Abstract
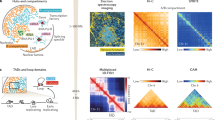
Chromosome conformation capture (3C) and fluorescence in situ hybridization (FISH) are two widely used technologies that provide distinct readouts of 3D chromosome organization. While both technologies can assay locus-specific organization, how to integrate views from 3C, or genome-wide Hi-C, and FISH is far from solved. Contact frequency, measured by Hi-C, and spatial distance, measured by FISH, are often assumed to quantify the same phenomena and used interchangeably. Here, however, we demonstrate that contact frequency is distinct from average spatial distance, both in polymer simulations and in experimental data. Performing a systematic analysis of the technologies, we show that this distinction can create a seemingly paradoxical relationship between 3C and FISH, both in minimal polymer models with dynamic looping interactions and in loop-extrusion simulations. Together, our results indicate that cross-validation of Hi-C and FISH should be carefully designed, and that jointly considering contact frequency and spatial distance is crucial for fully understanding chromosome organization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Dekker, J. & Mirny, L. The 3D genome as moderator of chromosomal communication. Cell 164, 1110–1121 (2016).
Denker, A. & de Laat, W. A long-distance chromatin affair. Cell 162, 942–943 (2015).
Bonev, B. & Cavalli, G. Organization and function of the 3D genome. Nat. Rev. Genet. 17, 661–678 (2016).
Schmitt, A.D., Hu, M. & Ren, B. Genome-wide mapping and analysis of chromosome architecture. Nat. Rev. Mol. Cell Biol. 17, 743–755 (2016).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Imakaev, M.V., Fudenberg, G. & Mirny, L.A. Modeling chromosomes: beyond pretty pictures. FEBS Lett. 589 20 Pt A, 3031–3036 (2015).
Fraser, J., Williamson, I., Bickmore, W.A. & Dostie, J. An overview of genome organization and how we got there: from FISH to Hi-C. Microbiol. Mol. Biol. Rev. 79, 347–372 (2015).
Sachs, R.K., van den Engh, G., Trask, B., Yokota, H. & Hearst, J.E. A random-walk/giant-loop model for interphase chromosomes. Proc. Natl. Acad. Sci. USA 92, 2710–2714 (1995).
Shopland, L.S. et al. Folding and organization of a contiguous chromosome region according to the gene distribution pattern in primary genomic sequence. J. Cell Biol. 174, 27–38 (2006).
Branco, M.R. & Pombo, A. Intermingling of chromosome territories in interphase suggests role in translocations and transcription-dependent associations. PLoS Biol. 4, e138 (2006).
Tanabe, H. et al. Evolutionary conservation of chromosome territory arrangements in cell nuclei from higher primates. Proc. Natl. Acad. Sci. USA 99, 4424–4429 (2002).
Bolzer, A. et al. Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol. 3, e157 (2005).
Shachar, S., Voss, T.C., Pegoraro, G., Sciascia, N. & Misteli, T. Identification of gene positioning factors using high-throughput imaging mapping. Cell 162, 911–923 (2015).
Joyce, E.F., Williams, B.R., Xie, T. & Wu, C.-t. Identification of genes that promote or antagonize somatic homolog pairing using a high-throughput FISH-based screen. PLoS Genet. 8, e1002667 (2012).
Boettiger, A.N. et al. Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature 529, 418–422 (2016).
Beliveau, B.J. et al. Single-molecule super-resolution imaging of chromosomes and in situ haplotype visualization using Oligopaint FISH probes. Nat. Commun. 6, 7147 (2015).
Fabre, P.J. et al. Nanoscale spatial organization of the HoxD gene cluster in distinct transcriptional states. Proc. Natl. Acad. Sci. USA 112, 13964–13969 (2015).
Giorgetti, L. & Heard, E. Closing the loop: 3C versus DNA FISH. Genome Biol. 17, 215 (2016).
Hakim, O. et al. Diverse gene reprogramming events occur in the same spatial clusters of distal regulatory elements. Genome Res. 21, 697–706 (2011).
Giorgetti, L. et al. Predictive polymer modeling reveals coupled fluctuations in chromosome conformation and transcription. Cell 157, 950–963 (2014).
Rao, S.S.P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Nora, E.P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Dixon, J.R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Wang, S. et al. Spatial organization of chromatin domains and compartments in single chromosomes. Science 353, 598–602 (2016).
Williamson, I. et al. Spatial genome organization: contrasting views from chromosome conformation capture and fluorescence in situ hybridization. Genes Dev. 28, 2778–2791 (2014).
Jost, D., Vaillant, C. & Meister, P. Coupling 1D modifications and 3D nuclear organization: data, models and function. Curr. Opin. Cell Biol. 44, 20–27 (2017).
Doyle, B., Fudenberg, G., Imakaev, M. & Mirny, L.A. Chromatin loops as allosteric modulators of enhancer-promoter interactions. PLoS Comput. Biol. 10, e1003867 (2014).
Eastman, P. et al. OpenMM 4: A reusable, extensible, hardware independent library for high performance molecular simulation. J. Chem. Theory Comput. 9, 461–469 (2013).
Eastman, P. et al. OpenMM 7: rapid development of high performance algorithms for molecular dynamics. Preprint at http://biorxiv.org/content/early/2016/12/06/091801 (2016).
Hofmann, A. & Heermann, D.W. The role of loops on the order of eukaryotes and prokaryotes. FEBS Lett. 589 20 Pt A, 2958–2965 (2015).
Gavrilov, A.A. et al. Disclosure of a structural milieu for the proximity ligation reveals the elusive nature of an active chromatin hub. Nucleic Acids Res. 41, 3563–3575 (2013).
Belmont, A.S. Large-scale chromatin organization: the good, the surprising, and the still perplexing. Curr. Opin. Cell Biol. 26, 69–78 (2014).
Nagano, T. et al. Comparison of Hi-C results using in-solution versus in-nucleus ligation. Genome Biol. 16, 175 (2015).
Lajoie, B.R., Dekker, J. & Kaplan, N. The Hitchhiker's guide to Hi-C analysis: practical guidelines. Methods 72, 65–75 (2015).
Hsieh, T.S., Fudenberg, G., Goloborodko, A. & Rando, O.J. Micro-C XL: assaying chromosome conformation from the nucleosome to the entire genome. Nat. Methods 13, 1009–1011 (2016).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Nichols, M.H. & Corces, V.G. A CTCF code for 3D genome architecture. Cell 162, 703–705 (2015).
Sanborn, A.L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl. Acad. Sci. USA 112, E6456–E6465 (2015).
Hansen, A.S., Pustova, I., Cattoglio, C., Tjian, R. & Darzacq, X. CTCF and cohesin regulate chromatin loop stability with distinct dynamics. eLife http://dx.doi.org/10.7554/eLife.25776 (2017).
Nora, E.P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Barrington, C., Finn, R. & Hadjur, S. Cohesin biology meets the loop extrusion model. Chromosom. Res. 25, 51–60 (2017).
Schwarzer, W. et al. Two independent modes of chromosome organization are revealed by cohesin removal. Preprint at http://biorxiv.org/content/early/2016/12/15/094185 (2016).
Brackley, C.A. et al. Non-equilibrium chromosome looping via molecular slip-links. Preprint at https://arxiv.org/abs/1612.07256 (2016).
Goloborodko, A., Marko, J.F. & Mirny, L.A. Chromosome compaction by active loop extrusion. Biophys. J. 110, 2162–2168 (2016).
Imakaev, M.V., Tchourine, K.M., Nechaev, S.K. & Mirny, L.A. Effects of topological constraints on globular polymers. Soft Matter 11, 665–671 (2015).
Halverson, J.D., Smrek, J., Kremer, K. & Grosberg, A.Y. From a melt of rings to chromosome territories: the role of topological constraints in genome folding. Rep. Prog. Phys. 77, 022601 (2014).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Acknowledgements
The authors thank A. Goloborodko for thoughtful comments regarding the statistical mechanics analogy of re-weighting loop conformations, G. Nir for helpful discussions regarding imaging, and anonymous reviewers for thoughtful and detailed feedback. The authors also thank J. Dekker and other members of the UMass–MIT Center for 3D Structure and Physics of the Genome for helpful feedback. Finally, the authors thank L. Mirny for comments on earlier drafts of this paper and for supporting their independent work. This work was supported by NSF 1504942 Physics of Chromosomes (PI: L. Mirny) and U54 DK107980 3D Structure and Physics of the Genome (PIs: J. Dekker and L. Mirny). During revisions, G.F. was supported by the San Simeon Fund (PI: K. Pollard).
Author information
Authors and Affiliations
Contributions
G.F. and M.I. jointly conceived of the study, performed analyses, and wrote the manuscript, and are listed in alphabetical order on the first page.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–3 and Supplementary Note (PDF 5037 kb)
Rights and permissions
About this article
Cite this article
Fudenberg, G., Imakaev, M. FISH-ing for captured contacts: towards reconciling FISH and 3C. Nat Methods 14, 673–678 (2017). https://doi.org/10.1038/nmeth.4329
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4329
This article is cited by
-
Increased enhancer–promoter interactions during developmental enhancer activation in mammals
Nature Genetics (2024)
-
Enhancer selectivity in space and time: from enhancer–promoter interactions to promoter activation
Nature Reviews Molecular Cell Biology (2024)
-
RNA polymerase II dynamics shape enhancer–promoter interactions
Nature Genetics (2023)
-
The spatial organization of transcriptional control
Nature Reviews Genetics (2023)
-
Sex-specific multi-level 3D genome dynamics in the mouse brain
Nature Communications (2022)