Abstract
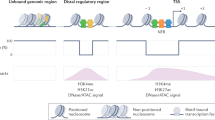
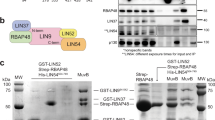
FoxP3 conditions the transcriptional signature and functional facets of regulatory T cells (Treg cells). Its mechanism of action, whether as an activator or a repressor, has remained unclear. Here, chromatin analysis showed that FoxP3 bound active enhancer elements, not repressed chromatin, around loci over- or under-expressed in Treg cells. We evaluated the impact of a panel of FoxP3 mutants on its transcriptional activity and interactions with DNA, transcriptional cofactors and chromatin. Computational integration, confirmed by biochemical interaction and size analyses, showed that FoxP3 existed in distinct multimolecular complexes. It was active and primarily an activator when complexed with the transcriptional factors RELA, IKZF2 and KAT5. In contrast, FoxP3 was inactive when complexed with the histone methyltransferase EZH2 and transcription factors YY1 and IKZF3. The latter complex partitioned to a peripheral region of the nucleus, as shown by super-resolution microscopy. Thus, FoxP3 acts in multimodal fashion to directly activate or repress transcription, in a context- and partner-dependent manner, to govern Treg cell phenotypes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Sakaguchi, S. Naturally arising CD4+ regulatory t cells for immunologic self-tolerance and negative control of immune responses. Annu. Rev. Immunol. 22, 531–562 (2004).
Josefowicz, S.Z., Lu, L.F. & Rudensky, A.Y. Regulatory T cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012).
Panduro, M., Benoist, C. & Mathis, D. Tissue Tregs. Annu. Rev. Immunol. 34, 609–633 (2016).
Ziegler, S.F. FOXP3: of mice and men. Annu. Rev. Immunol. 24, 209–226 (2006).
Fontenot, J.D. et al. Regulatory T cell lineage specification by the forkhead transcription factor foxp3. Immunity 22, 329–341 (2005).
Sugimoto, N. et al. Foxp3-dependent and -independent molecules specific for CD25+CD4+ natural regulatory T cells revealed by DNA microarray analysis. Int. Immunol. 18, 1197–1209 (2006).
Hill, J.A. et al. Foxp3 transcription-factor-dependent and -independent regulation of the regulatory T cell transcriptional signature. Immunity 27, 786–800 (2007).
Ferraro, A. et al. Interindividual variation in human T regulatory cells. Proc. Natl. Acad. Sci. USA 111, E1111–E1120 (2014).
Arvey, A. et al. Genetic and epigenetic variation in the lineage specification of regulatory T cells. eLife 4, e07571 (2015).
Wan, Y.Y. & Flavell, R.A. Regulatory T-cell functions are subverted and converted owing to attenuated Foxp3 expression. Nature 445, 766–770 (2007).
Gavin, M.A. et al. Foxp3-dependent programme of regulatory T-cell differentiation. Nature 445, 771–775 (2007).
Cipolletta, D. et al. PPAR-γ is a major driver of the accumulation and phenotype of adipose tissue Treg cells. Nature 486, 549–553 (2012).
Lopes, J.E. et al. Analysis of FOXP3 reveals multiple domains required for its function as a transcriptional repressor. J. Immunol. 177, 3133–3142 (2006).
Li, B. et al. FOXP3 is a homo-oligomer and a component of a supramolecular regulatory complex disabled in the human XLAAD/IPEX autoimmune disease. Int. Immunol. 19, 825–835 (2007).
Wu, Y. et al. FOXP3 controls regulatory T cell function through cooperation with NFAT. Cell 126, 375–387 (2006).
Bandukwala, H.S. et al. Structure of a domain-swapped FOXP3 dimer on DNA and its function in regulatory T cells. Immunity 34, 479–491 (2011).
Chen, Y. et al. DNA binding by FOXP3 domain-swapped dimer suggests mechanisms of long-range chromosomal interactions. Nucleic Acids Res. 43, 1268–1282 (2015).
Wright, P.E. & Dyson, H.J. Intrinsically disordered proteins in cellular signalling and regulation. Nat. Rev. Mol. Cell Biol. 16, 18–29 (2015).
Andersen, K.G., Nissen, J.K. & Betz, A.G. Comparative genomics reveals key gain-of-function events in Foxp3 during regulatory T cell evolution. Front. Immunol. 3, 113 (2012).
Xiao, Y. et al. Histone acetyltransferase mediated regulation of FOXP3 acetylation and Treg function. Curr. Opin. Immunol. 22, 583–591 (2010).
Rudra, D. et al. Transcription factor Foxp3 and its protein partners form a complex regulatory network. Nat. Immunol. 13, 1010–1019 (2012).
Bettini, M.L. et al. Loss of epigenetic modification driven by the Foxp3 transcription factor leads to regulatory T cell insufficiency. Immunity 36, 717–730 (2012).
Zheng, Y. et al. Regulatory T-cell suppressor program co-opts transcription factor IRF4 to control TH2 responses. Nature 458, 351–356 (2009).
Chaudhry, A. et al. CD4+ regulatory T cells control TH17 responses in a Stat3-dependent manner. Science 326, 986–991 (2009).
Darce, J. et al. An N-terminal mutation of the Foxp3 transcription factor alleviates arthritis but exacerbates diabetes. Immunity 36, 731–741 (2012).
Samstein, R.M. et al. Foxp3 exploits a pre-existent enhancer landscape for regulatory T cell lineage specification. Cell 151, 153–166 (2012).
Schubert, L.A., Jeffery, E., Zhang, Y., Ramsdell, F. & Ziegler, S.F. Scurfin (FOXP3) acts as a repressor of transcription and regulates T cell activation. J. Biol. Chem. 276, 37672–37679 (2001).
Bettelli, E., Dastrange, M. & Oukka, M. Foxp3 interacts with nuclear factor of activated T cells and NF-κB to repress cytokine gene expression and effector functions of T helper cells. Proc. Natl. Acad. Sci. USA 102, 5138–5143 (2005).
Li, B. et al. FOXP3 interactions with histone acetyltransferase and class II histone deacetylases are required for repression. Proc. Natl. Acad. Sci. USA 104, 4571–4576 (2007).
Chen, C., Rowell, E.A., Thomas, R.M., Hancock, W.W. & Wells, A.D. Transcriptional regulation by Foxp3 is associated with direct promoter occupancy and modulation of histone acetylation. J. Biol. Chem. 281, 36828–36834 (2006).
Arvey, A. et al. Inflammation-induced repression of chromatin bound by the transcription factor Foxp3 in regulatory T cells. Nat. Immunol. 15, 580–587 (2014).
Kitagawa, Y. et al. Guidance of regulatory T cell development by Satb1-dependent super-enhancer establishment. Nat. Immunol. 18, 173–183 (2017).
Fontenot, J.D., Gavin, M.A. & Rudensky, A.Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061 (2003).
Luo, C.T. & Li, M.O. Transcriptional control of regulatory T cell development and function. Trends Immunol. 34, 531–539 (2013).
Benoist, C. & Mathis, D. Treg cells, life history, and diversity. in Immune Tolerance (eds. Mathis, D. & Rudensky, A.) 31–44 (Cold Spring Harbor Press, Cold Spring Harbor, New York, 2013).
Quintana, F.J. et al. Aiolos promotes TH17 differentiation by directly silencing Il2 expression. Nat. Immunol. 13, 770–777 (2012).
Xiong, Y. et al. Polycomb antagonizes p300/CREB-binding protein-associated factor to silence FOXP3 in a Kruppel-like factor-dependent manner. J. Biol. Chem. 287, 34372–34385 (2012).
Gordon, S., Akopyan, G., Garban, H. & Bonavida, B. Transcription factor YY1: structure, function, and therapeutic implications in cancer biology. Oncogene 25, 1125–1142 (2006).
Hwang, S.S. et al. YY1 inhibits differentiation and function of regulatory T cells by blocking Foxp3 expression and activity. Nat. Commun. 7, 10789 (2016).
Cowell, I.G. Repression versus activation in the control of gene transcription. Trends Biochem. Sci. 19, 38–42 (1994).
Thiel, G., Lietz, M. & Hohl, M. How mammalian transcriptional repressors work. Eur. J. Biochem. 271, 2855–2862 (2004).
Hahn, S. Ellis Englesberg and the discovery of positive control in gene regulation. Genetics 198, 455–460 (2014).
Fischer, M., Steiner, L. & Engeland, K. The transcription factor p53: not a repressor, solely an activator. Cell Cycle 13, 3037–3058 (2014).
Mekhail, K. & Moazed, D. The nuclear envelope in genome organization, expression and stability. Nat. Rev. Mol. Cell Biol. 11, 317–328 (2010).
Loizou, L., Andersen, K.G. & Betz, A.G. Foxp3 interacts with c-Rel to mediate NF-κB repression. PLoS One 6, e18670 (2011).
Lee, S.M., Gao, B. & Fang, D. FoxP3 maintains Treg unresponsiveness by selectively inhibiting the promoter DNA-binding activity of AP-1. Blood 111, 3599–3606 (2008).
Gill, G. & Ptashne, M. Negative effect of the transcriptional activator GAL4. Nature 334, 721–724 (1988).
Fu, W. et al. A multiply redundant genetic switch 'locks in' the transcriptional signature of regulatory T cells. Nat. Immunol. 13, 972–980 (2012).
Blecher-Gonen, R. et al. High-throughput chromatin immunoprecipitation for genome-wide mapping of in vivo protein-DNA interactions and epigenomic states. Nat. Protoc. 8, 539–554 (2013).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B.A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Whyte, W.A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Shen, L., Shao, N., Liu, X. & Nestler, E. ngs.plot: Quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 15, 284 (2014).
Koh, K.P., Sundrud, M.S. & Rao, A. Domain requirements and sequence specificity of DNA binding for the forkhead transcription factor FOXP3. PLoS One 4, e8109 (2009).
Wakamatsu, E., Mathis, D. & Benoist, C. Convergent and divergent effects of costimulatory molecules in conventional and regulatory CD4+ T cells. Proc. Natl. Acad. Sci. USA 110, 1023–1028 (2013).
Hiraoka, Y., Agard, D.A. & Sedat, J.W. Temporal and spatial coordination of chromosome movement, spindle formation, and nuclear envelope breakdown during prometaphase in Drosophila melanogaster embryos. J. Cell Biol. 111, 2815–2828 (1990).
Gustafsson, M.G. et al. Three-dimensional resolution doubling in wide-field fluorescence microscopy by structured illumination. Biophys. J. 94, 4957–4970 (2008).
Acknowledgements
We thank A. Arvey, L. Chen, T. Chatila, K. Struhl, J. Waters and A. Rudensky for discussions: K. Hattori, C. Araneo and A. Rhoads for help with mice, cell sorting, profiling and software; and the HMS Cell Biology Microscopy Facility and L. Shao for super-resolution microscopy. Supported by the US National Institutes of Health (AI116834), GlaxoSmithKline (Sponsored Research Agreement+ and NRF (357-2011-1C00084 to H.-K.K.).
Author information
Authors and Affiliations
Contributions
H.-K.K. and H.-M.C. performed the experiments; all authors designed the study and analyzed and interpreted the data, and H.-K.K., C.B. and D.M. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Determinants of FoxP3’s transcriptional functions.
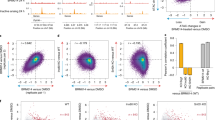
(a) Representative flow cytometry plots of FoxP3 expression after transduction of EV, WT and mutant FoxP3 vectors into activated CD4+ T cells. Surface staining detected the vector-encoded Thy1.1 reporter (which serves as a standard and to guide cell sorting in later experiments) and intracellular staining detected FoxP3 levels. Data are representative of biological triplicates (quantitative results tabulated in Table S4). (b) After retroviral transduction as in a, the size and expression of the FoxP3 proteins were detected by immunoblotting of nuclear extracts with anti-FLAG antibody. Data representative of biological triplicates. (c) WT or mutant FoxP3 were transduced into HEK293 cells, and after 72 hr of transfection, cells were stained with anti-FLAG (red) and counterstained for nuclear DNA with DAPI (blue). Data representative of biological triplicates. (d, e): Evaluation of DNA-binding capacity of WT or mutant FoxP3. Specificity controls: nuclear extracts prepared from CD4+ T cells 72 hrs after transduced with EV or WT FoxP3 vectors as in a were incubated with FKRE2 or control (scrambled sequence) oligos, and binding detected by ELISA with anti-FLAG (results shown as mean of biological duplicates, ± SD) (e) Binding activity of WT and mutant FoxP3 in the same assay (values standardized to binding by WT FoxP3 in each experiment; mean +/- SD, Data representative for 2-3 biological experiments). (f) Correlation between the suppressive activity of CD4+ T cells transduced with the mutant FoxP3 panel and the induction (or repression) of individual FoxP3 target genes by the same mutant panel (as Fig. 2f, except for individual transcripts). t.test, P* < 0.05, P** < 0.005 and P*** < 0.001
Supplementary Figure 2 Determinants of FoxP3’s transcriptional functions.
(a) Specificity controls for the co-IP experiments (Fig. 4). To validate the interaction between WT FoxP3 and 17 selected co-factors, FLAG-tagged FoxP3 and each co-factor (tagged with HA, 6xHIS or GFP) were co-transduced into HEK293 cells and immuno-precipitated with anti-FLAG or isotype-matched IgG. Specific co-IP of each co-factor was detected by specific anti-tag antibody. Data representative of biological triplicates. (b, c): Co-immunoprecipitation of FoxP3 and cofactors does not depend on DNA as an intermediate. Co-IPs were performed as in Fig. 4 in the presence of 10μg/ml Ethidium bromide (b) or 1 mg/ml DNAse I (c). Representative immunoblots from biological duplicates. (d) The co-IP experiments primarily detect pre-formed multimolecular complexes, not in vitro-association. To test for artefactual complex formation, lysates of cells singly or doubly transduced with WT FoxP3 and/or STAT3 or RELA were incubated alone or after mixing for 30 minutes under co-IP conditions, then co-IPed with anti-FLAG (FoxP3) and STAT3 or RELA detected by immunoblotting (as for Fig. 4 and Supplementary Fig. 2B). Interaction is only detected robustly with lysates from doubly-transduced cells, although a low amount of interaction can be detected by co-incubating lysates from singly-transduced cells. Results representative of biological duplicates.
Supplementary Figure 3 FoxP3 forms two distinct complexes.
3D-SIM detection of FoxP3, RELA and IKZF3 in ex vivo Treg cells. Image representative of >20 cells in biological triplicates as for Fig. 7. Scale bars (left and middle: 1 μm, right: 0.1 μm).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 (PDF 515 kb)
Supplementary Table 1
Probe sequences for each transcript in Treg specific code-set (XLSX 65 kb)
Supplementary Table 2
Mutant information (XLSX 11 kb)
Supplementary Table 3
FoxP3 expression (MFI) for WT and mutant constructs (XLSX 9 kb)
Supplementary Table 4
Nanostring data (XLSX 57 kb)
Supplementary Table 5
References for co-factor and FoxP3 interaction (XLSX 12 kb)
Supplementary Table 6
Co-factor interaction (XLSX 23 kb)
Supplementary Video 1
3D-SIM for FoxP3, RelA and EZH2 in FoxP3 transduced CD4+ T cells (MP4 15365 kb)
Rights and permissions
About this article
Cite this article
Kwon, HK., Chen, HM., Mathis, D. et al. Different molecular complexes that mediate transcriptional induction and repression by FoxP3. Nat Immunol 18, 1238–1248 (2017). https://doi.org/10.1038/ni.3835
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3835
This article is cited by
-
The regulation and differentiation of regulatory T cells and their dysfunction in autoimmune diseases
Nature Reviews Immunology (2024)
-
Targeting the epigenome to reinvigorate T cells for cancer immunotherapy
Military Medical Research (2023)
-
Three-dimensional structured illumination microscopy with enhanced axial resolution
Nature Biotechnology (2023)
-
Regulatory T cells in the face of the intestinal microbiota
Nature Reviews Immunology (2023)
-
Foxp3 orchestrates reorganization of chromatin architecture to establish regulatory T cell identity
Nature Communications (2023)