Abstract
Sex chromosomes are subject to sex-specific selective evolutionary forces1,2. One model predicts that genes with sex-biased expression should be enriched on the X chromosome2,3,4,5. In agreement with Rice's hypothesis3, spermatogonial genes are over-represented on the X chromosome of mice6 and sex- and reproduction-related genes are over-represented on the human X chromosome7,8. Male-biased genes are under-represented on the X chromosome in worms and flies9,10,11, however. Here we show that mouse spermatogenesis genes are relatively under-represented on the X chromosome and female-biased genes are enriched on it. We used Spo11−/− mice blocked in spermatogenesis early in meiosis12 to evaluate the temporal pattern of gene expression in sperm development. Genes expressed before the Spo11 block are enriched on the X chromosome, whereas those expressed later in spermatogenesis are depleted. Inactivation of the X chromosome in male meiosis may be a universal driving force for X-chromosome demasculinization.
Similar content being viewed by others
Main
To investigate the genomic distribution of mouse genes with patterns of sex-biased expression, we analyzed the chromosomal distribution of genes expressed in sexually dimorphic tissues. We consider genes that are preferentially expressed in such tissues as testis, ovary and placenta to be sex-biased genes. We used two approaches to define the tissue specificity of genes expressed in mice: analysis of large-scale microarray-based gene expression data and analysis of the distribution of expressed-sequence tags (ESTs) in different cDNA libraries. Instead of focusing on transcripts restricted to a single tissue, our analysis targeted the identification of tissue-enriched transcripts using a 'preferential expression measure' (PEM)13. As an uniform source of expression data, we used publicly available data sets of mouse RNA samples representing 49 tissues hybridized to Affymetrix mouse U74A microarrays (GNF data set)14 and 20 tissues analyzed with RIKEN cDNA microarrays (RIKEN data set)15.
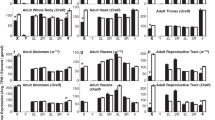
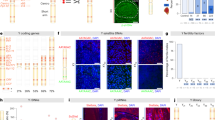
In the GNF data set14, expression of 624 genes (∼7% of all analyzed) in testis was three times greater than the median expression in other tissues, and expression of 361 genes in testis was five times greater (PEM values >1.58 and >2.32, respectively). These genes are considered testis-enriched genes. The chromosomal distribution of testis-enriched genes was significantly different from uniform (χ2 = 30.8, P = 0.05; Fig. 1a and Supplementary Table 1 online). The most significant deviation from the average gene density was on the X chromosome, where testis-enriched genes were 2.5 times less dense (the ratio of the density of testis-enriched genes on the X chromosome to the average density on all chromosomes, ρX, was 0.40; χ2 = 7.7, P = 0.006). The depletion of testis-enriched genes on the X chromosome was independent of the cut-off (three or five times greater expression) used in the selection process (Table 1). In addition, genes that were expressed most highly in testis and all the genes detected in testis were under-represented on the X chromosome (Table 1).
(a–e) The percentages of genes residing on individual chromosomes and satisfying given selection criteria at different stringency levels are shown. Dark blue, genes that are five times more abundant than median of all tissues; light blue, genes that are three times more abundant. (f,g) The chromosomal distribution of tissue-enriched genes based on an analysis of the RIKEN data set. Dark green, genes whose expression is five times greater than the median; light green, genes whose expression is three times greater. (h) The chromosomal distribution of transcripts enriched in testis as found in our EST analysis using R statistics. Dark yellow, R > 10, expression in testis is two times greater than median; light yellow, R > 20, expression in testis is two times greater. Chromosomes that have gene densities significantly deviating from the average are indicated (*P < 0.05, **P < 0.01, ***P < 0.001).
In contrast to testes-enriched genes, the chromosomal distribution of most tissue-enriched genes was not significantly different from random (Fig. 1d,e; χ2 = 23.1 and 13.4 for liver- and lung-enriched genes, respectively; P > 0.1). The chromosomal distribution of ovary-specific genes, however, was far from uniform (χ2 = 31.9, P = 0.03; Fig. 1b). Again, the chromosome with the most profound difference in gene density was the X chromosome, but unlike testis-enriched genes, ovary-enriched genes were more abundant on the X (ρX = 1.95; χ2 = 10.58, P = 0.001; Table 1). We also investigated the chromosome distribution for genes preferentially expressed in placenta (Fig. 1c), which was markedly similar to that of ovary-enriched genes (ρX = 1.66; χ2 = 7.24, P = 0.007). As with testis-enriched genes, the enrichment was independent of the cut-off expression level for both sets of female-biased genes (Table 1). In addition to tissue-enriched genes, genes that are highly expressed in placenta or ovary were also enriched on the X chromosome (ρX = 1.5, P = 0.03 and ρX = 1.7, P = 0.01, respectively; Table 1). An analysis of the independent RIKEN set of microarray expression data15 confirmed the trends observed with the GNF set: testis-enriched genes were under-represented on the X chromosome (ρX = 0.62; χ2 = 7.92, P = 0.005) and placenta-enriched genes were over-represented (ρX = 2.12; χ2 = 12.1, P < 0.001; Fig. 1f,g).
To complement and extend the microarray expression data, we analyzed the distribution of ESTs in different cDNA libraries available in GenBank. We selected genes that satisfied two criteria: (i) a statistically significant preferential expression in testis and (ii) expression at least two times higher in testis than in other tissues. The chromosomal distribution of testis-enriched genes based on the EST data confirmed the under-representation of these genes on the X chromosome (ρX = 0.45; χ2 = 5.0, P = 0.02; Fig. 1h).
The X chromosome is not the only chromosome with a skewed distribution of tissue-enriched genes. Analysis of both the GNF data set and EST data showed that testis-enriched genes are over-represented on chromosome 17. Approximately half (18 of 38) of the testis-enriched genes on chromosome 17 are located in the t-complex region that contains many male fertility genes16. In addition, we identified another cluster located centromeric to the t-complex (Supplementary Table 2 online).
To further define testis- and ovary-enriched genes, we compared the RNAs expressed in mouse gonads using direct competitive hybridization on cDNA microarrays. We identified 749 genes whose expression was at least three times higher in testes than in ovaries and 404 genes that were preferentially expressed in ovaries compared with testes (Fig. 2a). Again we found that the X chromosome is a disfavored location for testis-enriched genes (ρX = 0.28; P < 0.001) but a preferred site for ovary-enriched genes (ρX = 2.88; P < 0.001; Fig. 2d). We also compared the RNAs expressed in testes and ovaries with a reference RNA population, which gives a measure of the preferential expression in a tissue that is similar to PEM. We found that the expression of 858 genes was at least three times higher in testes than in reference RNA and that 243 genes were overexpressed in ovaries (Fig. 2b,c). As in the direct comparison, testis-expressed genes were under-represented on the X chromosome (ρX = 0.50; P = 0.01; Fig. 2d). The distribution of genes enriched in ovaries, however, was closer to random (ρX = 1.10).
Scatter plots of gene expression values (in arbitrary units) in testis versus ovary (a), ovary versus reference RNA (b) and testis versus reference RNA (c). Each gene is represented by a dot and the axes are the associated intensity values. Genes whose expression was at least three times greater in a given tissue are shown in pink (ovary-enriched) and blue (testis-enriched). (d) The percentages of expressed genes residing on the X chromosome and the autosomes.
Our analysis of several independent data sets consistently identified deviations in the chromosomal distribution of sex-biased genes. The PEM values calculated for sexually dimorphic tissues from the different data sets were positively correlated on a gene-by-gene basis (Supplementary Fig. 1 online).
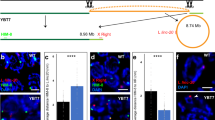
Testes contain more than 10 different cell types, most of which participate in spermatogenesis; over 90% are germ cells, most of which are past early meiosis (Fig. 3a)17,18. Sex chromosomes are condensed and transcriptionally repressed at the onset of male meiosis (meiotic sex chromosome inactivation, MSCI)19,20,21. To elucidate the effect of MSCI on the genomic distribution of male-biased genes, we analyzed the transcripts of testes from Spo11−/− mice which have a genetic ablation of cells past the early stages of spermatogenesis. The disruption of Spo11 in mice blocks meiosis before pachytene and interrupts normal testis development (Fig. 3a,b)12,22. In agreement with the marked changes in testis morphology in Spo11−/− mice (Fig. 3b), the expression of almost 30% of the transcripts changed by a factor of more than two (Fig. 3c). Consistent with the relative proportion of meiotic and postmeiotic cells in testis, most of these testis-enriched transcripts were depleted in testes of mature Spo11−/− mice (Supplementary Fig. 2 online). Like genes encoding testis-enriched transcripts, genes encoding transcripts depleted in the testes of Spo11−/− mice were strongly under-represented on the X chromosome (ρX = 0.34; χ2 = 12.9, P < 0.001; Fig. 3d). Next, we analyzed transcripts that were more abundant in testes of Spo11−/− mice than in testes of wild-type mice. Because testes of Spo11−/− mice are relatively enriched with premeiotic cells, the transcripts over-represented in these testes should include spermatogonial transcripts, reported to be over-represented on the X chromosome6. Like spermatogonial genes, Spo11−/−-enriched genes (which are not limited to spermatogonial genes) were significantly more dense on the X chromosome (ρX = 2.23; χ2 = 47.8, P < 0.001; Fig. 3d).
(a) Time line of spermatogenesis. (b) Hematoxylin and eosin–stained cross-section of seminiferous tubules from wild-type (WT) and Spo11−/− mice. Sperm development proceeds towards the lumen with the earliest cells with most dense nuclei at the periphery and later cells including elongated spermatids near the center. The seminiferous tubules of Spo11−/− mice are devoid of all but the earliest cells. (c) Gene expression profiles in testes of Spo11−/− and juvenile wild-type (15 d) mice. Testis RNA samples from adult (8–10 weeks) wild-type, adult Spo11−/− and 15-d-old wild-type mice were competitively hybridized to cDNA microarrays in four independent experiments in the combinations indicated. Intensities beyond a two-fold cutoff are shown in red (enriched in Spo11−/− or juvenile testis) and green (depleted in Spo11−/− or juvenile testis). Most genes reported by Wang and coworkers6 map to the group of ∼1,000 genes enriched in testes of Spo11−/− mice (13 of 20, compared with only 2 of 20 that mapped to the group of genes depleted in testes of Spo11−/− mice). (d) The chromosomal distributions of differentially expressed genes.
The first wave of meiosis in male mice is synchronous and starts at birth17 (Fig. 3a). Therefore, another approach to identify genes transcribed mostly in premeiotic cells is to work with testes from juvenile mice. At 15 d of age in wild-type mice, meiotic progression reaches approximately the stage of the block in Spo11−/− mice. We found that the expression profiles of 95% of the genes that were differentially expressed in testes of 15-d-old mice were similar to those of the Spo11 mutant (Supplementary Fig. 2 online). The chromosomal distribution of genes encoding transcripts that were enriched or depleted in juvenile mice followed the same trend as in the Spo11−/− mice (Fig. 3d; ρX = 0.44, χ2 = 9.96, P = 0.002 and ρX = 3.2, χ2 = 73.4, P < 0.001, respectively). Thus, both genetic ablation studies and analysis of testes from juvenile mice indicate that early genes are relatively abundant but that late genes are sparse on the mouse X chromosome.
A similar dissection of gene expression in spermatogenesis is possible using gene expression profiles of developing testes in mice23. We compared expression in samples from 1-, 4-, 8- and 11-d-old mice with samples from 18–26-d-old mice and found that the chromosome distribution of these genes was more skewed than that of Spo11−/− mice. Genes expressed at the earlier stages were abundant on the X chromosome (ρX = 2.32; χ2 = 125.9, P < 0.001) whereas late genes were nearly absent (ρX = 0.08; χ2 = 31.9, P < 0.001; Supplementary Table 1 online). We examined the differential genomic distribution of spermatogenic genes using existing data on the expression of these genes in the rat24. This analysis showed that meiotic-specific genes were completely absent from the X chromosome, whereas genes expressed in spermatogonia were significantly more abundant on the X chromosome (Fig. 4). Genes expressed in somatic cells of testis were also more abundant on the X chromosome.
Premeiotic, meiotic and postmeiotic germ cells and somatic Sertoli cells were purified from rat testes and the level of gene expression in individual cell fractions was measured using Affymetrix microarrays24. The percentage of genes on autosomes and on the X chromosome belonging to each cluster corresponding to a given cell population is plotted. Below the chart the relative gene density and the probability for observing such a distribution by chance are shown.
Next, we estimated the global effect of MSCI on transcription. Both the GNF and RIKEN data sets showed that genes located on the X chromosome had lower average expression in adult testes (Fig. 5a and Supplementary Fig. 3 online). Exploration of the mouse spermatogenesis data set23 showed that the decrease in net transcription from the X coincides with the onset of meiosis in developing testes (Fig. 5b). Although the X chromosome is inactivated in cells going through meiosis, several X-linked genes were reliably detected in more than one data set as testis-enriched (see Supplementary Note and Supplementary Table 3 online).
(a) Average expression level of genes residing on different chromosomes in testis, placenta and brain. The analysis was done on the RIKEN data set. (b) The transcriptional activity of chromosome 2 and the X and Y chromosomes during mouse spermatogenesis23. The average expression values were normalized to day 1 and plotted. An analysis of gene expression from other autosomes gave similar results to those obtained with chromosome 2.
In this study, we found that sex-biased genes had a nonrandom chromosomal distribution in the mouse. The accumulation of both male- and female-biased genes on the X chromosome is consistent with the predictions of W. Rice3 and previous reports on mice and humans6,7,8. We believe that the paucity of testis-enriched genes on the mouse X is mainly attributable to the effects of MSCI19,20. The under-representation of male-biased genes on the mouse X chromosome is similar to that reported for male-specific genes in Drosophila melanogaster and Caenorhabditis elegans9,10, where (in C. elegans, at least) MSCI is present25,26. The evolutionary roots of MSCI are deep, and MSCI is tightly linked to the evolution of the sex chromosomes. MSCI probably preceded somatic X-chromosome inactivation and may be the origin of paternal X inactivation in marsupials and in preimplantation mouse embryos27. Thus, there seem to be two main counteracting forces in evolution that affect the distribution of male-biased genes. One force removes male-biased genes from the X chromosome (MSCI), and the other adds them (the predicted and observed accumulation of male beneficial genes on the X chromosome). We believe that this hypothesis most parsimoniously accounts for and reconciles the previously published data on this subject from flies, worms, mice and humans.
Methods
Determining tissue-specificity from microarray data.
We defined genes as tissue-enriched or preferentially expressed if their expression in that tissue was several times greater than the median expression level calculated for all other tissues (the PEM is essentially the log2 relative excess over the median13). We obtained individual gene expression level data from data sets on hybridization intensity values of RNA samples from 49 different tissues on Affymetrix mouse U74A microarrays14 (GNF data set) and 20 tissues analyzed with RIKEN cDNA microarrays15 (RIKEN data set). The degree of correlation of PEM values from independent data sets is similar to that reported previously13. In the GNF data set, we defined genes expressed in a given tissue as those genes with an expression level higher than 200, estimated to correspond to three to five transcripts per cell14. We defined highly expressed genes in the GNF data set as those genes with an expression level higher than 1,200 in a tissue without regard to the expression in other tissues. We set this threshold so as to give ∼500 genes in every tissue.
EST analysis.
We counted the number of individual ESTs mapped to each Unigene cluster in two pools of cDNA libraries, one consisting of 7 testis cDNA libraries (pool A; 74,672 ESTs) and another a pool of 24 libraries from other tissues (pool B; 264,355 ESTs). The statistical likelihood analysis was based on a calculation of R statistics28 for each gene. We selected the genes according to two criteria: (i) representation in testis cDNA libraries (pool A) was two times higher than representation in the other libraries (pool B) and (ii) the R likelihood score was above a preset limit.
Analysis of chromosomal distributions.
We defined the gene density as the fraction of genes satisfying a given selection criterion of the total number of genes present on a chromosome (normalized gene number). We determined relative gene density (ρi) by dividing the gene density on chromosome i by the average gene density on all chromosomes. We used the χ2 test with 19 degrees of freedom for the genomic distribution as a whole or with one degree of freedom for individual chromosomes. We compared the observed number of genes on a given chromosome with the expected number based on a random distribution of all the genes in a data set.
Microarray procedures.
All the animal procedures were carried out according to the National Institutes of Health Guide for the Care and Use of Laboratory Animals. We surgically removed testes and ovaries from mice and stored them at −80 °C until use. We isolated RNA using a Trizol solution (Invitrogen). We used a mixture of total RNAs from 11 different cell lines from several different tissues (universal reference RNA, Stratagene) as reference RNA. We synthesized amino-allyl-modified cDNA from 10–20 μg of DNase-treated RNA using SuperScript II Reverse Transcriptase (Invitrogen), oligo(dT) (Invitrogen) and amino-allyl-modified dUTP (Sigma). We generated fluorescent probes by labeling amino-allyl-modified cDNA with Cy3 or Cy5 (Amersham). We carried out hybridizations on glass slide microarrays containing 15,000 genes from the NIA 15K mouse gene collection29 (Keck Biotechnology Resource Lab at Yale University). All microarray experiments were done in quadruplicate with different mouse pairs and dye-reversed hybridizations. We scanned the chips using the GenePix 4000A scanner (Axon Instruments) and analyzed primary data using Genepix 3.0 software (Axon Instruments). We carried out statistical analysis using a modified t-test implemented in SAM software30. We defined differentially expressed genes at a 1% false discovery rate confidence level and a cutoff for differential expression equal to three for the analysis of tissue specificity and two for Spo11 experiments. We identified early and late spermatogenic genes from a mouse spermatogenesis data set23 as those genes differentially expressed between groups of samples from 1-, 4-, 8-, 11- and 18-, 21-, 26-, 29-d-old mice (t-test, P < 0.01; 3,102 genes are differentially expressed at this confidence level).
GEO accession number.
Microarray data are available under the accession number GSE1138.
Note: Supplementary information is available on the Nature Genetics website.
Accession codes
References
Charlesworth, B. The evolution of chromosomal sex determination. Novartis Found. Symp. 244, 207–219 (2002).
Vallender, E.J. & Lahn, B.T. How mammalian sex chromosomes acquired their peculiar gene content. Bioessays 26, 159–169 (2004).
Rice, W.R. Sex chromosomes and the evolution of sexual dimorphism. Evolution 38, 735–742 (1984).
Fisher, R.A. The evolution of dominance. Biol. Rev. 6, 345–368 (1931).
Rice, W.R. Sexually antagonistic genes: experimental evidence. Science 256, 1436–1439 (1992).
Wang, P.J., Mccarrey, J.R., Yang, F. & Page, D.C. An abundance of X-linked genes expressed in spermatogonia. Nat. Genet. 27, 422–426 (2001).
Saifi, G.M. & Chandra, H.S. An apparent excess of sex- and reproduction-related genes on the human X chromosome. Proc. R. Soc. Lond. B Biol. Sci. 266, 203–209 (1999).
Lercher, M.J., Urrutia, A.O. & Hurst, L.D. Evidence that the human X chromosome is enriched for male-specific but not female-specific genes. Mol. Biol. Evol. 20, 1113–1116 (2003).
Parisi, M. et al. Paucity of genes on the Drosophila X chromosome showing male-biased expression. Science 299, 697–700 (2003).
Reinke, V. et al. A global profile of germline gene expression in C. elegans. Mol. Cell 6, 605–616 (2000).
Ranz, J.M., Castillo-Davis, C.I., Meiklejohn, C.D. & Hartl, D.L. Sex-dependent gene expression and evolution of the Drosophila transcriptome. Science 300, 1742–1745 (2003).
Romanienko, P.J. & Camerini-Otero, R.D. The mouse Spo11 gene is required for meiotic chromosome synapsis. Mol. Cell 6, 975–987 (2000).
Huminiecki, L., Lloyd, A.T. & Wolfe, K.H. Congruence of tissue expression profiles from Gene Expression Atlas, SAGEmap and TissueInfo databases. BMC Genomics 4, 31 (2003).
Su, A.I. et al. Large-scale analysis of the human and mouse transcriptomes. Proc. Natl. Acad. Sci. USA 99, 4465–4470 (2002).
Bono, H. et al. Systematic expression profiling of the mouse transcriptome using RIKEN cDNA microarrays. Genome Res. 13, 1318–1323 (2003).
Fraser, L.R. & Dudley, K. New insights into the t-complex and control of sperm function. Bioessays 21, 304–312 (1999).
Eddy, E.M. Male germ cell gene expression. Recent Prog. Horm. Res. 57, 103–128 (2002).
Bellve, A.R. Purification, culture, and fractionation of spermatogenic cells. Methods Enzymol. 225, 84–113 (1993).
Solari, A.J. The behavior of the XY pair in mammals. Int. Rev. Cytol. 38, 273–317 (1974).
Mckee, B.D. & Handel, M.A. Sex chromosomes, recombination, and chromatin conformation. Chromosoma 102, 71–80 (1993).
Lifschytz, E. & Lindsley, D.L. The role of X-chromosome inactivation during spermatogenesis. Proc. Natl. Acad. Sci. USA 69, 182–186 (1972).
Baudat, F., Manova, K., Yuen, J.P., Jasin, M. & Keeney, S. Chromosome synapsis defects and sexually dimorphic meiotic progression in mice lacking Spo11. Mol. Cell 6, 989–998 (2000).
Schultz, N., Hamra, F.K. & Garbers, D.L. A multitude of genes expressed solely in meiotic or postmeiotic spermatogenic cells offers a myriad of contraceptive targets. Proc. Natl. Acad. Sci. USA 100, 12201–12206 (2003).
Schlecht, U. et al. Expression profiling of Mammalian male meiosis and gametogenesis identifies novel candidate genes for roles in the regulation of fertility. Mol. Biol. Cell 15, 1031–1043 (2004).
Goldstein, P. The synaptonemal complexes of Caenorhabditis elegans: pachytene karyotype analysis of male and hermaphrodite wild-type and him mutants. Chromosoma 86, 577–593 (1982).
Kelly, W.G. et al. X-chromosome silencing in the germline of C. elegans. Development 129, 479–492 (2002).
Huynh, K.D. & Lee, J.T. Inheritance of a pre-inactivated paternal X chromosome in early mouse embryos. Nature 426, 857–862 (2003).
Stekel, D.J., Git, Y. & Falciani, F. The comparison of gene expression from multiple cDNA libraries. Genome Res. 10, 2055–2061 (2000).
Tanaka, T.S. et al. Genome-wide expression profiling of mid-gestation placenta and embryo using a 15,000 mouse developmental cDNA microarray. Proc. Natl. Acad. Sci. USA 97, 9127–9132 (2000).
Tusher, V.G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Acknowledgements
We thank P. Hsieh and B. Oliver for critically reading the manuscript, the members of the Camerini-Otero lab for discussions, L. Robinson and L. Moore for their help and M. Primig for communicating results before publication.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Khil, P., Smirnova, N., Romanienko, P. et al. The mouse X chromosome is enriched for sex-biased genes not subject to selection by meiotic sex chromosome inactivation. Nat Genet 36, 642–646 (2004). https://doi.org/10.1038/ng1368
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1368
This article is cited by
-
The X chromosome and male infertility
Human Genetics (2021)
-
Sex differences in the meiotic behavior of an XX sex chromosome pair in males and females of the mole vole Ellobius tancrei: turning an X into a Y chromosome?
Chromosoma (2021)
-
Dynamic sex chromosome expression in Drosophila male germ cells
Nature Communications (2021)
-
Male mice with large inversions or deletions of X-chromosome palindrome arms are fertile and express their associated genes during post-meiosis
Scientific Reports (2018)
-
Testis Transcriptome Modulation in Klinefelter Patients with Hypospermatogenesis
Scientific Reports (2017)