Abstract
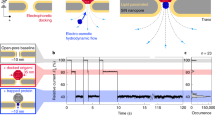
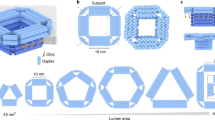
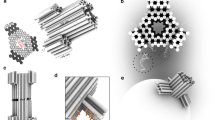
A nanometre-scale pore in a solid-state membrane provides a new way of electronically probing the structure of single linear polymers, including those of biological interest in their native environments. Previous work with biological protein pores wide enough to let through and sense single-stranded DNA molecules demonstrates the power of using nanopores, but many future tasks and applications call for a robust solid-state pore whose nanometre-scale dimensions and properties may be selected, as one selects the lenses of a microscope. Here we demonstrate a solid-state nanopore microscope capable of observing individual molecules of double-stranded DNA and their folding behaviour. We discuss extensions of the nanopore microscope concept to alternative probing mechanisms and applications, including the study of molecular structure and sequencing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Levene, M.J. et al. Zero-mode waveguides for single-molecule analysis at high concentrations. Science 299, 682–686 ( 2003).
Rees, W.A., Keller, R.W., Vesenka, J.P., Yang, G. & Bustamante, C. Evidence of DNA bending in transcription complexes imaged by scanning force microscopy. Science 260, 1646–1649 ( 1993).
Hansma, H.G. et al. Properties of biomolecules measured from atomic force microscope images: A review. J. Struct. Biol. 119, 99–108 ( 1997).
Quake, S.R., Babcock, H. & Chu, S. The dynamics of partially extended single molecules of DNA. Nature 388, 151–154 ( 1997).
Smith, S.B., Cui, Y. & Bustamante, C. Overstretching b-DNA: The elastic response of individual double-stranded and single-stranded DNA molecules. Science 271, 795–799 ( 1996).
Hille, B. Ionic Channels and Excitable Membranes (Sinauer, Sunderland, Massachusetts, 1992).
Bezrukov, S.M., Vodyanoy, I. & Parasegian, V.A. Counting polymers moving through a single ion channel. Nature 370, 279–281 ( 1994).
Kasianowicz, J.J., Brandin, E., Branton, D. & Deamer, D.W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl Acad. Sci. USA 93, 13770–13773 ( 1996).
Akeson, M., Branton, D., Kasianowicz, J.J., Brandin, E. & Deamer, D.W. Microsecond time-scale discrimination among polycytidylic acid, polyadenylic acid, and polyuridylic acid as homopolymers or as segments within single RNA molecules. Biophys. J. 77, 3227–3233 ( 1999).
Mellor, A., Nivon, L., Brandin, E., Golovchenko, J. & Branton, D. Rapid nanopore discrimination between single polynucleotide molecules. Proc. Natl Acad. Sci. USA 97, 1079–1084 ( 2000).
Li, J. et al. Ion-beam sculpting at nanometre length scales. Nature 412, 166–169 ( 2001).
Sauer-Budge, A.F. Unzipping Double-Stranded DNA Molecule by Molecule in a Nanopore Thesis, Harvard Univ. ( 2002).
Muthukumar, M. Dynamics of polyelctrolyte solutions. J. Chem. Phys. 107, 2619–2635 ( 1997).
Muthukumar, M. Polymer translocation through a hole. J. Chem. Phys. 111, 10371–10374 ( 1999).
Muthukumar, M. Polymer escape through a nanopore. J. Chem. Phys. 118, 5174–5184 ( 2003).
Lee, N. & Obukhov, S. Diffusion of a polymer chain through a thin membrane. J. Physique II 6, 195–204 ( 1996).
Evans, D.F. & Wennerström, H. The Colloidal Domain 361 (Wiley-VCH, New York, 1999).
Han, J. & Craighead, H.G. Separation of long DNA molecules in a microfabricated trap array. Science 288, 1026–1029 ( 2000).
Saleh, O.A. & Sohn, L.L. An artificial nanopore for molecular sensing. Nano Lett. 3, 37–38 ( 2003).
Kasianowicz, J.J., Henrickson, S.E., Weetall H.H. & Robertson, B. Simultaneous multianalyte detection with a nanopore. Anal. Chem. 73, 2268–2272 ( 2001).
Vercoutere, W. et al. Rapid discrimination among individual DNA hairpin molecules at single-nucleotide resolution using an ion channel. Nature Biotechnol. 19, 248–252 ( 2001).
Byers, T.J., Husain-Chishti, A., Dubreuil, R.R., Branton, D., & Goldstein, L.S. Sequence similarity of the amino-terminal domain of Drosophila β-spectrin to α-actinin and dystrophin. J. Cell Biol. 109, 1633–1641 ( 1989).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A Laboratory Manual 2nd edn (Cold Spring Harbor Laboratory, New York, 1989).
Acknowledgements
We acknowledge discussion and criticism of this work by D. Branton, M. Muthukumar and M. Aziz. Dr H. Wang prepared the 3-kb DNA used in the work. A. Kavcic, M. Burns, A. Huang and J. Gu assisted with software analysis. Q. Cai assisted with nanopore preparation and C. Russo provided assistance during preparation of this manuscript. Support for this research has been provided by DARPA, NSF, DOE, AFOSR and Agilent Technologies.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Li, J., Gershow, M., Stein, D. et al. DNA molecules and configurations in a solid-state nanopore microscope. Nature Mater 2, 611–615 (2003). https://doi.org/10.1038/nmat965
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat965
This article is cited by
-
On the origins of conductive pulse sensing inside a nanopore
Nature Communications (2022)
-
Nanopore microscope identifies RNA isoforms with structural colours
Nature Chemistry (2022)
-
Effects of symmetric and asymmetric salt conditions on a selective solid-state nanopore assay
Nano Research (2022)
-
Recent advances in properties and applications of nanoporous materials and porous carbons
Carbon Letters (2022)
-
Dynamics of driven polymer transport through a nanopore
Nature Physics (2021)