Abstract
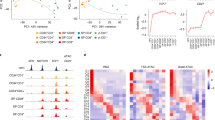
Type 1 helper T (TH1) cells are essential for cellular immunity, but their ontogeny, maturation and durability remain poorly understood. By constructing a dominant-negative form of T-bet, we were able to determine the role played by this lineage-inducing trans-activator in the establishment and maintenance of heritable TH1 gene expression. Optimal induction of interferon-γ (IFN-γ) expression required genetic interaction between T-bet and its target, the homeoprotein Hlx. In fully mature TH1 cells, reiteration of IFN-γ expression and stable chromatin remodeling became relatively independent of T-bet activity and coincided with demethylation of DNA. In contrast, some lineage attributes, such as expression of IL-12Rβ2 (interleukin 12 receptor β2), required ongoing T-bet activity in mature TH1 cells and their progeny. These findings suggest that heritable states of gene expression might be maintained by continued expression of the inducing factor or by a mechanism that confers a stable imprint of the induced state.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Glimcher, L.H. & Murphy, K.M. Lineage commitment in the immune system: the T helper lymphocyte grows up. Genes Dev. 14, 1693–1711 (2000).
Reiner, S.L. Helper T cell differentiation, inside and out. Curr. Opin. Immunol. 13, 351–355 (2001).
Kurata, H., Lee, H.J., O'Garra, A. & Arai, N. Ectopic expression of activated Stat6 induces the expression of Th2-specific cytokines and transcription factors in developing Th1 cells. Immunity 11, 677–688 (1999).
Kurt-Jones, E.A., Hamberg, S., Ohara, J., Paul, W.E. & Abbas, A.K. Heterogeneity of helper/inducer T lymphocytes. I. Lymphokine production and lymphokine responsiveness. J. Exp. Med. 166, 1774–1787 (1987).
Mullen, A.C. et al. Role of T-bet in commitment of TH1 cells before IL-12-dependent selection. Science 292, 1907–1910 (2001).
Mullen, A.C. et al. Cell cycle controlling the silencing and functioning of mammalian activators. Curr. Biol. 11, 1695–1699 (2001).
Kennedy, M.K., Picha, K.S., Shanebeck, K.D., Anderson, D.M. & Grabstein, K.H. Interleukin-12 regulates the proliferation of Th1, but not Th2 or Th0, clones. Eur. J. Immunol. 24, 2271–2278 (1994).
Ouyang, W. et al. Inhibition of Th1 development mediated by GATA-3 through an IL-4-independent mechanism. Immunity 9, 745–755 (1998).
Bird, J.J. et al. Helper T cell differentiation is controlled by the cell cycle. Immunity 9, 229–237 (1998).
Gett, A.V. & Hodgkin, P.D. Cell division regulates the T cell cytokine repertoire, revealing a mechanism underlying immune class regulation. Proc. Natl. Acad. Sci. USA 95, 9488–9493 (1998).
Richter, A., Lohning, M. & Radbruch, A. Instruction for cytokine expression in T helper lymphocytes in relation to proliferation and cell cycle progression. J. Exp. Med. 190, 1439–1450 (1999).
Grogan, J.L. et al. Early transcription and silencing of cytokine genes underlie polarization of T helper cell subsets. Immunity 14, 205–215 (2001).
Tada, M. & Smith, J.C. T-targets: clues to understanding the functions of T-box proteins. Dev. Growth Differ. 43, 1–11 (2001).
Szabo, S.J. et al. A novel transcription factor, T-bet, directs Th1 lineage commitment. Cell 100, 655–669 (2000).
Szabo, S.J. et al. Distinct effects of T-bet in TH1 lineage commitment and IFN-γ production in CD4 and CD8 T cells. Science 295, 338–342 (2002).
Conlon, F.L., Sedgwick, S.G., Weston, K.M. & Smith, J.C. Inhibition of Xbra transcription activation causes defects in mesodermal patterning and reveals autoregulation of Xbra in dorsal mesoderm. Development 122, 2427–2435 (1996).
Horb, M.E. & Thomsen, G.H. Tbx5 is essential for heart development. Development 126, 1739–1751 (1999).
Kispert, A., Koschorz, B. & Herrmann, B.G. The T protein encoded by Brachyury is a tissue-specific transcription factor. EMBO J. 14, 4763–4772 (1995).
Gajewski, T.F., Joyce, J. & Fitch, F.W. Antiproliferative effect of IFN-γ in immune regulation. III. Differential selection of TH1 and TH2 murine helper T lymphocyte clones using recombinant IL-2 and recombinant IFN-γ. J. Immunol. 143, 15–22 (1989).
Izon, D.J. et al. Notch1 regulates maturation of CD4+ and CD8+ thymocytes by modulating TCR signal strength. Immunity 14, 253–264 (2001).
DeKoter, R.P. & Singh, H. Regulation of B lymphocyte and macrophage development by graded expression of PU. 1. Science 288, 1439–1441 (2000).
Agarwal, S. & Rao, A. Modulation of chromatin structure regulates cytokine gene expression during T cell differentiation. Immunity 9, 765–775 (1998).
Solomon, M.J. & Varshavsky, A. A nuclease-hypersensitive region forms de novo after chromosome replication. Mol. Cell. Biol. 7, 3822–3825 (1987).
Wolffe, A.P. & Brown, D.D. DNA replication in vitro erases a Xenopus 5S RNA gene transcription complex. Cell 47, 217–227 (1986).
Lamolet, B. et al. A pituitary cell-restricted T box factor, Tpit, activates POMC transcription in cooperation with Pitx homeoproteins. Cell 104, 849–859 (2001).
Liu, J. et al. Tbx19, a tissue-selective regulator of POMC gene expression. Proc. Natl. Acad. Sci. USA 98, 8674–8679 (2001).
Hiroi, Y. et al. Tbx5 associates with Nkx2-5 and synergistically promotes cardiomyocyte differentiation. Nature Genet. 28, 276–280 (2001).
Bruneau, B.G. et al. A murine model of Holt-Oram syndrome defines roles of the T-box transcription factor Tbx5 in cardiogenesis and disease. Cell 106, 709–721 (2001).
Lipshutz, R.J., Fodor, S.P., Gingeras, T.R. & Lockhart, D.J. High density synthetic oligonucleotide arrays. Nature Genet. 21, 20–24 (1999).
Allen, J.D. et al. Novel murine homeo box gene on chromosome 1 expressed in specific hematopoietic lineages and during embryogenesis. Genes Dev. 5, 509–520 (1991).
Deguchi, Y. et al. Cloning of a human homeobox gene that resembles a diverged Drosophila homeobox gene and is expressed in activated lymphocytes. New Biol. 3, 353–363 (1991).
Hentsch, B. et al. Hlx homeo box gene is essential for an inductive tissue interaction that drives expansion of embryonic liver and gut. Genes Dev. 10, 70–79 (1996).
Kaplan, M.H., Sun, Y.L., Hoey, T. & Grusby, M.J. Impaired IL-12 responses and enhanced development of Th2 cells in Stat4-deficient mice. Nature 382, 174–177 (1996).
Horcher, M., Souabni, A. & Busslinger, M. Pax5/BSAP maintains the identity of B cells in late B lymphopoiesis. Immunity 14, 779–790 (2001).
Tanigaki, K. et al. Notch-RBP-J signaling is involved in cell fate determination of marginal zone B cells. Nature Immunol. 3, 443–450 (2002).
Reimold, A.M. et al. Plasma cell differentiation requires the transcription factor XBP-1. Nature 412, 300–307 (2001).
Calfon, M. et al. IRE1 couples endoplasmic reticulum load to secretory capacity by processing the XBP-1 mRNA. Nature 415, 92–96 (2002).
Agarwal, S., Avni, O. & Rao, A. Cell-type-restricted binding of the transcription factor NFAT to a distal IL-4 enhancer in vivo. Immunity 12, 643–652 (2000).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Yang, J., Zhu, H., Murphy, T.L., Ouyang, W. & Murphy, K.M., IL-18-stimulated GADD45β required in cytokine-induced, but not TCR-induced, IFN-γ production. Nature Immunol. 2, 157–164 (2001).
Lee, H.J. et al. GATA-3 induces T helper cell type 2 (Th2) cytokine expression and chromatin remodeling in committed Th1 cells. J. Exp. Med. 192, 105–115 (2000).
Zhang, D.H., Yang, L. & Ray, A. Differential responsiveness of the IL-5 and IL-4 genes to transcription factor GATA-3. J. Immunol. 161, 3817–382 (1998).
Cirillo, L.A. et al. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol. Cell 9, 279–289 (2002).
Lighvani, A.A. et al. T-bet is rapidly induced by interferon-γ in lymphoid and myeloid cells. Proc. Natl. Acad. Sci. USA 98, 15137–15142 (2001).
Kozak, M. Compilation and analysis of sequences upstream from the translational start site in eukaryotic mRNAs. Nucl. Acids Res. 12, 857–872 (1984).
Pear, W.S., Nolan, G.P., Scott, M.L. & Baltimore, D. Production of high-titer helper-free retroviruses by transient transfection. Proc. Natl. Acad. Sci. USA 90, 8392–8396 (1993).
Fitzpatrick, D.R. et al. Distinct methylation of the interferon γ (IFN-γ) and interleukin 3 (IL-3) genes in newly activated primary CD8+ T lymphocytes: regional IFN-γ promoter demethylation and mRNA expression are heritable in CD44highCD8+ T cells. J. Exp. Med. 188, 103–117 (1998).
Acknowledgements
We thank M. Weiss for helpful discussion; W. Pear and B. Cobb for advice on retrovirus production; V. Amorosa, J. Hartman, E. L. Pearce, M. Kessler and A. Villarino for assistance; D. Kessler for the cDNA of Drosophila engrailed repression domain; and W. DeMuth for cell sorting. Supported by the NIH (AI42370 to SLR).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Mullen, A., Hutchins, A., High, F. et al. Hlx is induced by and genetically interacts with T-bet to promote heritable TH1 gene induction. Nat Immunol 3, 652–658 (2002). https://doi.org/10.1038/ni807
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni807
This article is cited by
-
T Lymphocyte Characteristic Changes Under Serum Cytokine Deviations and Prognostic Factors of COVID-19 in Pregnant Women
Applied Biochemistry and Biotechnology (2023)
-
A comprehensive evaluation of the immune system response and type-I Interferon signaling pathway in hospitalized COVID-19 patients
Cell Communication and Signaling (2022)
-
T cell epigenetic remodeling and accelerated epigenetic aging are linked to long-term immune alterations in childhood cancer survivors
Clinical Epigenetics (2018)
-
IL-4 enhances IL-10 production in Th1 cells: implications for Th1 and Th2 regulation
Scientific Reports (2017)
-
DNA Methylation: a New Player in Multiple Sclerosis
Molecular Neurobiology (2017)


