Abstract
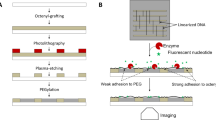
Biological molecules that self-assemble and interact with other molecules are attractive building blocks for engineering biological devices. DNA has been widely used for the creation of nanomaterials1, but the use of proteins remains largely unexplored. Here, we show that clathrin can form homogeneous and extended two-dimensional lattices on a variety of substrates, including glass, metal, carbon and plastic. Clathrin is a three-legged protein complex with unique self-assembling properties and is relevant in the formation of membrane transport vesicles in eukaryotic cells2,3. We used a fragment of the adaptor protein epsin to immobilize clathrin lattices on the substrates. The lattices span multiple square millimetres with a regular periodicity of 30 nm and can be functionalized via modified subunits of clathrin with either inorganic nanoparticles or active enzymes. The lattices can be stored for months after crosslinking and stabilization with uranyl acetate. They could be dehydrated and rehydrated without loss of function, offering potential applications in sensing and as biosynthetic reactors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Seeman, N. C. Nanomaterials based on DNA. Annu. Rev. Biochem. 79, 65–87 (2010).
Brodsky, F. M. Diversity of clathrin function: new tricks for an old protein. Annu. Rev. Cell Dev. Biol. 28, 309–336 (2012).
Pearse, B. M. Clathrin: a unique protein associated with intracellular transfer of membrane by coated vesicles. Proc. Natl Acad. Sci. USA 73, 1255–1259 (1976).
Goodman, R. P. et al. Rapid chiral assembly of rigid DNA building blocks for molecular nanofabrication. Science 310, 1661–1665 (2005).
Rothemund, P. W. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Selmi, D. N. et al. DNA-templated protein arrays for single-molecule imaging. Nano Lett. 11, 657–660 (2011).
Steinhauer, C., Jungmann, R., Sobey, T. L., Simmel, F. C. & Tinnefeld, P. DNA origami as a nanoscopic ruler for super-resolution microscopy. Angew. Chem. Int. Ed. 48, 8870–8873 (2009).
Douglas, T. & Young, M. Host–guest encapsulation of materials by assembled virus protein cages. Nature 393, 152–155 (1998).
Graveland-Bikker, J. F., Schaap, I. A., Schmidt, C. F. & de Kruif, C. G. Structural and mechanical study of a self-assembling protein nanotube. Nano Lett. 6, 616–621 (2006).
Sinclair, J. C., Davies, K. M., Venien-Bryan, C. & Noble, M. E. Generation of protein lattices by fusing proteins with matching rotational symmetry. Nature Nanotech. 6, 558–562 (2011).
Sleytr, U. B., Egelseer, E. M., Ilk, N., Pum, D. & Schuster, B. S-Layers as a basic building block in a molecular construction kit. FEBS J. 274, 323–334 (2007).
Yang, R. et al. Self-assembly of ferritin nanocages into linear chains induced by poly(α, L-lysine). Chem. Commun. 50, 481–483 (2014).
Ungewickell, E. & Branton, D. Assembly units of clathrin coats. Nature 289, 420–422 (1981).
Dannhauser, P. N. & Ungewickell, E. J. Reconstitution of clathrin-coated bud and vesicle formation with minimal components. Nature Cell Biol. 14, 634–639 (2012).
Saffarian, S., Cocucci, E. & Kirchhausen, T. Distinct dynamics of endocytic clathrin-coated pits and coated plaques. PLoS Biol. 7, e1000191 (2009).
Dannhauser, P. N. et al. Effect of clathrin light chains on the stiffness of clathrin lattices and membrane budding. Traffic 16, 519–533 (2015).
Nandi, P. K. & Edelhoch, H. The effects of lyotropic (Hofmeister) salts on the stability of clathrin coat structure in coated vesicles and baskets. J. Biol. Chem. 259, 11290–11296 (1984).
Hoffmann, A. et al. A comparison of GFP-tagged clathrin light chains with fluorochromated light chains in vivo and in vitro. Traffic 11, 1129–1140 (2010).
Wilbur, J. D. et al. Conformation switching of clathrin light chain regulates clathrin lattice assembly. Dev. Cell 18, 841–848 (2010).
Hom, N. et al. Anisotropic nanocrystal arrays organized on protein lattices formed by recombinant clathrin fragments. J. Mater. Chem. 22, 23335–23339 (2012).
Schoen, A. P., Schoen, D. T., Huggins, K. N., Arunagirinathan, M. A. & Heilshorn, S. C. Template engineering through epitope recognition: a modular, biomimetic strategy for inorganic nanomaterial synthesis. J. Am. Chem. Soc. 133, 18202–18207 (2011).
Fotin, A. et al. Molecular model for a complete clathrin lattice from electron cryomicroscopy. Nature 432, 573–579 (2004).
Holstein, S. E., Ungewickell, H. & Ungewickell, E. Mechanism of clathrin basket dissociation: separate functions of protein domains of the DnaJ homologue auxilin. J. Cell Biol. 135, 925–937 (1996).
Kalthoff, C., Alves, J., Urbanke, C., Knorr, R. & Ungewickell, E. J. Unusual structural organization of the endocytic proteins AP180 and epsin 1. J. Biol. Chem. 277, 8209–8216 (2002).
Keen, J. H., Willingham, M. C. & Pastan, I. H. Clathrin-coated vesicles: isolation, dissociation and factor-dependent reassociation of clathrin baskets. Cell 16, 303–312 (1979).
Ungewickell, E. & Ungewickell, H. Bovine brain clathrin light chains impede heavy chain assembly in vitro. J. Biol. Chem. 266, 12710–12714 (1991).
Winkler, F. K. & Stanley, K. K. Clathrin heavy chain, light chain interactions. EMBO J. 2, 1393–1400 (1983).
Hinrichsen, L., Meyerholz, A., Groos, S. & Ungewickell, E. J. Bending a membrane: how clathrin affects budding. Proc. Natl Acad. Sci. USA 103, 8715–8720 (2006).
Burnham, N. A. et al. Comparison of calibration methods for atomic-force microscopy cantilevers. Nanotechnology 14, 1–6 (2003).
Ortega-Esteban, A. et al. Minimizing tip–sample forces in jumping mode atomic force microscopy in liquid. Ultramicroscopy 114, 56–61 (2012).
Erhardt, J. & Dirr, H. Native dimer stabilizes the subunit tertiary structure of porcine class pi glutathione S-transferase. Eur. J. Biochem. 230, 614–620 (1995).
Schlossman, D. M., Schmid, S. L., Braell, W. A. & Rothman, J. E. An enzyme that removes clathrin coats: purification of an uncoating ATPase. J. Cell Biol. 99, 723–733 (1984).
Acknowledgements
The authors thank H. Ungewickell, G. Preiss and C. Lemke for expert technical assistance, the MHH core facility ‘Confocal Laser Microscopy’ for instrumental support and M. Breyvogel for the CAD model. The AFM instrumentation was funded through the ‘Cluster of Excellence and DFG Research Centre Nanoscale Microscopy and Molecular Physiology of the Brain’. M.P. thanks the Göttingen Graduate School for Neurosciences, Biophysics and Molecular Biosciences (GGNB) for a scholarship. All authors are grateful for the support of E.J. Ungewickell regarding protein purification, ideas and discussions.
Author information
Authors and Affiliations
Contributions
P.N.D. and H.B. prepared the samples. P.N.D. performed the electron and fluorescence microscopy experiments and analysed data. M.P. performed the AFM experiments and analysed data. P.N.D., M.P. and I.A.T.S. designed the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
A patent application has been filed by P.N. Dannhauser and E.J. Ungewickell (EP2812029 A1) that includes aspects of the procedure used for the coating of surfaces. So far, the application has exclusively proved to be of scientific value, with no assignable monetary value.
Supplementary information
Supplementary information
Supplementary information (PDF 1113 kb)
Rights and permissions
About this article
Cite this article
Dannhauser, P., Platen, M., Böning, H. et al. Durable protein lattices of clathrin that can be functionalized with nanoparticles and active biomolecules. Nature Nanotech 10, 954–957 (2015). https://doi.org/10.1038/nnano.2015.206
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2015.206
This article is cited by
-
Dynamical Majorana edge modes in a broad class of topological mechanical systems
Nature Communications (2017)
-
Concentration Dependent Ion-Protein Interaction Patterns Underlying Protein Oligomerization Behaviours
Scientific Reports (2016)