Abstract
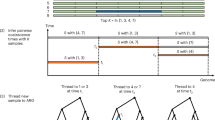
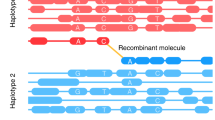
The study of complex genetic traits in humans is limited by the expense and difficulty of ascertaining populations of sufficient sample size to detect subtle genetic contributions to disease. Here we introduce an application of a somatic cell hybrid construction strategy called conversion1,2,3,4 that maximizes the genotypic information from each sampled individual. The approach permits direct observation of individual haplotypes, thereby eliminating the need for collecting and genotyping DNA from family members for haplotype-based analyses. We describe experimental data that validate the use of conversion as a whole-genome haplotyping tool and evaluate the theoretical efficiency of using conversion-derived haplotypes instead of conventional genotypes in the context of haplotype-frequency estimation. We show that, particularly when phenotyping is expensive, conversion-based haplotyping can be more efficient and cost-effective than standard genotyping.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Papadopolous, N., Leach, F.S., Kinzler, K.W. & Vogelstein, B. Monoallelic mutation analysis (MAMA) for identifying germline mutations. Nature Genet. 11, 99–102 (1995).
Laken, S.J. et al. Analysis of masked mutations in familial adenomatous polyposis. Proc. Natl. Acad. Sci. USA 96, 2322–2326 (1999).
Yan, H. et al. Conversion of diploidy to haploidy. Nature 403, 723–724 (2000).
Yan, H., Kinzler, K.W. & Vogelstein, B. Genetic testing—present and future. Science 289, 1890–1892 (2000).
MacLean, C.J. & Morton, N.E. Estimation of myriad haplotype frequencies. Genet. Epidemiol. 2, 263–272 (1985).
Excoffier, L. & Slatkin, M. Maximum-likelihood estimation of molecular haplotype frequencies in a diploid population. Mol. Biol. Evol. 12, 921–927 (1995).
Hawley, M.E. & Kidd, K.K. HAPLO: a program using the EM algorithm to estimate the frequencies of multi-site haplotypes. J. Hered. 86, 409–411 (1995).
Long, J.C., Williams, R.C. & Urbanek, M. An E-M algorithm and testing strategy for multiple-locus haplotypes. Am. J. Hum. Genet. 56, 799–810 (1995).
Michalatos-Beloin, S., Tishkoff, S.A., Bentley, K.L., Kidd, K.K. & Ruano, G. Molecular haplotyping of genetic markers 10 kb apart by allele-specific long-range PCR. Nucleic Acids Res. 24, 4841–4843 (1996).
Hodge, S.E., Boehnke, M. & Spence, M.A. Loss of information due to ambiguous haplotyping of SNPs. Nature Genet. 21, 360–361 (1999).
Weiss, M.C. & Green, H. Human-mouse hybrid cell lines containing partial complements of human chromosomes and functioning human genes. Proc. Natl. Acad. Sci. USA 58, 1104–1111 (1967).
Nishimura, D.Y. et al. The forkhead transcription factor gene FKHL7 is responsible for glaucoma phenotypes which map to 6p25. Nature Genet. 19, 140–147 (1998).
Glaser, T. et al. The beta-subunit of follicle-stimulating hormone is deleted in patients with aniridia and Wilms' tumour, allowing a further definition of the WAGR locus. Nature 321, 882–887 (1986).
Tishkoff, S.A., Pakstis, A.J., Ruano, G. & Kidd, K.K. The accuracy of statistical methods for estimation of haplotype frequencies: an example from the CD4 locus. Am. J. Hum. Genet. 67, 518–522 (2000).
Hill, W.G. Estimation of linkage disequilibrium in randomly mating populations. Heredity 33, 229–239 (1974).
McKeigue, P.M. Efficiency of estimation of haplotype frequencies: use of marker phenotypes of unrelated individuals versus gene counting of phase known gametes. Am. J. Hum. Genet. 67, 1626–1627 (2000).
Veldman, T., Vignon, C., Schrock, E., Rowley, J.D. & Ried, T. Hidden chromosome abnormalities in haematological malignancies detected by multicolour spectral karyotyping. Nature Genet. 15, 406–410 (1997).
Acknowledgements
We thank B. Vogelstein and H. Yan of Johns Hopkins University for generously donating the somatic cell hybrids. We would also like to acknowledge the technical assistance of T. Dennis and A. Dutra of the National Human Genome Research Institute Cytogenetic and Confocal Microscopy Core. This work was supported in part by a University of Michigan Predoctoral Fellowship to J.A.D. and by National Institutes of Health grants to M.B. (R01 HG00376) and S.B.G. (R01 CA81488).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Douglas, J., Boehnke, M., Gillanders, E. et al. Experimentally-derived haplotypes substantially increase the efficiency of linkage disequilibrium studies. Nat Genet 28, 361–364 (2001). https://doi.org/10.1038/ng582
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng582
This article is cited by
-
NCMHap: a novel method for haplotype reconstruction based on Neutrosophic c-means clustering
BMC Bioinformatics (2020)
-
DNA origami-based shape IDs for single-molecule nanomechanical genotyping
Nature Communications (2017)
-
The CRHR1 Gene Contributes to Genetic Susceptibility of Aggressive Behavior Towards Others in Chinese Southwest Han Population
Journal of Molecular Neuroscience (2014)
-
Whole-genome molecular haplotyping of single cells
Nature Biotechnology (2011)
-
Gold nanoparticles for high-throughput genotyping of long-range haplotypes
Nature Nanotechnology (2011)