Abstract
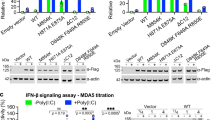
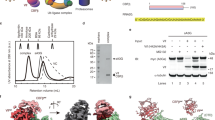
RIG-I is a cytosolic helicase that senses 5′-ppp RNA contained in negative-strand RNA viruses and triggers innate antiviral immune responses. Calorimetric binding studies established that the RIG-I C-terminal regulatory domain (CTD) binds to blunt-end double-stranded 5′-ppp RNA a factor of 17 more tightly than to its single-stranded counterpart. Here we report on the crystal structure of RIG-I CTD bound to both blunt ends of a self-complementary 5′-ppp dsRNA 12-mer, with interactions involving 5′-pp clearly visible in the complex. The structure, supported by mutation studies, defines how a lysine-rich basic cleft within the RIG-I CTD sequesters the observable 5′-pp of the bound RNA, with a stacked phenylalanine capping the terminal base pair. Key intermolecular interactions observed in the crystalline state are retained in the complex of 5′-ppp dsRNA 24-mer and full-length RIG-I under in vivo conditions, as evaluated from the impact of binding pocket RIG-I mutations and 2′-OCH3 RNA modifications on the interferon response.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Yoneyama, M. & Fujita, T. Recognition of viral nucleic acids in innate immunity. Rev. Med. Virol. 20, 4–22 (2010).
Wilkins, C. & Gale, M. Jr. Recognition of viruses by cytoplasmic sensors. Curr. Opin. Immunol. 22, 41–47 (2010).
Rehwinkel, J. & Reis e Sousa, C. RIGorous detection: exposing virus through RNA sensing. Science 327, 284–286 (2010).
Coch, C. et al. Higher activation of TLR9 in plasmacytoid dendritic cells by microbial DNA compared with self-DNA based on CpG-specific recognition of phosphodiester DNA. J. Leukoc. Biol. 86, 663–670 (2009).
Poeck, H. et al. 5′-triphosphate-siRNA: turning gene silencing and Rig-I activation against melanoma. Nat. Med. 14, 1256–1263 (2008).
Barchet, W., Wimmenauer, V., Schlee, M. & Hartmann, G. Accessing the therapeutic potential of immunostimulatory nucleic acids. Curr. Opin. Immunol. 20, 389–395 (2008).
Barral, P.M. et al. Functions of the cytoplasmic RNA sensors RIG-I and MDA-5: key regulators of innate immunity. Pharmacol. Ther. 124, 219–234 (2009).
Takeuchi, O. & Akira, S. Innate immunity to virus infection. Immunol. Rev. 227, 75–86 (2009).
Yoneyama, M. & Fujita, T. RNA recognition and signal transduction by RIG-I-like receptors. Immunol. Rev. 227, 54–65 (2009).
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in dsRNA-induced innate antiviral response. Nat. Immunol. 5, 730–737 (2004).
Besch, R. et al. Proapoptotic signaling induced by RIG-I and MDA-5 results in type I interferon-independent apoptosis in human melanoma cells. J. Clin. Invest. 119, 2399–2411 (2009).
Poeck, H. et al. Recognition of RNA virus by RIG-I results in activation of CARD9 and inflammasome signaling for interleukin 1 β production. Nat. Immunol. 11, 63–69 (2010).
Hornung, V. et al. 5′-triphosphate RNA is the ligand for RIG-I. Science 314, 994–997 (2006).
Pichlmair, A. et al. RIG-I mediated antiviral responds to single-stranded RNA bearing 5′-phosphates. Science 314, 997–1001 (2006).
Schmidt, A. et al. 5′-triphosphate RNA requires base-paired structures to activate antiviral signaling via RIG-I. Proc. Natl. Acad. Sci. USA 106, 12067–12072 (2009).
Schlee, M. et al. Recognition of the 5′-phosphate by RIG-I helicase requires short blunt dsRNA as contained in the panhandle of negative-strand virus. Immunity 31, 25–34 (2009).
Ablasser, A. et al. RIG-I dependent sensing of poly(dA:dT) through the induction of an RNA pol III-transcribed RNA intermediate. Nat. Immunol. 10, 1065–1072 (2009).
Chiu, Y.H. et al. RNA pol III detects cytosolic DNA and induces type I interferons through the RIG-I pathway. Cell 138, 576–591 (2009).
Myong, S. et al. Cytosolic viral sensor RIG-I is a 5′-triphosphate-dependent translocase on dsRNA. Science 323, 1070–1074 (2009).
Takahasi, K. et al. Non-self RNA-sensing mechanism of RNA-I helicase and activation of antiviral immune responses. Mol. Cell 29, 428–440 (2008).
Cui, S. et al. The C-terminal regulatory domain is the RNA 5′-triphosphate sensor of RIG-I. Mol. Cell 29, 169–179 (2008).
Ma, J.-B., Ye, K. & Patel, D.J. Structural basis for overhang-specific small interfering RNA recognition by the PAZ domain. Nature 429, 318–322 (2004).
Ma, J.B. et al. Structural basis for 5′-end-specific recognition of the guide RNA strand by the A. fugidus PIWI protein. Nature 434, 666–670 (2005).
Teplova, M. et al. Structural basis for recognition and sequestration of UUUOH 3′-terminii of nascent mRNA polymerase III transcripts by La autoantigen. Mol. Cell 21, 75–85 (2006).
Li, X. et al. The RIG-I-like receptor LGP2 recognizes the termini of dsRNA. J. Biol. Chem. 284, 13881–13891 (2009).
Leung, D.W. et al. Structural basis for dsRNA recognition and interferon antagonism by Ebola VP35. Nat. Struct. Mol. Biol. 17, 165–172 (2010).
Liu, L. et al. Structural basis of Toll-like receptor 3 signaling with double-stranded RNA. Science 320, 379–381 (2008).
Utyanskaya, E.Z., Lidskii, B.V., Neihaus, M.G. & Shilov, A.E. Mathematical modeling of kinetics of adenosine 5′-triphosphate hydrolysis catalyzed by Zn2+ ion in the pH range 7.1 to 7.4. J. Inorg. Biochem. 81, 239–258 (2000).
Ludwig, J. & Eckstein, F. Rapid and efficient synthesis of nucleoside 5′-O-(1-thiotriphosphates), 5′-triphosphates and 2′,3′-cyclophosphorothioates using 2-chloro-4h-1,3,2-benzodioxaphosphorin-4-one. J. Org. Chem. 54, 631–635 (1989).
Anderson, A.C. et al. HPLC purification of RNA for crystallography and NMR. RNA 2, 110–117 (1996).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Acknowledgements
The authors are grateful to the staff of X-29 beamline at Brookhaven National Laboratory for their assistance during synchrotron data collection, D. Shimanshu and Y. Tian for their assistance in synchrotron data collection, M. Aigner for chemical synthesis of 5′-ppp RNAs undertaken in Innsbruck and G. Wardle for MS analysis in the RU proteomics facility to quality control the 5′-ppp RNAs. D.J.P. was supported by funds from the Abby Rockefeller Mauze Trust and the Maloris Foundation. R.M. was supported by the Austrian Science fund FWF (I317). G.H. was supported by grants from the Bundesministerium für Bildung und Forschung Biofuture and GoBio and from the Deutsche Forschungsgemeinschaft (SFB704, SFB670, SFB832 and KFO177).
Author information
Authors and Affiliations
Contributions
Y.W. expressed the RIG-I CTD domain, with G.S. assisting with protein expression and purification; Y.W. crystallized and solved the structure of the complex; J.L. was responsible for chemical synthesis of palindromic and non-palindromic 5′-ppp RNAs and 2′-O-methyl-modified 5′-ppp RNAs; C.S., M.G. and M.S. were responsible for the in vivo functional assays, including expression of full-length RIG-I mutants; H.L. and Y.W. were responsible for ITC titration assays; S.J. was responsible for gel-shift assays and prepared the plasmids for mutant RIG-I CTD expression; R.M. provided earlier batches of short 5′-ppp RNAs for crystallization trials; the paper was written by D.J.P., G.H. and T.T. with the assistance of the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–3 and Supplementary Figures 1–4 (PDF 592 kb)
Rights and permissions
About this article
Cite this article
Wang, Y., Ludwig, J., Schuberth, C. et al. Structural and functional insights into 5′-ppp RNA pattern recognition by the innate immune receptor RIG-I. Nat Struct Mol Biol 17, 781–787 (2010). https://doi.org/10.1038/nsmb.1863
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1863
This article is cited by
-
mRNA ageing shapes the Cap2 methylome in mammalian mRNA
Nature (2023)
-
A non-canonical, interferon-independent signaling activity of cGAMP triggers DNA damage response signaling
Nature Communications (2021)
-
Multiple truncated isoforms of MAVS prevent its spontaneous aggregation in antiviral innate immune signalling
Nature Communications (2017)
-
Ube2D3 and Ube2N are essential for RIG-I-mediated MAVS aggregation in antiviral innate immunity
Nature Communications (2017)
-
Discriminating self from non-self in nucleic acid sensing
Nature Reviews Immunology (2016)