Key Points
-
One of the most enduring puzzles in evolutionary biology is why sexual reproduction, whereby the genomes of different individuals are mixed together, is so widespread.
-
Theoretical analyses have shown that genome mixing is a risky endeavour, and the conditions that favour the evolution of high rates of sex and recombination are often quite restrictive.
-
Most previous mathematical models have, however, focused on populations that are infinite in size, unstructured and isolated from other species.
-
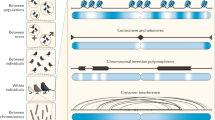
By incorporating realistic complexities, including species interactions, spatial structure and genetic drift, recent studies have broadened the conditions under which high rates of sex and recombination are expected to evolve.
Abstract
Sexual reproduction and recombination are ubiquitous. However, a large body of theoretical work has shown that these processes should only evolve under a restricted set of conditions. New studies indicate that this discrepancy might result from the fact that previous models have ignored important complexities that face natural populations, such as genetic drift and the spatial structure of populations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Chapman, T., Liddle, L. F., Kalb, J. M., Wolfner, M. F. & Partridge, L. Cost of mating in Drosophila melanogaster females is mediated by male accessory gland products. Nature 373, 241–244 (1995).
Redfield, R. J. Do bacteria have sex? Nature Rev. Genet. 2, 634–639 (2001).
Karlin, S. Detecting anomalous gene clusters and pathogenicity islands in diverse bacterial genomes. Trends Microbiol. 9, 335–343 (2001).
Kidwell, M. G. Lateral transfer in natural populations of eukaryotes. Annu. Rev. Genet. 27, 235–256 (1993).
de Koning, A. P., Brinkman, F. S., Jones, S. J. & Keeling, P. J. Lateral gene transfer and metabolic adaptation in the human parasite Trichomonas vaginalis. Mol. Biol. Evol. 17, 1769–1773 (2000).
Wolf, Y. I., Kondrashov, A. S. & Koonin, E. V. Interkingdom gene fusions. Genome Biol. 1, research0013.1–0013.13 (2000).
Bell, G. The Masterpiece of Nature: The Evolution and Genetics of Sexuality (Univ. California Press, Berkeley, 1982).A classic text that explores the vast array of ways in which organisms reproduce and the reasons for this diversity.
Vrijenhoek, R., Dawley, R., Cole, C. & Bogart, J. in Evolution and Cytology of Unisexual Vertebrates (eds Dawley, R. & Bogart, J.) 19–23 (Univ. State New York, New York, 1989).
Wilson, E. O. The Diversity of Life (W. W. Norton, New York, 1992).
Judson, O. P. & Normark, B. B. Ancient asexual scandals. Trends Ecol. Evol. 11, 41–46 (1996).
Mark Welch, D. & Meselson, M. Evidence for the evolution of bdelloid rotifers without sexual reproduction or genetic exchange. Science 288, 1211–1215 (2000).
Sogin, M. L. & Silberman, J. D. Evolution of the protists and protistan parasites from the perspective of molecular systematics. Int. J. Parasitol. 28, 11–20 (1998).
Morin, L. Long branch attraction effects and the status of 'basal eukaryotes': phylogeny and structural analysis of the ribosomal RNA gene cluster of the free-living diplomonad Trepomonas agilis. J. Eukaryot. Microbiol. 47, 167–177 (2000).
Hurst, L. D., Hamilton, W. D. & Ladle, R. J. Covert sex. Trends Ecol. Evol. 7, 144–145 (1992).
Burt, A., Carter, D. A., Koenig, G. L., White, T. J. & Taylor, J. W. Molecular markers reveal cryptic sex in the human pathogen Coccidioides immitis. Proc. Natl Acad. Sci. USA 93, 770–773 (1996).
Michod, R. E. & Levin, B. R. The Evolution of Sex (Sinauer Press, Sunderland, Massachusetts, 1988).An excellent review and source book for theoretical and empirical studies of the evolution of sex, with chapters written by authors from a variety of perspectives.
Kondrashov, A. S. Classification of hypotheses on the advantage of amphimixis. J. Hered. 84, 372–387 (1993).
Feldman, M. W., Otto, S. P. & Christiansen, F. B. Population genetic perspectives on the evolution of recombination. Annu. Rev. Genet. 30, 261–295 (1997).
Hickey, D. A. Selfish DNA: a sexually-transmitted nuclear parasite. Genetics 101, 519–531 (1982).
Reinke, V. et al. A global profile of germline gene expression in C. elegans. Mol. Cell 6, 605–616 (2000).
Bernstein, H., Byers, G. S. & Michod, R. E. Evolution of sexual reproduction: importance of DNA repair, complementation, and variation. Am. Nat. 117, 537–549 (1981).
Redfield, R. J. Evolution of natural transformation: testing the DNA repair hypothesis in Bacillus subtilis and Haemophilus influenzae. Genetics 133, 755–761 (1993).
Thuriaux, P. Is recombination confined to structural genes on the eukaryotic genome? Nature 268, 460–462 (1977).
Baker, B. S., Carpenter, A. T. C., Esposito, M. S., Esposito, R. E. & Sandler, L. The genetic control of meiosis. Annu. Rev. Genet. 10, 53–134 (1976).
Hawley, R. S. & Theurkauf, W. E. Requiem for distributive segregation: achiasmate segregation in Drosophila females. Trends Genet. 9, 310–317 (1993).
Merriam, J. R. & Frost, J. N. Exchange and nondisjunction of the X chromosomes in female Drosophila melanogaster. Genetics 49, 109–122 (1964).
Villeneuve, A. M. A cis-acting locus that promotes crossing over between X chromosomes in Caenorhabditis elegans. Genetics 136, 887–902 (1994).
Koehler, K. E., Hawley, R. S., Sherman, S. & Hassold, T. Recombination and nondisjunction in humans and flies. Hum. Mol. Genet. 5, 1495–1504 (1996).
Burt, A., Bell, G. & Harvey, P. H. Sex differences in recombination. J. Evol. Biol. 4, 259–277 (1991).
Otto, S. P. & Barton, N. H. Selection for recombination in small populations. Evolution 55, 1921–1931 (2001).An investigation of the relative importance of drift and epistasis to the evolution of sex and recombination with the use of a modifier model with directional selection.
Rice, W. R. Experimental tests of the adaptive significance of sexual reproduction. Nature Rev. Genet. 3, 241–251 (2002).
Altenberg, L. & Feldman, M. W. Selection, generalized transmission and the evolution of modifier genes. I. The reduction principle. Genetics 117, 559–572 (1987).
Feldman, M. W. Selection for linkage modification. I. Random mating populations. Theor. Popul. Biol. 21, 430–439 (1972).
Feldman, M. W., Christiansen, F. B. & Brooks, L. D. Evolution of recombination in a constant environment. Proc. Natl Acad. Sci. USA 77, 4838–4841 (1980).The first theoretical analysis to show that higher rates of recombination could evolve when deleterious mutations exhibit negative fitness interactions.
Kondrashov, A. S. Deleterious mutations as an evolutionary factor. I. The advantage of recombination. Genet. Res. 44, 199–217 (1984).
Charlesworth, B. Mutation–selection balance and the evolutionary advantage of sex and recombination. Genet. Res. 55, 199–221 (1990).
Barton, N. H. A general model for the evolution of recombination. Genet. Res. 65, 123–144 (1995).A ground-breaking theoretical study that generalized models of the evolution of recombination to a genome-wide level and to include several forms of selection.
Otto, S. P. & Feldman, M. W. Deleterious mutations, variable epistatic interactions, and the evolution of recombination. Theor. Popul. Biol. 51, 134–147 (1997).
Eshel, I. & Feldman, M. W. On the evolutionary effect of recombination. Theor. Popul. Biol. 1, 88–100 (1970).
Chasnov, J. R. Mutation–selection balance, dominance and the maintenance of sex. Genetics 156, 1419–1425 (2000).
Agrawal, A. F. & Chasnov, J. R. Recessive mutations and the maintenance of sex in structured populations. Genetics 158, 913–917 (2001).
Simmons, M. J. & Crow, J. F. Mutations affecting fitness in Drosophila populations. Annu. Rev. Genet. 11, 49–78 (1977).
Peters, A. D. & Lively, C. M. The Red Queen and fluctuating epistasis: a population genetic analysis of antagonistic coevolution. Am. Nat. 154, 393–405 (1999).
Lenormand, T. & Otto, S. P. The evolution of recombination in a heterogeneous environment. Genetics 156, 423–438 (2000).
Pylkov, K. V., Zhivotovsky, L. A. & Feldman, M. W. Migration versus mutation in the evolution of recombination under multilocus selection. Genet. Res. 71, 247–256 (1998).
Hill, W. G. & Robertson, A. The effect of linkage on the limits to artificial selection. Genet. Res. 8, 269–294 (1966).This paper showed that, in finite populations, selection at a locus is less efficient when neighboring loci are also under selection because of increased variation in the reproductive success of an allele (that is, increased random genetic drift).
Otto, S. P. & Barton, N. H. The evolution of recombination: removing the limits to natural selection. Genetics 147, 879–906 (1997).
Gessler, D. D. & Xu, S. On the evolution of recombination and meiosis. Genet. Res. 73, 119–131 (1999).
Peck, J. R. A ruby in the rubbish: beneficial mutations, deleterious mutations and the evolution of sex. Genetics 137, 597–606 (1994).
Martin, G. Évolution de la Recombinaison en Populations Subdivisées: Étude par Simulation de l'Influence de la Structure sur la Sélection pour la Recombinaison via l'Effet Hill–Robertson (DEA, Biologié, Université Montpellier II, Montpellier, 2001).
Foucault, M. The History of Sexuality. I. An Introduction (Vintage Books, New York, 1978).
Howard, R. S. & Lively, C. M. The maintenance of sex by parasitism and mutation accumulation under epistatic fitness functions. Evolution 52, 604–610 (1998).
Nei, M. Modification of linkage intensity by natural selection. Genetics 57, 625–641 (1967).
Acknowledgements
The authors gratefully acknowledge the comments and suggestions of J. Boughman, C. Griswold, T. Johnson, R. Redfield, H. Rundle, R. Sargent, D. Schluter, M. Whitlock and three anonymous reviewers. Funding was provided by the Natural Sciences and Engineering Research Council (Canada) to S.P.O. and the Centre National de la Recherche Scientifique (France) to S.P.O. and T. L.
Author information
Authors and Affiliations
Corresponding author
Related links
Glossary
- TRANSDUCTION
-
The exchange of genetic material from one cell to another that is mediated by a virus or phage.
- TRANSFORMATION
-
The uptake of DNA by a bacterium from the surrounding environment.
- CONJUGATION
-
In prokaryotes, the transfer of DNA from a donor cell to a recipient cell that is mediated by direct cell–cell contact.
- PROTIST
-
A eukaryote other than animals, plants and fungi; often single celled.
- CHIASMA
-
(pl. chiasmata). The cytological manifestation of genetic exchange between chromosomes, indicating that a crossover has occurred between homologous chromosomes.
- ANEUPLOIDY
-
The presence of extra copies, or fewer copies, of some chromosomes.
- EQUILIBRIUM
-
A state in which a system remains unchanged over time.
- LINKAGE DISEQUILIBRIUM
-
(D). A measure of genetic associations between alleles at different loci, which indicates whether particular haplotypes are more common than expected. We use the two-locus measure, D = frequency(AB) × frequency(ab) − frequency(Ab) × frequency(aB).
- HAPLOTYPE
-
A haploid genotype. A diploid genotype comprises a maternal and a paternal haplotype.
- GENETIC DRIFT
-
(also known as random drift). A phenomenon whereby the frequency of a gene in a population changes over time because the number of offspring born to parents that carry the gene is subject to chance variation.
- EPISTASIS
-
(ɛ). A measure of fitness interactions between alleles at different loci. In haploids, we use the two-locus measure, ɛ = fitness(AB) × fitness(ab) − fitness(Ab) × fitness(aB).
- MUTATION–SELECTION BALANCE
-
The equilibrium at which selection that increases the frequency of a favourable allele exactly balances mutations that decrease the frequency.
- SELECTION COEFFICIENT
-
A term that describes the difference in average fitness between two genotypes when fitness is measured relative to the average fitness of one of the genotypes (known as the reference genotype).
- DIRECTIONAL SELECTION
-
Selection that favours one allele over all other alleles of a gene. The frequency of this beneficial allele can rise or can be held in check by recurrent mutation.
- RECOMBINATION LOAD
-
The difference in fitness between offspring produced without recombination and those produced with recombination.
- HARDY–WEINBERG EQUILIBRIUM
-
A state in which the frequency of each diploid genotype at a locus equals that expected from the random union of alleles — that is, where the inbreeding coefficient (F) is zero.
- INBREEDING COEFFICIENT
-
(F). A measure of genetic associations between alleles at the same locus, which indicates whether homozygotes (positive F) or heterozygotes (negative F) are more common than expected. For ease of comparison with linkage disequilibrium, we write the standard measure for F as {frequency(AA) × frequency(aa) − [0.5 × frequency(Aa)]2}/{frequency(A) × frequency(a)}.
- DOMINANCE COEFFICIENT
-
The factor by which the selection coefficient is reduced in heterozygotes relative to homozygotes.
- SINGLE-LOCUS INTERACTION
-
(ı). A measure of fitness interactions between alleles at the same locus. We use the single-locus measure, ı = fitness(AA) × fitness(aa) − fitness(Aa)2.
Rights and permissions
About this article
Cite this article
Otto, S., Lenormand, T. Resolving the paradox of sex and recombination. Nat Rev Genet 3, 252–261 (2002). https://doi.org/10.1038/nrg761
Issue Date:
DOI: https://doi.org/10.1038/nrg761
This article is cited by
-
Genetic variation in released gametes produces genetic diversity in the offspring of the broadcast spawning coral Acropora tenuis
Scientific Reports (2022)
-
The origination events of gametic sexual reproduction and anisogamy
Journal of Ethology (2022)
-
Predicting global numbers of teleomorphic ascomycetes
Fungal Diversity (2022)
-
Sex determination mechanisms and sex control approaches in aquaculture animals
Science China Life Sciences (2022)
-
Starvation-induced cell fusion and heterokaryosis frequently escape imperfect allorecognition systems in an asexual fungal pathogen
BMC Biology (2021)