Abstract
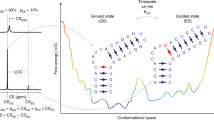
Protein structure is inherently dynamic, with function often predicated on excursions from low to higher energy conformations. For example, X-ray studies of a cavity mutant of T4 lysozyme, L99A, show that the cavity is sterically inaccessible to ligand, yet the protein is able to bind substituted benzenes rapidly. We have used novel relaxation dispersion NMR techniques to kinetically and thermodynamically characterize a transition between a highly populated (97%, 25 °C) ground state conformation and an excited state that is 2.0 kcal mol−1 higher in free energy. A temperature-dependent study of the rates of interconversion between ground and excited states allows the separation of the free energy change into enthalpic (ΔH = 7.1 kcal mol−1) and entropic (TΔS = 5.1 kcal mol−1, 25 °C) components. The residues involved cluster about the cavity, providing evidence that the excited state facilitates ligand entry.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Zhang, X.-J., Wozniak, J.A. & Matthews, B.W. J. Mol. Biol. 250, 527–552 (1995).
Morton, A. & Matthews, B.W. Biochemistry 34, 8576–8588 (1995).
Feher, V.A., Baldwin, E.P. & Dahlquist, F.W. Nature Struct. Biol. 3, 516–521 (1996).
Eriksson, A.E., Baase, W.A., Wozniak, J.A. & Matthews, B.W. Nature 355, 371–373 (1992).
Mulder, F.A.A., Hon, B., Muhandiram, D.R., Dahlquist, F.W. & Kay, L.E. Biochemistry 39, 12614–12622 (2000).
Goto, N.K., Skrynnikov, N.R., Dahlquist, F.W. & Kay, L.E. J. Mol. Biol. 308, 745–764 (2001).
Palmer, A.G., Kroenke, C.D. & Loria, J.P. Methods Enzymol. 339, 204–238 (2001).
Carver, J.P. & Richards, R.E. J. Magn. Reson. 6, 89–105 (1972).
Loria, J.P., Rance, M. & Palmer, A.G. J. Am. Chem. Soc. 121, 2331–2332 (1999).
Skrynnikov, N.R., Mulder, F.A.A., Hon, B., Dahlquist, F.W. & Kay, L.E. J. Am. Chem. Soc. 123, 4556–4566 (2001).
Rosen, M.K. et al. J. Mol. Biol. 263, 627–636 (1996).
Lee, A.L., Urbauer, J.L. & Wand, A.J. J. Biomol. NMR 9, 437–440 (1997).
Allerhand, A. & Thiele, E. J. Chem. Phys. 45, 902–916 (1966).
Ferry, J.D., Grandine, L.D. & Fitzgerald, E.R. J. Appl. Phys. 24, 911–921 (1953).
Denisov, V.P., Peters, J., Horlein, H.D. & Halle, B. Nature Struct. Biol. 3, 505–509 (1996).
Johnson, C.E. & Bovey, F.A. J. Chem. Phys. 29, 1012–1014 (1958).
Hard, T., Barnes, H.J., Larsson, C., Gustafsson, J.A. & Lund, J. Nature Struct. Biol. 2, 983–989 (1995).
Ernst, W.A. et al. Immunity 8, 331–340 (1998).
Cistola, D.P., Kim, K., Rogl, H. & Frieden, C. Biochemistry 35, 7559–7565 (1996).
Zheleznova, E.E., Markham, P.N., Neyfakh, A.A. & Brennan, R.G. Cell 96, 353–362 (1999).
Chen, X. et al. J. Mol. Biol. 278, 641–653 (1998).
Gee, A.C. & Katzenellenbogen, J.A. Mol. Endocr. 15, 421–428 (2001).
Luz, Z. & Meiboom, S. J. Chem. Phys. 39, 366–370 (1963).
Millet, O., Loria, J.P., Kroenke, C.D., Pons, M. & Palmer, A.G. J. Am. Chem. Soc. 122, 2867–2877 (2000).
Acknowledgements
This work was support by grants from the Medical Research Council of Canada (L.E.K.) and the National Institutes of Health (F.W.D.). L.E.K. is a foreign investigator of the Howard Hughes Medical Research Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mulder, F., Mittermaier, A., Hon, B. et al. Studying excited states of proteins by NMR spectroscopy. Nat Struct Mol Biol 8, 932–935 (2001). https://doi.org/10.1038/nsb1101-932
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb1101-932
This article is cited by
-
Application of High-Pressure Electron Paramagnetic Resonance (EPR) Spectroscopy in Protein Science
Applied Magnetic Resonance (2024)
-
Bound nucleotide can control the dynamic architecture of monomeric actin
Nature Structural & Molecular Biology (2022)
-
A litmus test for classifying recognition mechanisms of transiently binding proteins
Nature Communications (2022)
-
A quantitative model predicts how m6A reshapes the kinetic landscape of nucleic acid hybridization and conformational transitions
Nature Communications (2021)
-
Mutate-and-chemical-shift-fingerprint (MCSF) to characterize excited states in RNA using NMR spectroscopy
Nature Protocols (2021)