Abstract
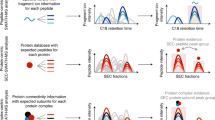
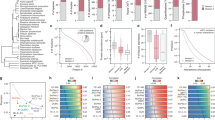
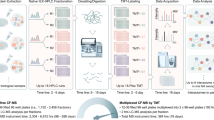
We present a mass spectrometry–based strategy for the absolute quantification of protein complex components isolated through affinity purification. We quantified bait proteins via isotope-labeled reference peptides corresponding to an affinity tag sequence and prey proteins by label-free correlational quantification using the precursor ion signal intensities of proteotypic peptides generated in reciprocal purifications. We used this method to quantitatively analyze interaction stoichiometries in the human protein phosphatase 2A network.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Gingras, A.C. et al. Nat. Rev. Mol. Cell Biol. 8, 645–654 (2007).
Kocher, T. & Superti-Furga, G. Nat. Methods 4, 807–815 (2007).
Bouwmeester, T. et al. Nat. Cell Biol. 6, 97–105 (2004).
Ewing, R.M. et al. Mol. Syst. Biol. 3, 89 (2007).
Jeronimo, C. et al. Mol. Cell 27, 262–274 (2007).
Desiderio, D.M. & Kai, M. Biomed. Mass Spectrom. 10, 471–479 (1983).
Gerber, S.A. et al. Proc. Natl. Acad. Sci. USA 100, 6940–6945 (2003).
Mallick, P. et al. Nat. Biotechnol. 25, 125–131 (2007).
Martens, L. et al. Proteomics 5, 3537–3545 (2005).
Deutsch, E.W., Lam, H. & Aebersold, R. EMBO Rep. 9, 429–434 (2008).
Schmidt, T.G. & Skerra, A. Nat. Protoc. 2, 1528–1535 (2007).
Virshup, D.M. Curr. Opin. Cell Biol. 12, 180–185 (2000).
Janssens, V. & Goris, J. Biochem. J. 353, 417–439 (2001).
Westermarck, J. & Hahn, W.C. Trends Mol. Med. 14, 152–160 (2008).
Xing, Y. et al. Cell 133, 154–163 (2008).
Acknowledgements
We thank O. Rinner and M. Beck for helpful discussions, M. Varjosalo for critically reading the manuscript, and the laboratory of E. Ogris (Max F. Perutz Laboratories) for providing the PPME1 antibody. This project has been funded in part by Eidgenössische Technische Hochschule Zurich, and the US National Heart, Lung, and Blood Institute, National Institutes of Health, under contract N01-HV-28179, and the European Union–funded large integrated FP7 project Systems Biology of T-cell Activation. A.W. and R.A. were supported in part by a grant from F. Hoffmann-La Roche Ltd. provided to the Competence Center for Systems Physiology and Metabolic Disease.
Author information
Authors and Affiliations
Contributions
A.W. performed the experimental and analytical work and wrote the manuscript together with M.G.; T.G. provided the cell lines, and A.S. helped with mass-spectrometry measurements. R.A. and M.G. provided conceptual advice. All authors discussed the results and commented on the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Tables 1–2, Supplementary Results, Supplementary Discussion, Supplementary Methods (PDF 1412 kb)
Rights and permissions
About this article
Cite this article
Wepf, A., Glatter, T., Schmidt, A. et al. Quantitative interaction proteomics using mass spectrometry. Nat Methods 6, 203–205 (2009). https://doi.org/10.1038/nmeth.1302
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1302
This article is cited by
-
DIP-MS: ultra-deep interaction proteomics for the deconvolution of protein complexes
Nature Methods (2024)
-
PME-1 sensitizes glioblastoma cells to oxidative stress-induced cell death by attenuating PP2A-B55α-mediated inactivation of MAPKAPK2-RIPK1 signaling
Cell Death Discovery (2023)
-
An AP-MS- and BioID-compatible MAC-tag enables comprehensive mapping of protein interactions and subcellular localizations
Nature Communications (2018)
-
Stepwise frontal affinity chromatography model for drug and protein interaction
Analytical and Bioanalytical Chemistry (2018)
-
Methylated-antibody affinity purification to improve proteomic identification of plant RNA polymerase Pol V complex and the interacting proteins
Scientific Reports (2017)