Abstract
The rate of translation of egg-laying hormone (ELH) in Aplysia californica bag cell neurons rises after an afterdischarge (AD), the physiological trigger for egg-laying1. We found that the 5′ untranslated region (5′ UTR) of ELH possessed an internal ribosomal entry site (IRES), and that an AD was accompanied by a switch to cap-independent, IRES-mediated translation, indicating that the use of a cellular IRES can be regulated by physiological activity. Furthermore, the AD caused dephosphorylation of the initiation factor eIF4E, which was sufficient to increase use of the ELH IRES, showing that dephosphorylation of eIF4E can trigger a switch to IRES-mediated translation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Lee, W. & Wayne, N.L. Endocrinology 139, 5109–5115 (1998).
Mahon, A.C. et al. J. Neurosci. 5, 1872–1880 (1985).
Pyronnet, S., Dostie, J. & Sonenberg, N. Genes Dev. 15, 2083–2093 (2001).
Scheper, G.C. et al. J. Biol. Chem. 277, 3303–3309 (2002).
Lachance, P.E.D., Miron, M., Raught, B., Sonenberg, N. & Lasko, P. Mol. Cell. Biol. 22, 1656–1663 (2002).
Dyer, J.R. & Sossin, W.S. J. Neurochem. 75, 872–881 (2000).
Ghirardi, M., Montarolo, P.G. & Kandel, E.R. Neuron 14, 413–420 (1995).
Yanow, S.K., Manseau, F., Hislop, J., Castellucci, V.F. & Sossin, W.S. J. Neurochem. 70, 572–583 (1998).
Pinkstaff, J.K., Chappell, S.A., Mauro, V.P., Edelman, G.M. & Krushel, L.A. Proc. Natl. Acad. Sci. USA 98, 2770–2775 (2001).
Manseau, F., Fan, X., Hueftlein, T., Sossin, W.S. & Castellucci, V.F. J. Neurosci. 21, 1247–1256 (2001).
Acknowledgements
This work is supported by Canadian Institute of Health Research (CIHR) grant MOP-15121 to W.S.S., CIHR grant MOP-14142 to V.F.C. and grants from the National Institutes of Health (R01 NS 33548) and the Whitehall Foundation (F98-35) to N.L.W. W.S.S. is a Killiam Scholar and a CIHR investigator. We thank P. McPherson and N. Sonenberg for comments on an earlier version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1.
LF has a significantly increased EGFP/EBFP ratio. The EGFP/EBFP ratios were calculated for expressing cells (defined by having maximal EBFP fluorescence >20% over background) from several experiments (n ≥ 3 cells/experiment: LF, n=7 experiments, 59 cells; LR, n=6 experiments, 49 cells; SF, n=5 experiments, 39 cells; SR, n=6 experiments, 41 cells; BxG, n=4 experiments, 18 cells). There was a significant difference between the groups, F4,23 =21.1, P < 0.0001. This difference was solely due to LF; LF versus all other groups, Tukey's test P<0.001 (**). (JPG 10 kb)
Supplementary Fig. 2.
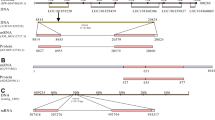
Green fluorescence is not due to a cryptic promoter or alternative splicing. (a) The position of the RNA probe for RNA protection assays (initial PCR was performed with a 5' primer in EBFP (GGCAGTGGACTCATCAGCTCGGCATGGACGAGCTGTA) and a 3' primer in EGFP (CTGCACTCAGCTGAGTGTGAACAGCTCCTCGCCCTT)), the positions of primers for the RT (ACCAGACCAACTGGTAATGGTA) and PCR reactions (5' primer, AACGCCATTTGACCATTCACCA; 3' primer A, ACCGGCGCTCAGCTGGAAT; 3' primer B, GTAGGCTTTGGCTTGTTGGAAA). (b) Examination of LF mRNAs by RNA protection assays. Dilutions of the probe alone showed the assay's sensitivity and the undigested probe's position. Only a single band can be seen in the protected lane. Molecular weight markers were biotinlyated RNA standards (Ambion, Austin, TX, USA). (c) Examination of LF mRNAs by RT-PCR. RT-PCR using either PCR primers A or B described in (a) demonstrated the presence of the expected LF band (large band), a smaller band (*) that is an alternative spliced form (see text) and from PCR A several PCR artifacts (o) due to mispriming at alternative sites. No bands were seen when the reverse transcriptase was omitted (no RT). Linearized LF vector (Standards, approximately 3x104, 1x105, 1x106 or 3x106 molecules of plasmid) was the positive control. M (molecular weight markers) were 100 BP+ (MBI-Fermentas, Flamborough, ON, Canada). Upper band is 3000 followed by 2000, 1500, 1200, 1031, 900 and then spaced at 100 bp intervals. (JPG 19 kb)
Supplementary Fig. 3.
LF, but not LR injected bag cell neurons express EGFP. This experiment was repeated three times with similar results. (JPG 12 kb)
Supplementary Fig. 4.
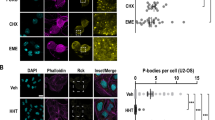
Expression of non-phosphorylatable eIF4E ST-AA depressed eIF4E phosphorylation in sensory cells. Three days after injecting cells with plasmids encoding EGFP and non-phosphorylatable eIF4E ST-AA (a) or EGFP and wild-type eIF4E (b), they were imaged for EGFP, fixed and stained with the phosphopeptide specific antibody to eIF4E. Cells with high levels of eIF4E ST-AA expression (as monitored by EGFP) had low levels of eIF4E phosphorylation (c) Staining is decreased after injecting with eIF4E ST-AA and EGFP, but not after vector and EGFP or eIF4E and EGFP. Combined data from three experiments: vector, n=20 cells; eIF4E, n=21 cells; eIF4EST-AA, n=29 cells; mean ± SEM; *, P<0.001 versus either of the two other groups, two-tailed Students t-test. (d) Phosphorylation of eIF4E in individual cells (anti-eIF4E-P staining) was negatively correlated (r2>0.5, P<0.01) with expression of eIF4E ST-AA (as monitored by EGFP expression) in injected sensory neurons. (JPG 52 kb)
Supplementary Fig. 5.
Absolute change in fluorescence after injection of eIF4E ST-AA. Within each experiment, the average blue and green fluorescence in the dye and eIF4E ST-AA groups were calculated, then the eIF4E ST-AA average was normalized against the dye average. (JPG 18 kb)
Rights and permissions
About this article
Cite this article
Dyer, J., Michel, S., Lee, W. et al. An activity-dependent switch to cap-independent translation triggered by eIF4E dephosphorylation. Nat Neurosci 6, 219–220 (2003). https://doi.org/10.1038/nn1018
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn1018
This article is cited by
-
Molecular targets for modulating the protein translation vital to proteostasis and neuron degeneration in Parkinson’s disease
Translational Neurodegeneration (2019)
-
RNA binding proteins: a common denominator of neuronal function and dysfunction
Neuroscience Bulletin (2014)
-
Pseudomonas HopU1 modulates plant immune receptor levels by blocking the interaction of their mRNAs with GRP7
The EMBO Journal (2013)
-
Translational control of localized mRNAs: restricting protein synthesis in space and time
Nature Reviews Molecular Cell Biology (2008)
-
Dendritic mRNA: transport, translation and function
Nature Reviews Neuroscience (2007)