Abstract
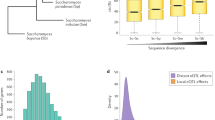
Evolution may depend more strongly on variation in gene expression than on differences between variant forms of proteins1. Regions of DNA that affect gene expression are highly variable, containing 0.6% polymorphic sites2. These naturally occurring polymorphic nucleotides can alter in vivo transcription rates3,4,5,6,7. Thus, one might expect substantial variation in gene expression between individuals. But the natural variation in mRNA expression for a large number of genes has not been measured. Here we report microarray studies addressing the variation in gene expression within and between natural populations of teleost fish of the genus Fundulus. We observed statistically significant differences in expression between individuals within the same population for approximately 18% of 907 genes. Expression typically differed by a factor of 1.5, and often more than 2.0. Differences between populations increased the variation. Much of the variation between populations was a positive function of the variation within populations and thus is most parsimoniously described as random. Some genes showed unexpected patterns of expression—changes unrelated to evolutionary distance. These data suggest that substantial natural variation exists in gene expression and that this quantitative variation is important in evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
King, M.C. & Wilson, A.C. Evolution at two levels: molecular similarities and biological differences between humans and chimpanzees. Science 188, 107–116 (1975).
Stephens, J.C. et al. Haplotype variation and linkage disequilibrium in 313 human genes. Science 293, 489–493 (2001).
Segal, J.A., Barnett, J.L. & Crawford, D.L. Functional analyses of natural variation in Sp1 binding sites of a TATA-less promoter. J. Mol. Evol. 49, 736–749 (1999).
Crawford, D.L., Segal, J.A. & Barnett, J.L. Evolutionary analysis of TATA-less proximal promoter function. Mol. Biol. Evol. 16, 194–207 (1999).
Beaty, J.S., West, K.A. & Nepom, G.T. Functional effects of a natural polymorphism in the transcriptional regulatory sequence of HLA-DQB1. Mol. Cell. Biol. 15, 4771–4782 (1995).
Antonarakis, S.E. et al. β-Thalassemia in American Blacks: novel mutations in the “TATA” box and an acceptor splice site. Proc. Natl Acad. Sci. USA 81, 1154–1158 (1984).
Koivisto, U.M., Palvimo, J.J., Janne, O.A. & Kontula, K. A single-base substitution in the proximal Sp1 site of the human low density lipoprotein receptor promoter as a cause of heterozygous familial hypercholesterolemia. Proc. Natl Acad. Sci. USA 91, 10526–10530 (1994).
Lewohl, J.M., Dodd, P.R., Mayfield, R.D. & Harris, R.A. Application of DNA microarrays to study human alcoholism [Review]. J. Biomed. Sci. 8, 28–36 (2001).
Golub, T.R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999).
Alizadeh, A.A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
DeRisi, J. et al. Use of a cDNA microarray to analyze gene expression patterns in human cancer. Nature Genet. 14, 457–460 (1996).
Lau, W.Y. et al. Differential gene expression of hepatocellular carcinoma using cDNA microarray analysis. Oncol. Res. 12, 59–69 (2000).
Lotrich, V.A. Summer home range and movements of Fundulus heteroclitus (Pisces: Cyprinodotidae) in tidal creek. Ecology 56, 191–198 (1975).
Brown, B.L. & Chapman, R.W. Gene flow and mitochondrial DNA variation in the killifish Fundulus heteroclitus. Evolution 45, 1147–1161 (1991).
Podrabsky, J.E., Javillonar, C., Hand, S.C. & Crawford, D.L. Intraspecific variation in aerobic metabolism and glycolytic enzyme expression in heart ventricles from Fundulus heteroclitus. Am. J. Physiol. 279, R2344–R2348 (2000).
Pierce, V.A. & Crawford, D.L. Phylogenetic analysis of glycolytic enzyme expression. Science 275, 256–259 (1997).
Kerr, M., Martin, M. & Churchill, G. Analysis of variance for gene expression microarray data. J. Comput. Biol. 7, 819–37 (2000).
Kerr, M.K. & Churchill, G.A. Statistical design and the analysis of gene expression microarray data. Genet. Res. 77, 123–128 (2001).
Jin, W. et al. The contributions of sex, genotype and age to transcriptional variance in Drosophila melanogaster. Nature Genet. 29, 389–395 (2001).
Sokal, R.R. & Rohlf, F.J. Biometry (W.H. Freeman, New York, 1981).
Brem, R.B., Yvert, G., Clinton, R. & Kruglyak, L. Genetic dissection of transcriptional regulation in budding yeast. Science 296, 752–755 (2002).
Enard, W. et al. Intra- and interspecific variation in primate gene expression patterns. Science 296, 340–343 (2002).
Crawford, D.L. & Powers, D.A. Molecular basis of evolutionary adaptation at the lactate dehydrogenase-B locus in the fish Fundulus heteroclitus. Proc. Natl Acad. Sci. USA 86, 9365–9369 (1989).
Nei, M. Molecular Evolutionary Genetics (Columbia Univ. Press, New York, 1987).
Bernardi, G. & Powers, D.A. Phylogenetic relationships among nine species from the genus Fundulus (Cyprinodontiformes, Fundulidae) inferred from sequences of the cytochrome B. Copeia 1995, 469–473 (1995).
Cashner, R.C., Rogers, J.S. & Grady, J.M. Phylogenetic studies of the genus Fundulus. in Systematic, Historical Ecology and North American Freshwater Fishes (ed. Mayden, R.L.) 421–437 (Stanford Univ. Press, Stanford, California, 1992).
Wiley, E.O. A study of the evolutionary relationships of Fundulus topminnows (Teleostei: Fundulidae). Am. Zool. 26, 121–130 (1986).
Westfall, P.H. & Young, S.S. Resampling-Based Multiple Testing: Examples and Methods for P-Value Adjustment (Wiley, New York, 1993).
Oleksiak, M.F., Kolell, K. & Crawford, D.L. The utility of natural populations for microarray analyses: isolation of genes necessary for functional genomic studies. Mar. Biotech. 3, S203–S211 (2001).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863–14868 (1998).
Acknowledgements
This research was supported by a National Science Foundation BioInformatics post-doctoral fellowship to M.F.O., a National Cancer Institute grant to G.A.C. and a National Science Foundation Integrative Biology and Neuroscience grant to D.L.C. Additional support was provided by the School of Biological Science through M. Marino-Carrion. We would like to thank K. Horgan of M.J. Research for arranging the donation of Tetrad-thermal cycler, Motorola for donating the activated slides used in the production of microarrays and AP-Biotech and specifically R. Feldman for use of MegaBace used to re-sequence cDNAs.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Oleksiak, M., Churchill, G. & Crawford, D. Variation in gene expression within and among natural populations. Nat Genet 32, 261–266 (2002). https://doi.org/10.1038/ng983
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng983
This article is cited by
-
The role and risks of selective adaptation in extreme coral habitats
Nature Communications (2023)
-
Cardiac physiology and metabolic gene expression during late organogenesis among F. heteroclitus embryo families from crosses between pollution-sensitive and -resistant parents
BMC Ecology and Evolution (2022)
-
A mathematical model of circadian rhythms and dopamine
Theoretical Biology and Medical Modelling (2021)
-
Establishment of a striped catfish skin explant model for studying the skin response in Aeromonas hydrophila infections
Scientific Reports (2021)
-
Generational sensitivity alteration in Chironomus yoshimatsui to carbamate and pharmaceutical chemicals and the effect on Catalase, CYP450, and hemoglobin gene transcription
Ecotoxicology (2021)