Abstract
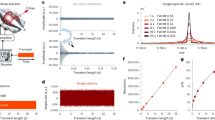
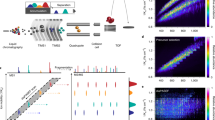
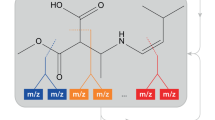
Peptide sequencing is the basis of mass spectrometry–driven proteomics. Here we show that in the linear ion trap–orbitrap mass spectrometer (LTQ Orbitrap) peptide ions can be efficiently fragmented by high-accuracy and full-mass-range tandem mass spectrometry (MS/MS) via higher-energy C-trap dissociation (HCD). Immonium ions generated via HCD pinpoint modifications such as phosphotyrosine with very high confidence. Additionally we show that an added octopole collision cell facilitates de novo sequencing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Aebersold, R. & Mann, M. Nature 422, 198–207 (2003).
Steen, H. & Mann, M. Nat. Rev. Mol. Cell Biol. 5, 699–711 (2004).
Louris, J.N. et al. Anal. Chem. 59, 1677–1685 (1987).
Hu, Q. et al. J. Mass Spectrom. 40, 430–443 (2005).
Makarov, A. et al. Anal. Chem. 78, 2113–2120 (2006).
Olsen, J.V. et al. Mol. Cell. Proteomics 4, 2010–2021 (2005).
Schwartz, J.C., Senko, M.W. & Syka, J.E. J. Am. Soc. Mass Spectrom. 13, 659–669 (2002).
Steen, H., Kuster, B., Fernandez, M., Pandey, A. & Mann, M. Anal. Chem. 73, 1440–1448 (2001).
Ong, S.E. et al. Mol. Cell. Proteomics 1, 376–386 (2002).
Blagoev, B., Ong, S.E., Kratchmarova, I. & Mann, M. Nat. Biotechnol. 22, 1139–1145 (2004).
Perkins, D.N., Pappin, D.J., Creasy, D.M. & Cottrell, J.S. Electrophoresis 20, 3551–3567 (1999).
Wisniewski, J.R., Zougman, A., Kruger, S. & Mann, M. Mol. Cell. Proteomics 6, 72–87 (2007).
Kim, J.Y., Kim, K.W., Kwon, H.J., Lee, D.W. & Yoo, J.S. Anal. Chem. 74, 5443–5449 (2002).
Carr, S.A., Annan, R.S. & Huddleston, M.J. Methods Enzymol. 405, 82–115 (2005).
Mann, M. & Wilm, M.S. Anal. Chem. 66, 4390–4399 (1994).
Acknowledgements
This work was supported by the EU research directorate (Interaction Proteome grant LSHG-CT-2003-505520). We thank other members of the department for Proteomics and Signal Transduction and of Thermo Fisher Scientific, especially R. Pesch and K. Strupat, for constructive comments and discussion.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
O.L., A.M. and S.H are employees of Thermo Fisher, the manufacturer of the LTQ Orbitrap instrument used in this research.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Table 1, Supplementary Methods (PDF 3566 kb)
Rights and permissions
About this article
Cite this article
Olsen, J., Macek, B., Lange, O. et al. Higher-energy C-trap dissociation for peptide modification analysis. Nat Methods 4, 709–712 (2007). https://doi.org/10.1038/nmeth1060
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1060
This article is cited by
-
Prediction of peptide mass spectral libraries with machine learning
Nature Biotechnology (2023)
-
Pentafluorobenzylpyridinium: new thermometer ion for characterizing the ions produced by collisional activation during tandem mass spectrometry
Analytical Sciences (2023)
-
A pathway to peptides in space through the condensation of atomic carbon
Nature Astronomy (2022)
-
Cyclic immonium ion of lactyllysine reveals widespread lactylation in the human proteome
Nature Methods (2022)
-
Glycoproteomics
Nature Reviews Methods Primers (2022)