Abstract
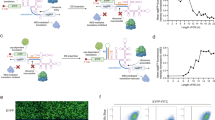
The regulation of cell signaling pathways and the reconstruction of genetic circuits are important aspects of bioengineering research. Both of these goals require molecular devices to transmit information from an input biomacromolecule to the desired outputs. Here, we show that an RNA-protein (RNP)-containing L7Ae–kink-turn interaction can be used to construct translational regulators under control of an input protein that regulates the expression of desired output proteins. We built a system in which L7Ae, an archaeal ribosomal protein, regulates the translation of a designed mRNA in vitro and in human cells. The translational regulator composed of the RNP might provide new therapeutic strategies based on the detection, repair or rewiring of intrinsic cellular defects, and it may also serve as an invaluable tool for the dissection of the behavior of complex, higher-order circuits in the cell.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Gottesman, S. The small RNA regulators of Escherichia coli: roles and mechanisms. Annu. Rev. Microbiol. 58, 303–328 (2004).
Bartel, D.P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Wang, D.O. et al. Synapse- and stimulus-specific local translation during long-term neuronal plasticity. Science 324, 1536–1540 (2009).
St. Johnston, D. Moving messages: the intracellular localization of mRNAs. Nat. Rev. Mol. Cell Biol. 6, 363–375 (2005).
Hentze, M.W. et al. Identification of the iron-responsive element for the translational regulation of human ferritin mRNA. Science 238, 1570–1573 (1987).
Ray, P.S. et al. A stress-responsive RNA switch regulates VEGFA expression. Nature 457, 915–919 (2009).
Endy, D. Foundations for engineering biology. Nature 438, 449–453 (2005).
Isaacs, F.J., Dwyer, D.J. & Collins, J.J. RNA synthetic biology. Nat. Biotechnol. 24, 545–554 (2006).
Drubin, D.A., Way, J.C. & Silver, P.A. Designing biological systems. Genes Dev. 21, 242–254 (2007).
Purnick, P.E. & Weiss, R. The second wave of synthetic biology: from modules to systems. Nat. Rev. Mol. Cell Biol. 10, 410–422 (2009).
Win, M.N., Liang, J.C. & Smolke, C.D. Frameworks for programming biological function through RNA parts and devices. Chem. Biol. 16, 298–310 (2009).
Dueber, J.E., Yeh, B.J., Bhattacharyya, R.P. & Lim, W.A. Rewiring cell signaling: the logic and plasticity of eukaryotic protein circuitry. Curr. Opin. Struct. Biol. 14, 690–699 (2004).
Winkler, W.C. Riboswitches and the role of noncoding RNAs in bacterial metabolic control. Curr. Opin. Chem. Biol. 9, 594–602 (2005).
Gallivan, J.P. Toward reprogramming bacteria with small molecules and RNA. Curr. Opin. Chem. Biol. 11, 612–619 (2007).
Sharma, V., Nomura, Y. & Yokobayashi, Y. Engineering complex riboswitch regulation by dual genetic selection. J. Am. Chem. Soc. 130, 16310–16315 (2008).
Win, M.N. & Smolke, C.D. Higher-order cellular information processing with synthetic RNA devices. Science 322, 456–460 (2008).
Suess, B. & Weigand, J.E. Engineered riboswitches—overview, problems and trends. RNA Biol. 5, 24–29 (2008).
Yamauchi, T. et al. Riboswitches for enhancing target gene expression in eukaryotes. ChemBioChem 9, 1040–1043 (2008).
Ogawa, A. & Maeda, M. An artificial aptazyme-based riboswitch and its cascading system in E. coli. ChemBioChem 9, 206–209 (2008).
Topp, S. & Gallivan, J.P. Riboswitches in unexpected places—a synthetic riboswitch in a protein coding region. RNA 14, 2498–2503 (2008).
Wieland, M. & Hartig, J.S. Improved aptazyme design and in vivo screening enable riboswitching in bacteria. Angew. Chem. Int. Edn Engl. 47, 2604–2607 (2008).
Stripecke, R. & Hentze, M.W. Bacteriophage and spliceosomal proteins function as position-dependent cis/trans repressors of mRNA translation in vitro. Nucleic Acids Res. 20, 5555–5564 (1992).
Paraskeva, E., Atzberger, A. & Hentze, M.W. A translational repression assay procedure (TRAP) for RNA-protein interactions in vivo. Proc. Natl. Acad. Sci. USA 95, 951–956 (1998).
Nie, M. & Htun, H. Different modes and potencies of translational repression by sequence-specific RNA-protein interaction at the 5′-UTR. Nucleic Acids Res. 34, 5528–5540 (2006).
Atsumi, S., Ikawa, Y., Shiraishi, H. & Inoue, T. Design and development of a catalytic ribonucleoprotein. EMBO J. 20, 5453–5460 (2001).
Saito, H. & Inoue, T. RNA and RNP as new molecular parts in synthetic biology. J. Biotechnol. 132, 1–7 (2007).
Saito, H. & Inoue, T. Synthetic biology with RNA motifs. Int. J. Biochem. Cell Biol. 41, 398–404 (2009).
Shimizu, Y. et al. Cell-free translation reconstituted with purified components. Nat. Biotechnol. 19, 751–755 (2001).
Szostak, J.W., Bartel, D.P. & Luisi, P.L. Synthesizing life. Nature 409, 387–390 (2001).
Saito, H. et al. Time-resolved tracking of a minimum gene expression system reconstituted in giant liposomes. ChemBioChem 10, 1640–1643 (2009).
Rozhdestvensky, T.S. et al. Binding of L7Ae protein to the K-turn of archaeal snoRNAs: a shared RNA binding motif for C/D and H/ACA box snoRNAs in Archaea. Nucleic Acids Res. 31, 869–877 (2003).
Moore, T., Zhang, Y., Fenley, M.O. & Li, H. Molecular basis of box C/D RNA-protein interactions; cocrystal structure of archaeal L7Ae and a box C/D RNA. Structure 12, 807–818 (2004).
Turner, B., Melcher, S.E., Wilson, T.J., Norman, D.G. & Lilley, D.M. Induced fit of RNA on binding the L7Ae protein to the kink-turn motif. RNA 11, 1192–1200 (2005).
Klein, D.J., Schmeing, T.M., Moore, P.B. & Steitz, T.A. The kink-turn: a new RNA secondary structure motif. EMBO J. 20, 4214–4221 (2001).
Matsumura, S., Ikawa, Y. & Inoue, T. Biochemical characterization of the kink-turn RNA motif. Nucleic Acids Res. 31, 5544–5551 (2003).
Hamma, T. & Ferre-D'Amare, A.R. Structure of protein L7Ae bound to a K-turn derived from an archaeal box H/ACA sRNA at 1.8 A resolution. Structure 12, 893–903 (2004).
Nolivos, S., Carpousis, A.J. & Clouet-d'Orval, B. The K-loop, a general feature of the Pyrococcus C/D guide RNAs, is an RNA structural motif related to the K-turn. Nucleic Acids Res. 33, 6507–6514 (2005).
Reichow, S.L., Hamma, T., Ferre-D'Amare, A.R. & Varani, G. The structure and function of small nucleolar ribonucleoproteins. Nucleic Acids Res. 35, 1452–1464 (2007).
Macias, S., Bragulat, M., Tardiff, D.F. & Vilardell, J. L30 binds the nascent RPL30 transcript to repress U2 snRNP recruitment. Mol. Cell 30, 732–742 (2008).
Chavatte, L., Brown, B.A. & Driscoll, D.M. Ribosomal protein L30 is a component of the UGA-selenocysteine recoding machinery in eukaryotes. Nat. Struct. Mol. Biol. 12, 408–416 (2005).
Gebauer, F. & Hentze, M.W. Molecular mechanisms of translational control. Nat. Rev. Mol. Cell Biol. 5, 827–835 (2004).
Babitzke, P., Baker, C.S. & Romeo, T. Regulation of translation initiation by RNA binding proteins. Annu. Rev. Microbiol. 63, 27–44 (2009).
Katoh, T. & Suzuki, T. Specific residues at every third position of siRNA shape its efficient RNAi activity. Nucleic Acids Res. 35, e27 (2007).
Weber, W. & Fussenegger, M. Engineering of synthetic mammalian gene networks. Chem. Biol. 16, 287–297 (2009).
Kaern, M. & Weiss, R. System Modeling in Cellular Biology Ch. 13 (The MIT Press, Cambridge, Massachusetts, 2006).
Niwa, H., Miyazaki, J. & Smith, A.G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet. 24, 372–376 (2000).
Acknowledgements
The authors thank Y. Shimizu (University of Tokyo) for providing us with the PURE system solutions, A. Huttenhofer (Innsbruck Medical University, Biocenter) and T.S. Rozhdestvensky (University of Muenster) for providing us with the L7Ae plasmids, and S. Kashida (Kyoto University) for providing us with the p481 and pSk-7 plasmids. The authors also thank A. Kitamura (JST) for RNP structural modeling. This work was supported by a Grant-in-Aid for Young Scientists (A) (H.S.) and the International Cooperative Research Project, Japan Science and Technology Agency (H.S. and T.I.).
Author information
Authors and Affiliations
Contributions
H.S., T.K. and T.I. designed the experiments and analyzed the results. H.S., T.K., T.H. and R.F. performed the in vitro translation assays and the RNP interaction assays. Y.F. and K.H. performed the in vivo translation assays. H.S. and T.I. wrote the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 and Supplementary Methods (PDF 5157 kb)
Rights and permissions
About this article
Cite this article
Saito, H., Kobayashi, T., Hara, T. et al. Synthetic translational regulation by an L7Ae–kink-turn RNP switch. Nat Chem Biol 6, 71–78 (2010). https://doi.org/10.1038/nchembio.273
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.273
This article is cited by
-
Engineered poly(A)-surrogates for translational regulation and therapeutic biocomputation in mammalian cells
Cell Research (2024)
-
PERSIST platform provides programmable RNA regulation using CRISPR endoRNases
Nature Communications (2022)
-
Regulated control of gene therapies by drug-induced splicing
Nature (2021)
-
Cell-free gene expression: an expanded repertoire of applications
Nature Reviews Genetics (2020)
-
Engineering protein-protein devices for multilayered regulation of mRNA translation using orthogonal proteases in mammalian cells
Nature Communications (2018)