Abstract
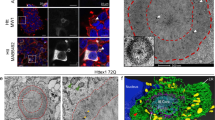
Post-translational modification by the lipid palmitate is crucial for the correct targeting and function of many proteins. Here we show that huntingtin (htt) is normally palmitoylated at cysteine 214, which is essential for its trafficking and function. The palmitoylation and distribution of htt are regulated by the palmitoyl transferase huntingtin interacting protein 14 (HIP14). Expansion of the polyglutamine tract of htt, which causes Huntington disease, results in reduced interaction between mutant htt and HIP14 and consequently in a marked reduction in palmitoylation. Mutation of the palmitoylation site of htt, making it palmitoylation resistant, accelerates inclusion formation and increases neuronal toxicity. Downregulation of HIP14 in mouse neurons expressing wild-type and mutant htt increases inclusion formation, whereas overexpression of HIP14 substantially reduces inclusions. These results suggest that the expansion of the polyglutamine tract in htt results in decreased palmitoylation, which contributes to the formation of inclusion bodies and enhanced neuronal toxicity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
El-Husseini, A.E. et al. Dual palmitoylation of PSD-95 mediates its vesiculotubular sorting, postsynaptic targeting, and ion channel clustering. J. Cell Biol. 148, 159–172 (2000).
El-Husseini, A. & Bredt, D.S. Protein palmitoylation: a regulator of neuronal development and function. Nat. Rev. Neurosci. 3, 791–802 (2002).
Huang, K. & El-Husseini, A. Modulation of neuronal protein trafficking and function by palmitoylation. Curr. Opin. Neurobiol. 15, 527–535 (2005).
Huang, K. et al. Huntingtin-interacting protein HIP14 is a palmitoyl transferase involved in palmitoylation and trafficking of multiple neuronal proteins. Neuron 44, 977–986 (2004).
The Huntington's Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 72, 971–983 (1993).
Hackam, A.S. et al. The influence of huntingtin protein size on nuclear localization and cellular toxicity. J. Cell Biol. 141, 1097–1105 (1998).
Wheeler, V.C. et al. Early phenotypes that presage late-onset neurodegenerative disease allow testing of modifiers in Hdh CAG knock-in mice. Hum. Mol. Genet. 11, 633–640 (2002).
Van Raamsdonk, J.M., Murphy, Z., Slow, E.J., Leavitt, B.R. & Hayden, M.R. Selective degeneration and nuclear localization of mutant huntingtin in the YAC128 mouse model of Huntington disease. Hum. Mol. Genet. 14, 3823–3835 (2005).
Velier, J. et al. Wild-type and mutant huntingtins function in vesicle trafficking in the secretory and endocytic pathways. Exp. Neurol. 152, 34–40 (1998).
DiFiglia, M. et al. Huntingtin is a cytoplasmic protein associated with vesicles in human and rat brain neurons. Neuron 14, 1075–1081 (1995).
Kegel, K.B. et al. Huntingtin associates with acidic phospholipids at the plasma membrane. J. Biol. Chem. 280, 36464–36473 (2005).
Kalchman, M.A. et al. Huntingtin is ubiquitinated and interacts with a specific ubiquitin-conjugating enzyme. J. Biol. Chem. 271, 19385–19394 (1996).
Steffan, J.S. et al. SUMO modification of Huntingtin and Huntington's disease pathology. Science 304, 100–104 (2004).
Humbert, S. et al. The IGF-1/Akt pathway is neuroprotective in Huntington's disease and involves Huntingtin phosphorylation by Akt. Dev. Cell 2, 831–837 (2002).
Warby, S.C. et al. Huntingtin phosphorylation on serine 421 is significantly reduced in the striatum and by polyglutamine expansion in vivo. Hum. Mol. Genet. 14, 1569–1577 (2005).
Kopito, R.R. Aggresomes, inclusion bodies and protein aggregation. Trends Cell Biol. 10, 524–530 (2000).
Waelter, S. et al. Accumulation of mutant huntingtin fragments in aggresome-like inclusion bodies as a result of insufficient protein degradation. Mol. Biol. Cell 12, 1393–1407 (2001).
Wyttenbach, A. et al. Heat shock protein 27 prevents cellular polyglutamine toxicity and suppresses the increase of reactive oxygen species caused by huntingtin. Hum. Mol. Genet. 11, 1137–1151 (2002).
Sapp, E. et al. Huntingtin localization in brains of normal and Huntington's disease patients. Ann. Neurol. 42, 604–612 (1997).
Zeron, M.M. et al. Increased sensitivity to N-methyl-D-aspartate receptor-mediated excitotoxicity in a mouse model of Huntington's disease. Neuron 33, 849–860 (2002).
Singaraja, R.R. et al. HIP14, a novel ankyrin domain-containing protein, links huntingtin to intracellular trafficking and endocytosis. Hum. Mol. Genet. 11, 2815–2828 (2002).
Slow, E.J. et al. Selective striatal neuronal loss in a YAC128 mouse model of Huntington disease. Hum. Mol. Genet. 12, 1555–1567 (2003).
Drisdel, R.C. & Green, W.N. Labeling and quantifying sites of protein palmitoylation. Biotechniques 36, 276–285 (2004).
Drisdel, R.C., Manzana, E. & Green, W.N. The role of palmitoylation in functional expression of nicotinic alpha7 receptors. J. Neurosci. 24, 10502–10510 (2004).
El-Husseini, A. et al. Synaptic strength regulated by palmitate cycling on PSD-95. Cell 108, 849–863 (2002).
Rakhilin, S. et al. Alpha-bungarotoxin receptors contain alpha7 subunits in two different disulfide-bonded conformations. J. Cell Biol. 146, 203–218 (1999).
Rocks, O. et al. An acylation cycle regulates localization and activity of palmitoylated Ras isoforms. Science 307, 1746–1752 (2005).
Drenan, R.M. et al. Palmitoylation regulates plasma membrane-nuclear shuttling of R7BP, a novel membrane anchor for the RGS7 family. J. Cell Biol. 169, 623–633 (2005).
Ferrante, R.J. et al. Histone deacetylase inhibition by sodium butyrate chemotherapy ameliorates the neurodegenerative phenotype in Huntington's disease mice. J. Neurosci. 23, 9418–9427 (2003).
Mastroberardino, P.G. et al. 'Tissue' transglutaminase ablation reduces neuronal death and prolongs survival in a mouse model of Huntington's disease. Cell Death Differ. 9, 873–880 (2002).
Slow, E.J. et al. Absence of behavioral abnormalities and neurodegeneration in vivo despite widespread neuronal huntingtin inclusions. Proc. Natl. Acad. Sci. USA 102, 11402–11407 (2005).
Arrasate, M., Mitra, S., Schweitzer, E.S., Segal, M.R. & Finkbeiner, S. Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 431, 805–810 (2004).
Soyombo, A.A. & Hofmann, S.L. Molecular cloning and expression of palmitoyl-protein thioesterase 2 (PPT2), a homolog of lysosomal palmitoyl-protein thioesterase with a distinct substrate specificity. J. Biol. Chem. 272, 27456–27463 (1997).
Verkruyse, L.A. & Hofmann, S.L. Lysosomal targeting of palmitoyl-protein thioesterase. J. Biol. Chem. 271, 15831–15836 (1996).
Wellington, C.L. et al. Caspase cleavage of gene products associated with triplet expansion disorders generates truncated fragments containing the polyglutamine tract. J. Biol. Chem. 273, 9158–9167 (1998).
Janas, J., Skowronski, J. & Van, A.L. Lentiviral delivery of RNAi in hippocampal neurons. Methods Enzymol. 406, 593–605 (2006).
Acknowledgements
We thank O. Sadiq and E. Eyu for technical assistance. M.R.H. is supported by grants from the Canadian Institutes for Health Research, the Huntington Disease Society of America, the Jack and Doris Brown Foundation and the Huntington Society of Canada. A.E.H. is supported by grants from the Canadian Institutes for Health Research, the EJLB foundation and Neuroscience Canada. R.C.D. and W.N.G. are supported by funding from the National Institute of Neurological Diseases and Stroke, the National Institute of Drug Abuse and the Alzheimer's Association. D.C.R. is funded by a Wellcome Trust Senior Fellowship in Clinical Science, an M.R.C. programme grant and E.U. Framework VI (EUROSCA). A.Y. is supported by funding from the Michael Smith Foundation for Health Research. KH is supported by University Graduate Fellowship and HighQ. M.R.H. and A.E.H. are supported by funding from the HighQ foundation and are investigators of the Fundamental Innovation in Neurodegenerative Diseases (FIND) Research Infrastructure Unit, funded by the Michael Smith Foundation for Health Research. M.R.H. holds a Canada Research Chair in Human Genetics and is a University Killam Professor.
Author information
Authors and Affiliations
Contributions
A.Y. performed all DNA manipulations to generate truncated and full-length mutant htt and HIP14 proteins, and performed most of the [3H]palmitoylation assays and data analyses. K.H. conducted all the Btn-BMCC palmitoylation assays for full-length htt, scored cells for inclusions in htt-transfected COS cells and neurons, designed and characterized HIP14 siRNA, performed all experiments (and corresponding data analysis) in which the alteration of htt trafficking was investigated by knocking down HIP14, conducted the virus production and infection experiments, and scored for TUNEL assay. R.K. performed endogenous htt [3H]palmitoylation assay, scored inclusions in neurons transfected with full-length htt, and performed fragment-htt toxicity assay and the corresponding data analysis. R.R.S. and L.G. performed the htt and HIP14 coimmunoprecipitation experiment. P.C.O. generated HIP14 siRNA lentiviral constructs and supervised the virus production experiment. P.A. conducted the time-lapse experiments. A.M. assisted in the characterization of full-length htt palmitoylation. C.M.C. and L.A.R. were involved in developing the NMDA-induced excitotoxicity TUNEL assay. R.K. and K.H. conducted the TUNEL assays. W.N.G. and R.C.D. developed and supervised the Btn-BMCC palmitoylation assays. B.R. and D.C.R. performed toxicity assays in COS cells. M.R.H., A.E.H., A.Y. and K.H. wrote the manuscript. M.R.H. and A.E.H. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 2
Altered distribution of palmitoylation-resistant htt in HEK-293 cells. (PDF 1609 kb)
Rights and permissions
About this article
Cite this article
Yanai, A., Huang, K., Kang, R. et al. Palmitoylation of huntingtin by HIP14is essential for its trafficking and function. Nat Neurosci 9, 824–831 (2006). https://doi.org/10.1038/nn1702
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn1702
This article is cited by
-
Protein acylation: mechanisms, biological functions and therapeutic targets
Signal Transduction and Targeted Therapy (2022)
-
Roles of palmitoylation in structural long-term synaptic plasticity
Molecular Brain (2021)
-
Protein expression of prenyltransferase subunits in postmortem schizophrenia dorsolateral prefrontal cortex
Translational Psychiatry (2020)
-
Palmitoylation-mediated synaptic regulation of AMPA receptor trafficking and function
Archives of Pharmacal Research (2019)
-
Therapeutic approaches to Huntington disease: from the bench to the clinic
Nature Reviews Drug Discovery (2018)