Abstract
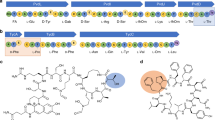
Pyrrolysyl-tRNA synthetase (PylRS) is a major tool in genetic code expansion using noncanonical amino acids, yet its structure and function are not completely understood. Here we describe the crystal structure of the previously uncharacterized essential N-terminal domain of this unique enzyme in complex with tRNAPyl. This structure explains why PylRS remains orthogonal in a broad range of organisms, from bacteria to humans. The structure also illustrates why tRNAPyl recognition by PylRS is anticodon independent: the anticodon does not contact the enzyme. Then, using standard microbiological culture equipment, we established a new method for laboratory evolution—a noncontinuous counterpart of the previously developed phage-assisted continuous evolution. With this method, we evolved novel PylRS variants with enhanced activity and amino acid specificity. Finally, we employed an evolved PylRS variant to determine its N-terminal domain structure and show how its mutations improve PylRS activity in the genetic encoding of a noncanonical amino acid.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Mukai, T. et al. RNA-dependent cysteine biosynthesis in bacteria and archaea. MBio 8, e00561–17 (2017).
Wan, W., Tharp, J.M. & Liu, W.R. Pyrrolysyl-tRNA synthetase: an ordinary enzyme but an outstanding genetic code expansion tool. Biochim. Biophys. Acta 1844, 1059–1070 (2014).
Crnković, A., Suzuki, T., Söll, D. & Reynolds, N.M. Pyrrolysyl-tRNA synthetase, an aminoacyl-tRNA synthetase for genetic code expansion. Croat. Chem. Acta 89, 163–174 (2016).
Gaston, M.A., Jiang, R. & Krzycki, J.A. Functional context, biosynthesis, and genetic encoding of pyrrolysine. Curr. Opin. Microbiol. 14, 342–349 (2011).
Mukai, T. et al. Adding L-lysine derivatives to the genetic code of mammalian cells with engineered pyrrolysyl-tRNA synthetases. Biochem. Biophys. Res. Commun. 371, 818–822 (2008).
Chin, J.W. Expanding and reprogramming the genetic code of cells and animals. Annu. Rev. Biochem. 83, 379–408 (2014).
Ambrogelly, A. et al. Pyrrolysine is not hardwired for cotranslational insertion at UAG codons. Proc. Natl. Acad. Sci. USA 104, 3141–3146 (2007).
Ho, J.M. et al. Efficient reassignment of a frequent serine codon in wild-type Escherichia coli. ACS Synth. Biol. 5, 163–171 (2016).
Yanagisawa, T., Umehara, T., Sakamoto, K. & Yokoyama, S. Expanded genetic code technologies for incorporating modified lysine at multiple sites. ChemBioChem 15, 2181–2187 (2014).
Guo, L.T. et al. Polyspecific pyrrolysyl-tRNA synthetases from directed evolution. Proc. Natl. Acad. Sci. USA 111, 16724–16729 (2014).
Jiang, R. & Krzycki, J.A. PylSn and the homologous N-terminal domain of pyrrolysyl-tRNA synthetase bind the tRNA that is essential for the genetic encoding of pyrrolysine. J. Biol. Chem. 287, 32738–32746 (2012).
Herring, S. et al. The amino-terminal domain of pyrrolysyl-tRNA synthetase is dispensable in vitro but required for in vivo activity. FEBS Lett. 581, 3197–3203 (2007).
Yanagisawa, T., Ishii, R., Fukunaga, R., Nureki, O. & Yokoyama, S. Crystallization and preliminary X-ray crystallographic analysis of the catalytic domain of pyrrolysyl-tRNA synthetase from the methanogenic archaeon Methanosarcina mazei. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 62, 1031–1033 (2006).
Kavran, J.M. et al. Structure of pyrrolysyl-tRNA synthetase, an archaeal enzyme for genetic code innovation. Proc. Natl. Acad. Sci. USA 104, 11268–11273 (2007).
Nozawa, K. et al. Pyrrolysyl-tRNA synthetase-tRNAPyl structure reveals the molecular basis of orthogonality. Nature 457, 1163–1167 (2009).
Esvelt, K.M., Carlson, J.C. & Liu, D.R. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011).
Badran, A.H. et al. Continuous evolution of Bacillus thuringiensis toxins overcomes insect resistance. Nature 533, 58–63 (2016).
Dickinson, B.C., Packer, M.S., Badran, A.H. & Liu, D.R. A system for the continuous directed evolution of proteases rapidly reveals drug-resistance mutations. Nat. Commun. 5, 5352 (2014).
Bryson, D. et al. Continuous directed evolution of aminoacyl-tRNA synthetases. Nat Chem. Biol. 13 http://dx.doi.org/10.1038/nchembio.2474 (2017).
Meyer, J.R. et al. Repeatability and contingency in the evolution of a key innovation in phage lambda. Science 335, 428–432 (2012).
Hammerling, M.J. et al. Bacteriophages use an expanded genetic code on evolutionary paths to higher fitness. Nat. Chem. Biol. 10, 178–180 (2014).
Badran, A.H. & Liu, D.R. Development of potent in vivo mutagenesis plasmids with broad mutational spectra. Nat. Commun. 6, 8425 (2015).
Mogk, A. et al. Roles of individual domains and conserved motifs of the AAA+ chaperone ClpB in oligomerization, ATP hydrolysis, and chaperone activity. J. Biol. Chem. 278, 17615–17624 (2003).
Yamamoto, H. et al. 70S-scanning initiation is a novel and frequent initiation mode of ribosomal translation in bacteria. Proc. Natl. Acad. Sci. USA 113, E1180–E1189 (2016).
Owens, A.E., Grasso, K.T., Ziegler, C.A. & Fasan, R. Two-tier screening platform for directed evolution of aminoacyl-tRNA synthetases with enhanced stop codon suppression efficiency. ChemBioChem 18, 1109–1116 (2017).
Yanagisawa, T. et al. Multistep engineering of pyrrolysyl-tRNA synthetase to genetically encode Nɛ-(o-azidobenzyloxycarbonyl) lysine for site-specific protein modification. Chem. Biol. 15, 1187–1197 (2008).
Sorokin, D.Y. et al. Discovery of extremely halophilic, methyl-reducing euryarchaea provides insights into the evolutionary origin of methanogenesis. Nat. Microbiol. 2, 17081 (2017).
O'Donoghue, P., Ling, J., Wang, Y.S. & Söll, D. Upgrading protein synthesis for synthetic biology. Nat. Chem. Biol. 9, 594–598 (2013).
Wolfson, A.D., Pleiss, J.A. & Uhlenbeck, O.C. A new assay for tRNA aminoacylation kinetics. RNA 4, 1019–1023 (1998).
Schachtele, C.F., Anderson, D.L. & Rogers, P. Mechanism of canavanine death in Escherichia coli. II. Membrane-bound canavanyl-protein and nuclear disruption. J. Mol. Biol. 33, 861–872 (1968).
Fan, C., Ho, J.M.L., Chirathivat, N., Söll, D. & Wang, Y.S. Exploring the substrate range of wild-type aminoacyl-tRNA synthetases. ChemBioChem 15, 1805–1809 (2014).
Hong, K.W. et al. Transfer RNA-dependent cognate amino acid recognition by an aminoacyl-tRNA synthetase. EMBO J. 15, 1983–1991 (1996).
Gresham, D. & Dunham, M.J. The enduring utility of continuous culturing in experimental evolution. Genomics 104 6 Pt A, 399–405 (2014).
Dickinson, B.C., Leconte, A.M., Allen, B., Esvelt, K.M. & Liu, D.R. Experimental interrogation of the path dependence and stochasticity of protein evolution using phage-assisted continuous evolution. Proc. Natl. Acad. Sci. USA 110, 9007–9012 (2013).
Suzuki, T., Yamashita, K., Tanaka, Y., Tanaka, I. & Yao, M. Crystallization and preliminary X-ray crystallographic analysis of a bacterial Asn-transamidosome. Acta Crystallogr. F Struct. Biol. Commun. 70, 790–793 (2014).
Easton, L.E., Shibata, Y. & Lukavsky, P.J. Rapid, nondenaturing RNA purification using weak anion-exchange fast performance liquid chromatography. RNA 16, 647–653 (2010).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Terwilliger, T.C. et al. Decision-making in structure solution using Bayesian estimates of map quality: the PHENIX AutoSol wizard. Acta Crystallogr. D Biol. Crystallogr. 65, 582–601 (2009).
Terwilliger, T.C. et al. Iterative model building, structure refinement and density modification with the PHENIX AutoBuild wizard. Acta Crystallogr. D Biol. Crystallogr. 64, 61–69 (2008).
Afonine, P.V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Bricogne, G. et al. BUSTER v. 2.10.2 (Global Phasing Ltd., Cambridge, UK, 2016).
Yamashita, K., Zhou, Y., Tanaka, I. & Yao, M. New model-fitting and model-completion programs for automated iterative nucleic acid refinement. Acta Crystallogr. D Biol. Crystallogr. 69, 1171–1179 (2013).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Wong, M.L., Guzei, I.A. & Kiessling, L.L. An asymmetric synthesis of L-pyrrolysine. Org. Lett. 14, 1378–1381 (2012).
Carlson, J.C., Badran, A.H., Guggiana-Nilo, D.A. & Liu, D.R. Negative selection and stringency modulation in phage-assisted continuous evolution. Nat. Chem. Biol. 10, 216–222 (2014).
Hubbard, B.P. et al. Continuous directed evolution of DNA-binding proteins to improve TALEN specificity. Nat. Methods 12, 939–942 (2015).
Acknowledgements
The authors thank S. Melnikov and Y. Xiong (Yale University) for insightful discussions and intellectual contributions, A. Shinoda (Paul Scherrer Institute) and K. Yamashita (RIKEN) for advice on structure analysis, and S. Trauger (Small Molecule Mass Spectrometry Laboratory at Harvard University) for providing expertise with intact protein mass spectrometry analysis. This work was supported the US National Institutes of Health (NIH) R01EB022376 and R35GM118062 (to D.R.L.), and R01GM022854 and R35GM122560 (to D.S.), by the Defense Advanced Research Projects Agency N66001-12-C-4207 (to D.R.L.), by the Department of Energy DE-FG02-98ER20311 (to D.S.), and the Howard Hughes Medical Institute. D.I.B is supported by the National Institutes of Health under a Ruth L. Kirschstein National Research Service Award (F32GM106621). This research used resources of the Advanced Photon Source, a US Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357.
Author information
Authors and Affiliations
Contributions
T.S. purified and crystallized the PylRS–tRNAPyl complexes, solved structures, and analyzed data. C.M. designed the PANCE research, performed experiments, analyzed data, and wrote the manuscript. D.S. designed and supervised the research and wrote the manuscript. D.R.L. designed and supervised the research and edited the manuscript. L.-T.G. designed the chimeric chPylRS variant for evolution in PANCE, performed protein purification and in vitro aminoacylation assays, and analyzed data. J.M.L.H. assisted with design and refinement of PANCE procedure. D.I.B. established initial selection conditions, performed read-through assays, and performed mass spectrometry and western blot analyses. All authors contributed to editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–9, Supplementary Tables 1–4, and Supplementary Note (PDF 1893 kb)
Rights and permissions
About this article
Cite this article
Suzuki, T., Miller, C., Guo, LT. et al. Crystal structures reveal an elusive functional domain of pyrrolysyl-tRNA synthetase. Nat Chem Biol 13, 1261–1266 (2017). https://doi.org/10.1038/nchembio.2497
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2497
This article is cited by
-
An evolved pyrrolysyl-tRNA synthetase with polysubstrate specificity expands the toolbox for engineering enzymes with incorporation of noncanonical amino acids
Bioresources and Bioprocessing (2023)
-
Quintuply orthogonal pyrrolysyl-tRNA synthetase/tRNAPyl pairs
Nature Chemistry (2023)
-
Structural basis for a degenerate tRNA identity code and the evolution of bimodal specificity in human mitochondrial tRNA recognition
Nature Communications (2023)
-
Systematic molecular evolution enables robust biomolecule discovery
Nature Methods (2022)
-
Restoration of dystrophin expression in mice by suppressing a nonsense mutation through the incorporation of unnatural amino acids
Nature Biomedical Engineering (2021)