Abstract
Apoptosis is often accompanied by degradation of chromosomal DNA. CAD, caspase-activated DNase, was identified in 1998 as a DNase that is responsible for this process. In the last several years, mice deficient in the CAD system have been generated. Studies with these mice indicated that apoptotic DNA degradation occurs in two different systems. In one, the DNA fragmentation is carried out by CAD in the dying cells and in the other, by lysosomal DNase II after the dying cells are phagocytosed. Several other endonucleases have also been suggested as candidate effectors for the apoptotic degradation of chromosomal DNA. In this review, we will discuss the mechanism and role of DNA degradation during apoptosis.
Similar content being viewed by others
Introduction
Programmed cell death or apoptosis is a process that removes toxic or useless cells that are produced during animal development.1 During tissue turnover, senescent cells die via apoptosis to maintain homeostasis. Apoptosis can be induced by a variety of external stimuli, including death factors such as Fas ligand, γ-irradiation, anticancer drugs, and factor deprivation.2,3,4,5 Failure of apoptosis causes an accumulation of abnormal cells leading to cancer and autoimmune diseases, whereas its exacerbation causes tissue destruction such as that seen in hepatitis, AIDS, and neurodegenerative diseases.
Apoptosis is defined by morphological changes of cells and nuclei, such as cell shrinkage, condensation and fragmentation of nuclei, and blebbing of the plasma membranes.6 This process is accompanied by the degradation of chromosomal DNA in most cases. In contrast to the necrotic cell death that accompanies the rupture of plasma membranes, the cellular contents are thought not to be released during the apoptotic process. However, if dying cells are not properly disposed of, they rupture and undergo secondary necrosis. Thus, apoptotic cells must be swiftly recognized and engulfed by professional phagocytes (macrophages and dendritic cells) or neighboring cells.7,8 Extensive studies of the signal trans-duction of apoptosis identified caspases (cysteine-protease) as the effectors.9 Whatever the apoptotic stimuli are, a set of caspases seems to be activated to cleave cellular substrates to kill the cells10 and to present one or more ‘eat me’ signals to phagocytes.7
Identification of the DNase responsible for the apoptotic DNA degradation
In 1980, Wyllie reported11 that when mouse thymocytes were treated with glucocorticoid, a substantial amount of chromatin complex was released from the nucleus. The DNA in the released complex was double stranded, and consisted of multimers of about 180 bp, which is the size of the DNA comprising a nucleosome. Since the protein composition in the complex was not different from that in intact nucleosomes, it was concluded that a double-strand specific endonuclease(s) is activated during apoptosis and cleaves the nucleosomal spacer regions to yield a DNA ladder. Sub-sequently, the application of pulsed-field gel electrophoresis showed that prior to or in the absence of internucleosomal cleavage, chromosomal DNA is cleaved into large fragments of 50–300 kb.12,13 The sequence of the cleavage sites is A/T-rich, as found in the nuclear scaffold region,14 suggesting that chromatins are cleaved by a nuclease at the nuclear scaffold attachment sites. A different requirement for metal ions (Mg2+ and Ca2+) initially suggested that the DNase(s) for internucleosomal cleavage and large-scale degradation might differ.15
Many groups since then have searched for the nuclease(s) responsible for this process. However, the DNase was labile and scanty, making purification of the enzyme extremely difficult.16 Nonetheless, several enzymes were proposed as candidates, including DNase I,17 DNase II,18 cyclophilins,19 and DNase γ;20 none of them, however, appeared to fulfill the criteria for the apoptotic DNase. For example, DNase I is a secreted protein with a signal sequence, expressed only in a limited set of tissues, such as the kidney, pancreas, and seminal plasma. It is not, however, expressed in lymphocytes, which efficiently undergo apoptotic DNA fragmentation. DNase I-mutant mice were recently established by gene targeting, and the analysis of these mice showed that DNase I is responsible for the removal of nuclear debris at sites of high cell turnover,21 but not for the degradation of DNA in apoptotic cells (personal communication from Dr. Hans G Mannherz, Ruhr-Universität Bochum, Germany). DNase II is a lysosomal enzyme, and apoptotic DNA degradation occurs at the early stage of apoptosis when lysosomes have not yet been destroyed, suggesting that DNase II might not function to fragment DNA in apoptotic cells. However, as discussed below, DNase II seems to work in phagocytes to cleave the DNA of the apoptotic cells after phagocytes engulf them.
Meanwhile, caspases have been identified as effectors of the apoptotic process,22 and DNA fragmentation as well as nuclear morphological changes have been placed downstream of caspase activity.23,24,25,26,27,28 When cytosolic extracts from proliferating cells are treated with caspases, they induce DNA fragmentation in isolated nuclei.29,30,31 The factor responsible, designated as DNA fragmentation factor (DFF), has been purified from HeLa cells to homogeneity as a heterodimer of 45 and 40-kDa subunits.30 When DFF was treated with caspase 3, the 45-kDa subunit (DFF45) was specifically cleaved, and DFF then caused DNA fragmentation in nuclei. However, since the DNase activity was not detected in this preparation, Liu et al.30 initially concluded that the DNase responsible for the DNA fragmentation resided in the nuclei, and that DFF activated the DNase(s). A similar approach by us32 and Williams' group33 has led purification of a DNase of 40 kDa from mouse lymphoma WR19L and human Jurkat cells, respectively. This molecule was designated as caspase-activated DNase (CAD) by us or caspase-activated nuclease (CPAN) by Williams' group. The DNase activity of CAD/CPAN could be inhibited by ICAD (inhibitor of CAD, or DFF45), and the inhibitory activity of ICAD was destroyed by caspase 3. In addition to action on naked DNA, CAD/CPAN caused the fragmentation of chromosomal DNA in nuclei. These similar-ities between DFF and CAD eventually led us to conclude that DFF is a complex of CAD and ICAD.34,35 The initial failure to detect the DNase activity of human DFF40 was apparently because of its extreme instability.33,34
Caspase-activated DNase (CAD, DFF40)
CAD has so far been identified in six species (human, mouse, rat, Zebrafish, Drosophila, and chicken) (Figure 1). Except for Drosophila CAD, which is a 52-kDa protein with an isoelectric point of 7.2, the CADs from other species are basic proteins of about 40 kDa with a pI of 8.7–9.5. Drosophila CAD contains a caspase-recognition sequence (EETD) in the middle of the molecule, which is cleaved by caspases to become an active enzyme comprised of large and small subunits (p32 and p20).36 In contrast, the CAD from other species does not undergo post-translational cleavage, and has no subunit structure. The enzymatic properties of CAD have been mainly characterized with the mouse and human proteins. CAD binds to DNA without Mg2+,37 and has an endonuclease activity at neutral pH in the presence of Mg2+.38 It is specific for double-stranded DNA with a preference for A/T-rich region, and it does not cleave single-stranded DNA or RNA. CAD mainly generates DNA fragments with blunt ends carrying a 5′-phosphate and 3′ hydroxyl group,38 which can be substrates for the terminal transferase39 used for the TUNEL (TdT-mediated dUTP-biotin nick end labeling) reaction.
Alignment of the amino-acid sequences of CAD from six different organisms. The amino-acid sequences of CAD from Drosophila melanogaster,36 mouse,32 rat,103 human,34,35 zebrafish (GenBank Accession number: AF286179), and chicken75 are aligned to give a maximum homology by introducing several gaps (−). The amino-acid residues conserved among six members are shown in bold. The caspase-recognition sequence in Drosophila CAD is boxed. Three lysine residues and one tyrosine residue essential for the CAD's DNase activity are shown in blue and brown, respectively. Four histidine residues required for the DNase activity are shown in red. Among these histidine residues, His-313 in mouse sequence is not conserved in Drosophila, but His-433 of Drosophila CAD may have a similar function. A putative nuclear localization signal is underlined. The total numbers of amino acids for each proteins, and their isoelectric points are listed at the end
CAD has a high specific activity. The homogeneous recombinant mouse CAD (0.1 pmol) generates about 30 pmol of 3′ hydroxyl ends after 30 min at 30°C when it is assayed using optimal conditions.39 Human and mouse CADs behave peculiarly in gel filtration: when analyzed using a low-salt buffer, they stick to the gel (H Sakahira and SN, unpublished result) or elute as aggregates,33,40 probably because of their extremely positive charge. On the other hand, active Drosophila CAD behaves like a protein of about 100 kDa consisting of (p32)2(p19)2.36 Several histidine, lysine, and tyrosine residues in the C-terminal half of CAD are well conserved among different species (Figure 1).39,41 Chemical and site-directed mutagenesis showed that these conserved residues are essential for its DNase activity, indicating that CAD is a member of the histidine nuclease family that includes RNase T1, DNase I, and S1 nuclease.39,41,42 Histone H1, high mobility group protein (HMG)-1, HMG-2, and DNA topo-isomerase IIα were reported to bind CAD and stimulate its DNase or DNA fragmentation activity.34,40,43 On the other hand, when the recombinant CAD, free of the residual ICAD, was assayed under optimal conditions, histone HI had no effect on CAD's DNase activity (H Sakahira and SN, unpublished result). Although we cannot rule out the possibility that these molecules regulate the CAD activity in cells, it is likely that the effect of these proteins is to stabilize CAD during the assay, or to remove caspase-cleaved ICAD from CAD.
Inhibitor of CAD (ICAD, DFF45)
ICAD has been identified in human, mouse, rat, and Drosophila. In human and mouse, there are two forms of ICAD that are encoded by alternatively spliced mRNAs; these are designated ICAD-L and ICAD-S for the long and short forms, respectively.32,44 ICAD-L and ICAD-S are acidic proteins of 45 and 30 kDa, and they inhibit CAD's DNase activity with the same efficiency.37,45 They inhibit CAD by blocking its binding to DNA.39 In proliferating cells, ICAD-L but not ICAD-S exists as a heterodimer (CAD/ICAD-L) with CAD in mouse and human cells,32,40 or as a heterotetramer (CAD)2/(ICAD-L)2 in Drosophila cells.36 This is because ICAD-L but not ICAD-S binds to the nascent polypeptide of CAD while it is on ribosomes.46 During its synthesis, ICAD-L, together with general chaperones of hsp70 and hsp40, helps CAD to be folded correctly. It is released from ribosomes as a complex with CAD when the CAD synthesis is completed. Thus, the functionally folded CAD protein cannot be produced without ICAD-L,47 and ICAD-L must be coexpressed in mammalian, insect, or E. coli cells to produce functional CAD.32,37,40 In addition to the regulation of CAD by ICAD-L, we recently found that the expression of ICAD-L but not ICAD-S is regulated by CAD at the post-translational level, suggesting that their expression is mutually regulated.104 No specific function has been assigned to ICAD-S, and it is probably dispensable in most if not all species, as has been shown in Drosophila.48 In this regard, results by Chen et al.49 indicating that rat and mouse tissues express only ICAD-S are difficult to explain.
Human and mouse ICADs carry two caspase-recognition sites, at Asp-117 and 224,30,44 while there is a single caspase recognition site in Drosophila ICAD at Asp-119.48 Among members of the mammalian caspase family, caspases 3 and 7 cleave ICAD.50,51 Mammalian ICAD must be cleaved at both sites to lose its CAD-inhibiting activity. That is, ICAD fragments carrying amino-acid residues from 1 to 224 or 118 to 331 still have a substantial affinity for CAD and can inhibit its DNase activity.44,52,53 The human breast carcinoma MCF-7 cell line carrying a deletion in the caspase 3 gene,28 or mouse cells derived from caspase 3-null mice54 do not undergo DNA fragmentation upon exposure to apoptotic stimuli. In these cells, ICAD is cleaved at Asp-117 but not Asp-224, suggesting an absolute requirement of caspase 3 to cleave ICAD at Asp-224 for full activation of CAD.55 Differing from the initial report,51 two reports have claimed that granzyme B, a serine protease released from cytotoxic T cells (CTL), cleaves ICAD at Asp-117 to activate CAD.56,57 However, the fact that ICAD-L can still inhibit the CAD's DNase when cleaved only at Asp-117 may not support the idea that ICAD-L is cleaved in this way in CTL-induced apoptotic DNA fragmentation.
Interaction between CAD and ICAD
As described above, ICAD-L can be divided into three domains (amino acids 1–117, 118–224, and 225–331) that can be separated by cleavage with caspase 3. These three domains synergistically contribute to ICAD's ability to bind CAD and inhibit its DNase activity.52,53 Among these domains, the N-terminal domain of about 80 amino acids is significantly homologous to the N-terminal domain of CAD, and is called the CIDE or CAD (CD) domain.35,58 The ICAD-CD and CAD-CD specifically interact with each other, and this interaction is important for ICAD-L to recognize the nascent polypeptide of CAD on ribosomes.46 The tertiary structures of the CAD-CD and its complex with the ICAD-CD have been resolved by the NMR analysis.59,60,61 Both CAD-CD and ICAD-CD are composed of one α-helix and five β-strands, and belong to the ubiquitin-superfold family. In the complex, they are aligned side by side through ionic interactions between basic residues in CAD-CD and acidic residues in ICAD-CD.59 Thus, point mutations in the acidic residues of the ICAD-CD abolish the ability of ICAD-L to facilitate the correct folding of CAD during its synthesis, although the mutants can still inhibit the CAD's DNase.
Cellular Localization of CAD and ICAD
CAD must function in the nucleus to digest chromosomal DNA. CAD from human, mouse, rat, and zebrafish carries a typical nuclear localization signal (NLS) at the C terminus (Figure 1). An NLS-like sequence has also been identified in the C-terminal region of ICAD-L.62,63 However, there is contradictory evidence regarding which cellular compartments contain CAD and ICAD in proliferating cells. The CAD/ICAD (DFF) complex was purified from the cytosolic fraction of HeLa and mouse lymphoma cells.30,32 The fractionation of cellular organelles followed by Western blotting showed that ICAD-L is mainly localized in the cytosol.64 From these results, we initially suggested that CAD/ICAD complex resides in the cytoplasm in proliferating cells and that CAD translocates into the nucleus after ICAD is cleaved by caspase 3.32 CAD but not the CAD/ICAD complex enters nuclei in a cell-free system, or in a semipermeabilized cell system, which seemed to agree with this proposal. On the other hand, immunostaining of endogenous ICAD showed it to be in the nucleus of human HeLa and epithelial cells,34,63 but in the cytoplasm of activated thymocytes and Hodgkin's lymphoma cells.65,66 The expression pattern of exogenous ICAD tagged with Flag or GFP also gave contradictory results. Two reports showed that the exogenously expressed ICAD is mainly localized in the nucleus,63,67 but one report showed a cytoplasmic localization.65 Whether these contradictory results indicate that ICAD is localized in the nucleus or cytoplasm in a cell-specific manner, or whether they are because of technical problems in the cell fractionation and/or immunostaining procedure remains to be clarified. In this regard, it may be worth noting that Drosophila CAD and ICAD do not carry an NLS at the position corresponding to the NLS of their mammalian counterparts (Figure 1).36,48
Contribution of CAD to DNA Degradation
During apoptosis, DNA is cleaved into blunt end double-stranded fragments carrying a 5′ phosphate and 3′ hydroxyl group.68 Using this knowledge, two methods – TUNEL69 and LM-PCR (linker-mediated PCR)70 – were developed to label the cleaved DNA. The application of these methods indicated that DNA fragmentation takes place in most apoptotic processes in mammals, Drosophila, and C. elegans.69,70,71
CAD and ICAD are expressed at the mRNA and protein levels in various tissues with abundant expression in lymphoid tissues such as the thymus, spleen, and lymph nodes.72,73 Thymocytes, lymphocytes, and lymphoid cell lines undergo massive DNA fragmentation upon exposure to apoptotic stimuli (Figure 2). In contrast, mouse embryonal fibroblasts (MEF) express a low level of CAD, and the presence of apoptotic DNA fragmentation is not clear.74 The contribution of CAD to DNA fragmentation was examined by expressing a caspase-resistant ICAD, or by deleting the ICAD or CAD gene by the gene-targeting method (KK, HF, and SN, unpublished results).47,50,75 ICAD that carries mutations (Asp to Glu mutation at P1 position) in the caspase-recognition sites binds CAD and inhibits its DNase, but since this molecule cannot be cleaved by caspases, CAD is not activated. Since ICAD-L is indispensable for the production of functional CAD protein, ICAD-null cells behave as if they lacked CAD. Thus, cells expressing the caspase-resistant ICAD, or that are deficient in the ICAD or CAD gene, show similar phenotypes. Splenocytes, thymocytes, and granulocytes from ICAD (DFF45)-null mice are resistant to DNA fragmentation upon exposure to various apoptotic stimuli such as staurosporine, etoposide, and actinomycin D.47,76 Human cell lines (Jurkat lymphoma and IL-3-dependent TF-1 myeloid lymphoma) and the Drosophila BG-2 neuronal cell line that were transformed with the caspase-resistant ICAD, or thymocytes from the transgenic mice carrying the caspase-resistant ICAD gene, do not undergo DNA fragmentation after treatment with Fas ligand, factor deprivation, anticancer drugs, or kinase inhibitors.48,50,73 Chicken DT40 cells that lack the CAD gene do not show fragmented DNA after treatment with etoposide.75 Lymphocytes and hepatocytes from the CAD-null mice are also resistant to the apoptotic DNA fragmentation (Figure 2, data not shown). From these results, it is clear that CAD is responsible for the degradation of DNA into nucleosomal units during apoptosis.
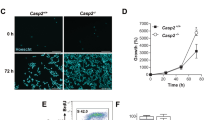
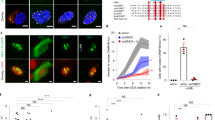
CAD-dependent apoptotic DNA fragmentation in mouse splenocytes and embryonal fibroblasts. (a) Apoptotic DNA fragmentation in mouse splenocytes. The S-30 fractions (20 μg protein each) from splenocytes of the wild-type (WT) and CAD−/− (CAD KO) mice were analyzed by Western blotting using the anti-CAD mAb (panel a). The relative molecular weights of the standard proteins are shown in kDa on the left. The band for CAD is indicated by an arrow on the right. Mouse splenocytes (1 × 106 cells/ml) from wild-type (□) and CAD−/− (▪) mice were incubated at 37°C with 500 U/ml of human Fas ligand. At the indicated time, cells were stained with FITC-labeled annexin V, and the percentage of annexin V-positive cells is shown in panel b. The chromosomal DNA was analyzed by electrophoresis on an agarose gel and is shown in panel c. The sizes of the marker DNAs are given in bp on the left. (b) Apoptotic DNA fragmentation in MEF. The S-30 fractions (40 μg protein each) from the wild-type (WT), CAD−/− (CAD-KO), and CAD-transformed (CAD) MEFs were analyzed by Western blotting using the anti-CAD mAb (panel a). The wild-type (○), CAD−/− (▴), and CAD-transformed MEFs (□) (1 × 104 cells) were incubated in 96-well microtiter plates at 37°C for the indicated periods with 5000 U/ml Fas ligand in the presence of 1.5 μg/ml cycloheximide. The cell viability was analyzed by staining the cells with crystal violet, and is shown as the percentage (panel b). The fragmentation of chromosomal DNA is shown in panel c. Similar results were obtained with treatment with staurosporine
There are, however, reports that challenge the idea that CAD is indispensable in apoptotic DNA fragmentation. During apoptosis induced by a set of stimuli, many proteins are released from the mitochondria.77 One is endonuclease G, which is usually localized to the intermembrane space of the mitochondria. Since this nuclease caused DNA fragmentation in isolated nuclei, two groups suggested that endo-nuclease G works as an apoptotic endonuclease.78,79 However, this nuclease is (dG)n/(dC)n-specific,80 preferentially generates single-stranded nicks, and cleaves not only DNA in internucleosomal regions but also DNA within nucleosomes.81 As discussed by Wildlak et al.,81 it is unlikely that endonuclease G alone is involved in the apoptotic cleavage of chromosomal DNA. The observation that the number of TUNEL-positive cells increases in an endonuclease G-deficient mutant of C. elegans82 supports this notion.
As described above, the apoptotic fragmentation of DNA into nucleosomal units is preceded by the degradation of chromosomal DNA into a high-Mr DNA of 50–300 kb.12,13 Cells expressing the caspase-resistant ICAD,83 or thymocytes and embryonal fibroblasts from ICAD (DFF45)-null mice, lack this type of chromosomal DNA degradation upon treatment with various apoptotic stimuli,74,84 suggesting that CAD is also responsible for the large-scale degradation of chromosomal DNA. On the other hand, the large-scale degradation of chromosomal DNA was observed in chicken DT40 cells lacking CAD, and Samejima et al.75 proposed that an ICAD-inhibitable non-CAD DNase causes this type of degradation. No caspase-dependent DNase activity was detected when the extracts from the thymocytes from CAD-null mice were treated with caspase 3 (HN, and SN, unpublished results), and all the ICAD seems to be associated with CAD in Drosophila cells,48 indicating that there are no additional ICAD-inhibitable DNases in mouse and Drosophila cells. Purification and characterization of the nuclease that causes the large-scale DNA degradation from chicken cells will be necessary to clarify this apparent controversy. In addition, apoptosis-inducing factor (AIF), which is also released from mitochondria during apoptosis, has been reported to cause the large-scale DNA degradation.85 However, AIF has no nuclease activity, and the nuclease that is activated by AIF has not been identified. It is therefore unclear whether AIF has a role in the apoptotic DNA degradation.
Contribution of the DNA Degradation to Cell Death
When CAD is injected into cells, chromatins quickly condense and are fragmented.86 We are certain that the degradation of chromosomal DNA by CAD is an efficient way to kill the cells. However, this system is apparently dispensable for cell death. That is, cells expressing caspase-resistant ICAD or cells deficient in ICAD or CAD are still efficiently killed by treatment with apoptotic stimuli (Figure 2). Caspases that are activated during the apoptotic process cleave more than 100 cellular substrates such as actin and cyclin A.87 Without the function of these molecules, cells will not survive. In some cases such as glucocorticoid-treated thymocytes, kainic acid-treated neuronal cells, or tumor necrosis factor (TNF)-treated MEF, CAD deficiency causes a delay in the cell death processes.74,76,88 Treatment of MEF cells with TNF does not cause DNA fragmentation, but induces the large-scale degradation of chromosomal DNA.74 This is followed by activation of poly(ADP-ribose) polymerase (PARP), which leads to the depletion of NAD and ATP that kills the cells. Since the large-scale DNA degradation does not occur in DFF45 (ICAD)-null cells, NAD is not depleted and the cell death is delayed. On the other hand, staurosporine or Fas ligand, which induces apoptosis more strongly than TNF does, kills CAD-null, wild-type, or CAD-transformed MEF cells with the same efficiency (Figure 2). It is possible that strong apoptosis inducers such as staurosporine and Fas ligand activate caspases sufficiently to kill the cells directly. In contrast, if the caspases are not sufficiently activated by weak stimuli such as TNF, another pathway, such as CAD-dependent PARP activation, may contribute to the killing of the cells.
Mice deficient in CAD do not show adverse phenotypes, and do not accumulate undigested nuclei or DNA in their tissues. Although one report claims that spatial learning and memory are enhanced in DFF45 (ICAD)-null mice,89 this was not confirmed by another group.90 At a minumum, the longevity and fertility of CAD- or ICAD-null mice kept in specific pathogen-free conditions in our Medical School are not different from those of wild-type mice, indicating that the CAD-mediated apoptotic DNA degradation is not required for normal animal development.
DNA Degradation of Apoptotic Cells in Phagocytes
With a few exceptions,78,90 cells deficient in CAD do not undergo apoptotic DNA degradation in vitro, and do not become TUNEL-positive. However, very surprisingly, the thymus, ovary, and brain of CAD-deficient mice were found to contain TUNEL-positive cells to the same extent as in wild-type mice.73 These TUNEL-positive cells were always inside macrophages, as revealed by electron microscopy. When wild-type mice were irradiated with γ-rays, massive TUNEL-positive cells were generated outside the macrophages in the thymus. The same treatment of the CAD-deficient mice killed many thymocytes, but they were not TUNEL-positive. This was explained as follows: During development, cells undergoing negative or positive selection in the thymus die in an orderly manner, and the dying cells, whether their DNA is fragmented or not, are immediately engulfed by phagocytes. The engulfed dead cells will enter lysosomes, and their DNA is degraded by a lysosomal enzyme(s). On the other hand, if massive cell death is induced by γ-rays, the capacity of phagocytes to engulf the cells is exceeded, and the corpses with their TUNEL-positive nuclei are left unengulfed. In any case, these results indicate that the DNA of apoptotic cells can be degraded in phagocytes after the cells are engulfed.73,91 In support of this idea, when the apoptotic CAD-deficient cells were cocultured with macrophages, their DNA was degraded in the lysosomes of macrophages.73,92
Lysosomes carry a DNase called DNase II. It generates DNA fragments carrying 5′ hydroxyl and 3′ phosphate ends.93 Thus, it is unlikely that the action of DNase II produces TUNEL-positive cells. However, lysosomes also carry acid phosphatases that may remove the 3′ phosphate from the DNA fragments and allow them to become TUNEL-reactive at a certain time point. The DNase II-null mice accumulate many undigested nuclei in the macrophages of the fetal liver, and die as fetuses because of the failure of definitive erythropoiesis.94 In addition, these embryos harbor a large amount of DNA that is left undigested in the macrophages of the tissues where apoptotic cell death occurs.95 A similar deposit of DNA was previously reported in a C. elegans mutant lacking DNase II (Nuc-1).96 Although there is a possibility that Nuc-1 may function in the dying cells in C. elegans,71 its localization in lysosomes strongly suggests that DNase II functions in phagocytes, at least in the mouse system.
Conclusion and perspective
The degradation of chromosomal DNA is a well-conserved apoptotic process. It can be seen in mammals, Drosophila, C. elegans, and plants. As discussed above, this process is mediated in vivo by two systems, one operating cell-autonomously in the dying cells, and the other in phagocytes after the dying cells are engulfed. In cell-autonomous DNA degradation, DNA is first cleaved into fragments of about 50 kb with intact nucleosomes, followed by degradation into nucleosomal units. Although we cannot rule out the possibility that other nucleases (endonuclease G, unidentified AIF-activated nuclease, and others) degrade DNA in certain circumstances, CAD is mainly responsible for this cell-autonomous DNA degradation. A simple explanation is, when CAD is activated, it first digests DNA at the nuclear scaffold region since this region is most exposed and A/T-rich. This unpacks the chromatin, and CAD then gains access to the spacer regions of nucleosomes to produce DNA fragments in nucleosomal units. There is a single gene for CAD in mammals and Drosophila. In C. elegans, Nuc-1 is known as a nuclease involved in DNA degradation,96 but no CAD homolog has been identified. However, TUNEL-positive cells can be observed in C. elegans, in particular when Nuc-1 is mutated,71 suggesting that this animal also carries a CAD-like DNase.
One of the most surprising findings in our laboratory in recent years is that apoptotic DNA degradation or the generation of TUNEL-positive cells takes place in vivo in mice that are deficient in the CAD system. This and the identification of Nuc-1 as a DNase II-homolog by Wu et al.71 clarified the role for DNase II in digesting the DNA of apoptotic cells in macrophages. In C. elegans, Nuc-1 works in a Ced-3 (caspase)-dependent manner.71 Macrophages engulf only apoptotic cells by recognizing phosphatidylserine that is exposed on the cell surface in a caspase-dependent manner.7 In this regard, DNase II in macrophages also works in a caspase-dependent manner in the apoptotic process. In addition to DNase II, lysosomes contain abundant proteases that can digest the proteins of the phagocytosed apoptotic cells. Thus, the DNA of the engulfed apoptotic cells will be completely digested into nucleotides by DNase II in lysosomes.97 We recently established Drosophila mutants lacking both CAD and DNase II.105 In the CAD single mutant, there was no accumulation of DNA in embryonal tissues. In contrast, in the DNase II mutant, fragmented DNA accumulated in macrophages, and in the double mutant for the CAD and DNase II genes, large, probably intact, chromosomal DNA accumulated in macrophages. These results support the two-step model for apoptotic DNA degradation.
DNA is an immunogenic molecule, and activates immune responses.98 If the DNA of apoptotic cells is left undigested, it may have adverse effects on the homeostasis of animals, as found in DNase I-null mice.21 The accumulation of undigested DNA in various tissues in DNase II-null mice but not CAD-deficient mice suggests that DNA degradation in macrophages may play a more vital role than CAD-mediated DNA degradation. If so, why have cells evolved an elaborate system to degrade chromosomal DNA by themselves? This system is dispensable for cells to die, and it is not necessary for phagocytes to engulf dying cells. During development, apoptotic cells are recognized by phagocytes at a very early stage, when no apparent morphological change or DNA fragmentation has occurred.99,100 Once apoptotic cells present an ‘eat me’ signal, phagocytes can degrade the proteins and DNA of apoptotic cells, and may actively kill the cells.101,102 In this regard, the role of caspases may be just to present the ‘eat me’ signal to phagocytes. It is not clear to us, however, why caspases cleave so many cellular proteins, and CAD cleaves the chromosomal DNA if phagocytes can do the job. On the other hand, in pathological settings such as inflammation, ischemia, and acute infection, a large load of apoptotic cells may exceed the local phagocytic clearance capacity. In these situations, cell-autonomous degradation of chromosomal DNA by CAD, and the cleavage of cellular proteins by caspases may be more beneficial for organisms than releasing intact DNA or proteins into the circulation. We hope that in the near future, we will have a clearer picture of the role of DNA degradation in mammalian homeostasis and in various human diseases.
Abbreviations
- CAD:
-
caspase-activated DNase
- ICAD:
-
inhibitor of CAD
- DFF:
-
DNA fragmentation factor
References
Jacobson MD, Weil M and Raff MC (1997) Programmed cell death in animal development. Cell 88: 347–354
Evan G and Littlewood T (1998) A matter of life and cell death. Science 281: 1317–1322
Nagata S (1997) Apoptosis by death factor. Cell 88: 355–365
Ashkenazi A and Dixit VM (1998) Death receptors: signaling and modulation. Science 281: 1305–1308
Green DR and Reed JC (1998) Mitochondria and apoptosis. Science 281: 1309–1312
Kerr JF, Wyllie AH and Currie AR (1972) Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br. J. Cancer 26: 239–257
Fadok VA, Bratton DL, Frasch SC, Warner ML and Henson PM (1998) The role of phosphatidylserine in recognition of apoptotic cells by phagocytes. Cell Death Differ. 5: 551–562
Savill J and Fadok V (2000) Corpse clearance defines the meaning of cell death. Nature 407: 784–788
Thornberry NA and Lazebnik Y (1998) Caspases: enemies within. Science 281: 1312–1316
Samali A, Zhivotovsky B, Jones D, Nagata S and Orrenius S (1999) Apoptosis: cell death defined by caspase activation. Cell Death Differ. 6: 495–496
Wyllie AH (1980) Glucocorticoid-induced thymocyte apoptosis is associated with endogenous endonuclease activation. Nature 284: 555–556
Lagarkova MA, Iarovaia OV and Razin SV (1995) Large-scale fragmentation of mammalian DNA in the course of apoptosis proceeds via excision of chromosomal DNA loops and their oligomers. J. Biol. Chem. 270: 20239–20241
Oberhammer F, Wilson JW, Dive C, Morris ID, Hickman JA, Wakeling AE, Walker PR and Sikorska M (1993) Apoptotic death in epithelial cells: cleavage of DNA to 300 and/or 50 kb fragments prior to or in the absence of internucleosomal fragmentation. EMBO J. 12: 3679–3684
Khodarev NN, Bennett T, Shearing N, Sokolova I, Koudelik J, Walter S, Villalobos M and Vaughan AT (2000) LINE L1 retrotransposable element is targeted during the initial stages of apoptotic DNA fragmentation. J. Cell Biochem. 79: 486–495
Sun XM and Cohen GM (1994) Mg(2+)-dependent cleavage of DNA into kilobase pair fragments is responsible for the initial degradation of DNA in apoptosis. J. Biol. Chem. 269: 14857–14860
Cohen JJ, Duke RC, Fadok VA and Sellins KS (1992) Apoptosis and programmed cell death in immunity. Annu. Rev. Immunol. 10: 267–293
Peitsch MC, Polzar B, Stephan H, Crompton T, MacDonald HR, Mannherz HG and Tschopp J (1993) Characterization of the endogenous deoxyribonuclease involved in nuclear DNA degradation during apoptosis (programmed cell death). EMBO J. 12: 371–377
Barry MA and Eastman A (1993) Identification of deoxyribonuclease II as an endonuclease involved in apoptosis. Arch. Biochem. Biophys. 300: 440–450
Montague JW, Hughes FJ and Cidlowski JA (1997) Native recombinant cyclophilins A, B, and C degrade DNA independently of peptidylprolyl cis–trans-isomerase activity. Potential roles of cyclophilins in apoptosis. J. Biol. Chem. 272: 6677–6684
Shiokawa D, Iwamatsu A and Tanuma S (1997) Purification, characterization and amino acid sequencing of DNase γ from rat spleen. Arch. Biochem. Biophys. 346: 15–20
Napirei M, Karsunky H, Zevnik B, Stephan H, Mannherz HG and Moroy T (2000) Features of systemic lupus erythematosus in DNase1-deficient mice. Nat. Genet. 25: 177–181
Nicholson DW, Ali A, Thornberry NA, Vaillancourt JP, Ding CK, Gallant M, Gareau Y, Griffin PR, Labelle M, Lazebnik YA, Munday NA, Raju SM, Smulson ME, Yamin T-T, Yu VL and Miller DK (1995) Identification and inhibition of the ICE/CED-3 protease necessary for mammalian apoptosis. Nature 376: 37–43
Chow SC, Weis M, Kass GEN, Holmstrom TH, Eriksson JE and Orrenius S (1995) Involvement of multiple proteases during Fas-mediated apoptosis in T lymphocytes. FEBS Lett. 364: 134–138
Dubrez L, Savoy I, Hamman A and Solary E (1996) Pivotal role of a DEVD-sensitive step in etoposide-induced and Fas-mediated apoptotic pathways. EMBO J. 15: 5504–5512
Enari M, Hug H and Nagata S (1995) Involvement of an ICE-like protease in Fas-mediated apoptosis. Nature 375: 78–81
Enari M, Hase A and Nagata S (1995) Apoptosis by a cytosolic extract from Fas-activated cells. EMBO J. 14: 5201–5208
Martin SJ, Newmeyer DD, Mathias S, Farschon DM, Wang H-G, Reed JC, Kolesnik RN and Green DR (1995) Cell-free reconstitution of Fas-, UV radiation- and ceramide-induced apoptosis. EMBO J. 14: 5191–5200
Janicke RU, Sprengart ML, Wati MR and Porter AG (1998) Caspase-3 Is required for DNA fragmentation and morphological changes associated with apoptosis. J. Biol. Chem. 273: 9357–9360
Enari M, Talanian RV, Wong WW and Nagata S (1996) Sequential activation of ICE-like and CPP32-like proteases during Fas-mediated apoptosis. Nature 380: 723–726
Liu X, Zou H, Slaughter C and Wang X (1997) DFF, a heterodimeric protein that functions downstream of caspase-3 to trigger DNA fragmentation during apoptosis. Cell 89: 175–184
Mitamura S, Ikawa H, Mizuno N, Kaziro Y and Itoh H (1998) Cytosolic nuclease activated by caspase-3 and inhibited by DFF-45. Biochem. Biophys. Res. Commun. 243: 480–484
Enari M, Sakahira H, Yokoyama H, Okawa K, Iwamatsu A and Nagata S (1998) A caspase-activated DNase that degrades DNA during apoptosis, and its inhibitor ICAD. Nature 391: 43–50
Halenbeck R, MacDonald H, Roulston A, Chen TT, Conroy L and Williams LT (1998) CPAN, a human nuclease regulated by the caspase-sensitive inhibitor DFF45. Curr. Biol. 8: 537–540
Liu X, Li P, Widlak P, Zou H, Luo X, Garrard WT and Wang X (1998) The 40-kDa subunit of DNA fragmentation factor induces DNA fragmentation and chromatin condensation during apoptosis. Proc. Natl. Acad. Sci. USA 95: 8461–8466
Mukae N, Enari M, Sakahira H, Fukuda Y, Inazawa J, Toh H and Nagata S (1998) Molecular cloning and characterization of human caspase-activated DNase. Proc. Natl. Acad. Sci. USA 95: 9123–9128
Yokoyama H, Mukae N, Sakahira H, Okawa K, Iwamatsu A and Nagata S (2000) A novel activation mechanism of caspase-activated DNase from Drosophila melanogaster. J. Biol. Chem. 275: 12978–12986
Sakahira H, Enari M and Nagata S (1999) Functional differences of two forms of the inhibitor of caspase- activated DNase, ICAD-L, and ICAD-S. J. Biol. Chem. 274: 15740–15744
Widlak P, Li P, Wang X and Garrard WT (2000) Cleavage preferences of the apoptotic endonuclease DFF40 (caspase-activated DNase or nuclease) on naked DNA and chromatin substrates. J. Biol. Chem. 275: 8226–8232
Sakahira H, Takemura Y and Nagata S (2001) Enzymatic active site of caspase-activated DNase (CAD) and its inhibition by inhibitor of CAD (ICAD). Arch. Biochem. Biophys. 388: 91–99
Liu X, Zou H, Widlak P, Garrard W and Wang X (1999) Activation of the apoptotic endonuclease DFF40 (caspase-activated DNase or nuclease). Oligomerization and direct interaction with histone H1. J. Biol. Chem. 274: 13836–13840
Meiss G, Scholz SR, Korn C, Gimadutdinow O and Pingoud A (2001) Identification of functionally relevant histidine residues in the apoptotic nuclease CAD. Nucleic Acids Res. 29: 3901–3909
Korn C, Scholz SR, Gimadutdinow O, Pingoud A and Meiss G (2002) Involvement of conserved histidine, lysine and tyrosine residues in the mechanism of DNA cleavage by the caspase-3 activated DNase CAD. Nucleic Acids Res. 30: 1325–1332
Durrieu F, Samejima K, Fortune JM, Kandels-Lewis S, Osheroff N and Earnshaw WC (2000) DNA topoisomerase IIalpha interacts with CAD nuclease and is involved in chromatin condensation during apoptotic execution. Curr. Biol. 10: 923–926
Sakahira H, Enari M and Nagata S (1998) Cleavage of CAD inhibitor in CAD activation and DNA degradation during apoptosis. Nature 391: 96–99
Gu J, Dong RP, Zhang C, McLaughlin DF, Wu MX and Schlossman SF (1999) Functional interaction of DFF35 and DFF45 with caspase-activated DNA fragmentation nuclease DFF40. J. Biol. Chem. 274: 20759–20762
Sakahira H and Nagata S (2002) Co-translational folding of caspase-activated DNase with Hsp70, Hsp40 and inhibitor of caspase-activated DNase. J. Biol. Chem. 277: 3364–3370
Zhang J, Liu X, Scherer DC, van Kaer L, Wang X and Xu M (1998) Resistance to DNA fragmentation and chromatin condensation in mice lacking the DNA fragmentation factor 45. Proc. Natl. Acad. Sci. USA 95: 12480–12485
Mukae N, Yokoyama H, Yokokura T, Sakoyama Y, Sakahira H and Nagata S (2000) Identification and developmental expression of inhibitor of caspase-activated DNase (ICAD) in Drosophila melanogaster. J. Biol. Chem. 275: 21402–21408
Chen D, Stetler RA, Cao G, Pei W, O'Horo C, Yin XM and Chen J (2000) Characterization of the rat DNA fragmentation factor 35/Inhibitor of caspase-activated DNase (Short form). The endogenous inhibitor of caspase-dependent DNA fragmentation in neuronal apoptosis. J. Biol. Chem. 275: 38508–38517
McIlroy D, Sakahira H, Talanian RV and Nagata S (1999) Involvement of caspase 3-activated DNase in internucleosomal DNA cleavage induced by diverse apoptotic stimuli. Oncogene 18: 4401–4408
Wolf BB, Schuler M, Echeverri F and Green DR (1999) Caspase-3 is the primary activator of apoptotic DNA fragmentation via DNA fragmentation factor-45/inhibitor of caspase-activated DNase inactivation. J. Biol. Chem. 274: 30651–30656
McCarty JS, Toh SY and Li P (1999) Study of DFF45 in its role of chaperone and inhibitor: two independent inhibitory domains of DFF40 nuclease activity. Biochem. Biophys. Res. Commun. 264: 176–180
McCarty JS, Toh SY and Li P (1999) Multiple domains of DFF45 bind synergistically to DFF40: roles of caspase cleavage and sequestration of activator domain of DFF40. Biochem. Biophys. Res. Commun. 264: 181–185
Woo M, Hakem R, Soengas MS, Duncan GS, Shahinian A, Kagi D, Hakem A, McCrrach M, Khoo W, Kaufman SA, Senaldi G, Howard T, Lowe SW and Mak TW (1998) Essential contribution of caspase 3/CPP32 to apoptosis and its associated nuclear changes. Genes Dev. 12: 806–819
Tang D and Kidd VJ (1998) Cleavage of DFF-45/ICAD by multiple caspases is essential for its function during apoptosis. J. Biol. Chem. 273: 28549–28552
Thomas DA, Du C, Xu M, Wang X and Ley T (2000) DFF45/ICAD can be directly processed by granzyme B during the induction of apoptosis. Immunity 12: 621–632
Sharif-Askari E, Alam A, Rheaume E, Beresford PJ, Scotto C, Sharma K, Lee D, DeWolf WE, Nuttall ME, Lieberman J and Sekaly RP (2001) Direct cleavage of the human DNA fragmentation factor-45 by granzyme B induces caspase-activated DNase release and DNA fragmentation. EMBO J. 20: 3101–3113
Inohara N, Koseki T, Chen S, Wu X and Nunez G (1998) CIDE, a novel family of cell death activators with homology to the 45 kDa subunit of the DNA fragmentation factor. EMBO J. 17: 2526–2533
Otomo T, Sakahira H, Uegaki K, Nagata S and Yamazaki T (2000) Structure of the heterodimeric complex between CAD domains of CAD and ICAD. Nat. Struct. Biol. 7: 658–662
Uegaki K, Otomo T, Sakahira H, Shimizu M, Yumoto N, Kyogoku Y, Nagata S and Yamazaki T (2000) Structure of the CAD domain of caspase-activated DNase and interaction with the CAD domain of its inhibitor. J. Mol. Biol. 297: 1121–1128
Zhou P, Lugovskoy AA, McCarty JS, Li P and Wagner G (2001) Solution structure of DFF40 and DFF45 N-terminal domain complex and mutual chaperone activity of DFF40 and DFF45. Proc. Natl. Acad. Sci. USA 98: 6051–6055
Samejima K and Earnshaw WC (2000) Differential localization of ICAD-L and ICAD-S in cells due to removal of a C-terminal NLS from ICAD-L by alternative splicing. Exp. Cell Res. 255: 314–320
Lechardeur D, Drzymala L, Sharma M, Zylka D, Kinach R, Pacia J, Hicks C, Usmani N, Rommens JM and Lukacs GL (2000) Determinants of the nuclear localization of the heterodimeric DNA fragmentation factor (ICAD/CAD). J. Cell Biol. 150: 321–334
Zhivotovsky B, Samali A, Gahm A and Orrenius S (1999) Caspases: their intracellular localization and translocation during apoptosis. Cell Death Differ. 6: 644–651
Nagata T, Kishi H, Liu QL, Matsuda T, Imanaka T, Tsukada K, Kang D and Muraguchi A (2002) The regulation of DNAse activities in subcellular compartments of activated thymocytes. Immunology 105: 399–406
Xerri L, Palmerini F, Devilard E, Defrance T, Bouabdallah R, Hassoun J and Birg F (2000) Frequent nuclear localization of ICAD and cytoplasmic co-expression of caspase-8 and caspase-3 in human lymphomas. J. Pathol. 192: 194–202
Samejima K and Earnshaw WC (1998) ICAD/DFF regulator of apoptotic nuclease is nuclear. Exp. Cell Res. 243: 453–459
Rosl F (1992) A simple and rapid method for detection of apoptosis in human cells. Nucleic Acids Res. 20: 5243
Gavrieli Y, Sherman Y and Ben-Sasson SA (1992) Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J. Cell Biol. 119: 493–501
Staley K, Blaschke A and Chun J (1997) Apoptotic DNA fragmentation is detected by a semi-quantitative ligation-mediated PCR of blunt DNA ends. Cell Death Differ. 4: 66–75
Wu YC, Stanfield GM and Horvitz HR (2000) NUC-1, a Caenorhabditis elegans DNase II homolog, functions in an intermediate step of DNA degradation during apoptosis. Genes Dev. 14: 536–548
Kawane K, Fukuyama H, Adachi M, Sakahira H, Copeland NG, Gilbert DJ, Jenkin NA and Nagata S (1999) Structure and promoter analysis of murine CAD and ICAD genes. Cell Death Differ. 6: 745–752
McIlroy D, Tanaka M, Sakahira H, Fukuyama H, Suzuki M, Yamamura K-I, Ohsawa Y and Uchiyama Y, Nagata S (2000) An auxiliary mode of apoptotic DNA fragmentation provided by phagocytes. Genes Dev. 14: 549–558
Boulares AH, Zoltoski AJ, Yakovlev A, Xu M and Smulson ME (2001) Roles of DNA fragmentation factor and poly(ADP-ribose) polymerase in an amplification phase of tumor necrosis factor-induced apoptosis. J. Biol. Chem. 276: 38185–38192
Samejima K, Tone S and Earnshaw WC (2001) CAD/DFF40 nuclease is dispensable for high molecular weight DNA cleavage and stage I chromatin condensation in apoptosis. J. Biol. Chem. 276: 45427–45432
Zhang J, Wang X, Bove K and Xu M (1999) DNA fragmentation factor 45-deficient cells are more resistant to apoptosis and exhibit different dying morphology than wild-type control cells. J. Biol. Chem. 274: 37450–37454
Patterson SD, Spahr CS, Daugas E, Susin SA, Irinopoulou T, Koehler C and Kroemer G (2000) Mass spectrometric identification of proteins released from mitochondria undergoing permeability transition. Cell Death Differ. 7: 137–144
Li LY, Luo X and Wang X (2001) Endonuclease G is an apoptotic DNase when released from mitochondria. Nature 412: 95–99
van Loo G, Schotte P, van Gurp M, Demol H, Hoorelbeke B, Gevaert K, Rodriguez I, Ruiz-Carrillo A, Vandekerckhove J, Declercq W, Beyaert R and Vandenabeele P (2001) Endonuclease G: a mitochondrial protein released in apoptosis and involved in caspase-independent DNA degradation. Cell Death Differ. 8: 1136–1142
Ruiz-Carrillo A and Renaud J (1987) Endonuclease G: a (dG)n X (dC)n-specific DNase from higher eukaryotes. EMBO J. 6: 401–407
Widlak P, Li LY, Wang X and Garrard WT (2001) Action of recombinant human apoptotic endonuclease G on naked DNA and chromatin substrates: cooperation with exonuclease and DNase I. J. Biol. Chem. 276: 48404–48409
Parrish J, Li L, Klotz K, Ledwich D, Wang X and Xue D (2001) Mitochondrial endonuclease G is important for apoptosis in C. elegans. Nature 412: 90–94
Sakahira H, Enari M, Ohsawa Y, Uchiyama Y, Nagata S (1999) Apoptotic nuclear morphological change without DNA fragmentation. Curr. Biol. 9: 543–546
Zhang J, Lee H, Lou DW, Bovin GP and Xu M (2000) Lack of obvious 50 kilobase pair DNA fragments in DNA fragmentation factor 45-deficient thymocytes upon activation of apoptosis. Biochem. Biophys. Res. Commun. 274: 225–229
Susin SA, Lorenzo HK, Zamzami N, Marzo I, Snow BE, Brothers GM, Mangion J, Jacotot E, Costantini P, Loeffler M, Larochette N, Goodlett DR, Aebersold R, Siderovski DP, Penninger JM and Kroemer G (1999) Molecular characterization of mitochondrial apoptosis-inducing factor. Nature 397: 441–446
Susin SA, Daugas E, Ravagnan L, Samejima K, Zamzami N, Loeffler M, Costantini P, Ferri KF, Irinopoulou T, Prevost MC, Brothers G, Mak TW, Penninger J, Earnshaw WC and Kroemer G (2000) Two distinct pathways leading to nuclear apoptosis. J. Exp. Med. 192: 571–580
Stroh C and Schulze-Osthoff K (1998) Death by a thousand cuts: an ever increasing list of caspase substrates. Cell Death Differ. 5: 997–1000
Zhang J, Lee H, Agarwala A, Wen Lou D and Xu M (2001) DNA fragmentation factor 45 mutant mice exhibit resistance to kainic acid-induced neuronal cell death. Biochem. Biophys. Res. Commun. 285: 1143–1149
Slane JM, Lee HS, Vorhees CV, Zhang J and Xu M (2000) DNA fragmentation factor 45 deficient mice exhibit enhanced spatial learning and memory compared to wild-type control mice. Brain Res. 867: 70–79
Yakovlev AG, Di X, Movsesyan V, Mullins PG, Wang G, Boulares H, Zhang J, Xu M and Faden AI (2001) Presence of DNA fragmentation and lack of neuroprotective effect in DFF45 knockout mice subjected to traumatic brain injury. Mol. Med. 7: 205–216
Odaka C and Mizuochi T (2002) Macrophages are involved in DNA degradation of apoptotic cells in murine thymus after administration of hydrocortisone. Cell Death Differ. 9: 104–112
Hanayama R, Tanaka M, Miwa K, Shinohara A, Iwamatsu A and Nagata S (2002) Identification of a factor that links apoptotic cells to phagocytes. Nature 417: 182–187
Bernardi G (1971) Spleen acid deoxyribonuclease. In The Enzymes Boyer PD, ed. New York and London: Academic Press, pp 271–287
Kawane K, Fukuyama H, Kondoh G, Takeda J, Ohsawa Y, Uchiyama Y and Nagata S (2001) Requirement of DNase II for definitive erythropoiesis in the mouse fetal liver. Science 292: 1546–1549
Krieser RJ, MacLea KS, Longnecker DS, Fields JL, Fiering S and Eastman A (2002) Deoxyribonuclease IIa is required during the phagocytic phase of apoptosis and its loss causes lethality. Cell Death Differ. 9: 956–962
Hedgecock EM, Sulston JE and Thomson JN (1983) Mutations affecting programmed cell deaths in the nematode Caenorhabditis elegans. Science 220: 1277–1279
Odaka C and Mizuochi T (1999) Role of macrophage lysosomal enzymes in the degradation of nucleosomes of apoptotic cells. J. Immunol. 163: 5346–5352
Leadbetter EA, Rifkin IR, Hohlbaum AM, Beaudette BC, Shlomchik MJ and Marshak-Rothstein A (2002) Chromatin–IgG complexes activate B cells by dual engagement of IgM and Toll-like receptors. Nature 416: 603–607
Aliprantis AO, Diez-Roux G, Mulder LC, Zychlinsky A and Lang RA (1996) Do macrophages kill through apoptosis? Immunol. Today 17: 573–576
Diez-Roux G and Lang RA (1997) Macrophages induce apoptosis in normal cells in vivo. Development 124: 3633–3638
Hoeppner DJ, Hengartner MO and Schnabel R (2001) Engulfment genes cooperate with ced-3 to promote cell death in Caenorhabditis elegans. Nature 412: 202–206
Reddien PW, Cameron S and Horvitz HR (2001) Phagocytosis promotes programmed cell death in C. elegans. Nature 412: 198–202
Cao G, Pei W, Lan J, Stetler RA, Luo Y, Nagayama T, Graham SH, Yin XM, Simon RP and Chen J (2001) Caspase-activated DNase/DNA fragmentation factor 40 mediates apoptotic DNA fragmentation in transient cerebral ischemia and in neuronal cultures. J. Neurosci. 21: 4678–4690
Nagase H, Fukuyama H, Tanaka M, Kawane K and Nagata S (2002) Mutually regulated expression of caspase-activated DNase and its inhibitor for apoptotic DNA fragmentation. Cell Death Differ. (in press).
Mukae N, Yokoyama H, Yokokura T, Sakoyama Y and Nagata S (2002) Activation of the innate immunity in Drosophila by endogenous chromosomal DNA that escaped apoptotic degradation. Genes & Develop 16: 2662–2671
Acknowledgements
The work in our laboratory was supported in part by Grants-in-aid from the Ministry of Education, Science, Sports, and Culture in Japan. KK and NM were supported by a Research fellowship of the Japan Society for the Promotion of Science. We thank Drs. L Migletta and G Gray of Clarity Editing for the careful editing of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by M Hengartner
Rights and permissions
About this article
Cite this article
Nagata, S., Nagase, H., Kawane, K. et al. Degradation of chromosomal DNA during apoptosis. Cell Death Differ 10, 108–116 (2003). https://doi.org/10.1038/sj.cdd.4401161
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401161