrying a BCR–ABL1 gene rearrangement exhibit an undifferentiated phenotype. Comparing the genome-wide gene expression profiles of normal B-cell subsets and BCR–ABL1+ pre-B lymphoblastic leukemia cells by SAGE, the leukemia cells show loss of B lymphoid identity and aberrant expression of myeloid lineage-specific molecules. Consistent with this, BCR–ABL1+ pre-B lymphoblastic leukemia cells exhibit defective expression of IKAROS, a transcription factor needed for early lymphoid lineage commitment. As shown by inducible expression of BCR–ABL1 in human and murine B-cell precursor cell lines, BCR–ABL1 induces the expression of a dominant-negative IKAROS splice variant, termed IK6. Comparing matched leukemia sample pairs from patients before and during therapy with the BCR–ABL1 kinase inhibitor STI571 (Imatinib), inhibition of BCR–ABL1 partially corrected aberrant expression of IK6 and lineage infidelity of the leukemia cells. To elucidate the contribution of IK6 to lineage infidelity in BCR–ABL1+ cell lines, IK6 expression was silenced by RNA interference. Upon inhibition of IK6, BCR–ABL1+ leukemia cells partially restored B lymphoid lineage commitment. Therefore, we propose that BCR–ABL1 induces aberrant splicing of IKAROS, which interferes with lineage identity and differentiation of pre-B lymphoblastic leukemia cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Abbreviations
- IGH :
-
immunoglobulin heavy chain
- SAGE:
-
serial analysis of gene expression
- siRNA:
-
short-interfering RNA
References
Busslinger M . (2004). Annu Rev Immunol 22: 55–79.
Feldhahn N, Klein F, Mooster JL, Hadweh P, Sprangers M, Wartenberg M et al. (2005). J Exp Med 201: 1837–1852.
Feldhahn N, Schwering I, Lee S, Wartenberg M, Klein F, Wang H et al. (2002). J Exp Med 196: 1291–1305.
Georgopoulos K, Bigby M, Wang JH, Molnar A, Wu P, Winandy S et al. (1994). Cell 79: 143–156.
Huettner CS, Zhang P, Van Etten RA, Tenen DG . (2000). Nat Genet 24: 57–60.
Jumaa H, Bossaller L, Portugal K, Storch B, Lotz M, Flemming A et al. (2003). Nature 423: 452–456.
Klein F, Feldhahn N, Harder L, Wang H, Wartenberg M, Hofmann WK et al. (2004). J Exp Med 199: 673–685.
Klein F, Feldhahn N, Lee S, Wang H, Ciuffi F, von Elstermann M et al. (2003). Proc Natl Acad Sci USA 100: 6747–6752.
Klein F, Feldhahn N, Mooster JL, Sprangers M, Hofmann WK, Wernet P et al. (2005). J Immunol 174: 367–375.
Klucher KM, Lopez DV, Daley GQ . (1998). Blood 91: 3927–3934.
Kirstetter P, Thomas M, Dierich A, Kastner P, Chan S . (2002). Eur J Immunol 32: 720–730.
Laurent E, Talpaz M, Kantarjian H, Kurzrock R . (2001). Cancer Res 61: 2343–2355.
Look AT . (1997). Science 278: 1059–1064.
Müschen M, Lee S, Zhou G, Feldhahn N, Barath VS, Chen J et al. (2002). Proc Natl Acad Sci USA 99: 10014–10019.
Nakase K, Ishimaru F, Avitahl N, Dansako H, Matsuo K, Fujii K et al. (2000). Cancer Res 60: 4062–4065.
Perrotti D, Calabretta B . (2002). Oncogene 21: 8577–8583.
Salesse S, Dylla SJ, Verfaillie CM . (2004). Leukemia 18: 727–733.
Sun L, Heerema N, Crotty L, Wu X, Navara C, Vassilev A et al. (1999). Proc Natl Acad Sci USA 96: 680–685.
Tonnelle C, Bardin F, Maroc C, Imbert AM, Campa F, Dalloul A et al. (2001). Blood 98: 2673–2680.
Velculescu VE, Zhang L, Vogelstein B, Kinzler KW . (1995). Science 270: 484–487.
Acknowledgements
We would like to thank Martin Krönke (Köln), Janet D Rowley (Chicago) and Klaus Rajewsky (Boston) for continuous support, Wolf-Karsten Hofmann (Berlin) for provision of leukemia samples, George Q Daley (Cambridge) for provision of TONB210, and Stefanie Jauch and Peter Wurst for excellent technical assistance. FK is supported by the Studienstiftung des deutschen Volkes and the Köln Fortune program of the Faculty of Medicine of the University of Cologne. NF is supported by a fellowship of the José-Carreras-Leukemia Foundation. MM is supported by the Deutsche Forschungsgemeinschaft through the Emmy-Noether-Programm, the German José-Carreras-Leukemia-Foundation (grant to MM), the Deutsche Krebshilfe through joint project grant ‘Mechanisms of Malignant Lymphoma’ (MM) and the Ministry of Science and Research for North Rhine-Westphalia through the Stem Cell Network NRW (to MM).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Supplementary information
Rights and permissions
About this article
Cite this article
Klein, F., Feldhahn, N., Herzog, S. et al. BCR–ABL1 induces aberrant splicing of IKAROS and lineage infidelity in pre-B lymphoblastic leukemia cells. Oncogene 25, 1118–1124 (2006). https://doi.org/10.1038/sj.onc.1209133
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209133
Keywords
This article is cited by
-
Biochemical and cellular characterization of transcription factors binding to the hyperconserved core promoter-associated M4 motif
BMC Genomics (2016)
-
Prognostic value of rare IKZF1 deletion in childhood B-cell precursor acute lymphoblastic leukemia: an international collaborative study
Leukemia (2016)
-
Newly diagnosed acute lymphoblastic leukemia in China (I): abnormal genetic patterns in 1346 childhood and adult cases and their comparison with the reports from Western countries
Leukemia (2012)
-
Expression of dominant-negative Ikaros isoforms and associated genetic alterations in Chinese adult patients with leukemia
Annals of Hematology (2012)
-
Genomic profiling of high-risk acute lymphoblastic leukemia
Leukemia (2010)