Abstract
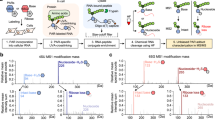
RNA G-quadruplex (G4) structures are thought to affect biological processes, including translation and pre-mRNA splicing, but it is not possible at present to demonstrate that they form naturally at specific sequences in long functional RNA molecules. We developed a new strategy, footprinting of long 7-deazaguanine-substituted RNAs (FOLDeR), that allows the formation of G4s to be confirmed in long RNAs and under functional conditions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Huppert, J.L. FEBS J. 277, 3452–3458 (2010).
Biffi, G., Tannahill, D., McCafferty, J. & Balasubramanian, S. Nat. Chem. 5, 182–186 (2013).
Rhodes, D. & Lipps, H.J. Nucleic Acids Res. 43, 8627–8637 (2015).
Biffi, G., Di Antonio, M., Tannahill, D. & Balasubramanian, S. Nat. Chem. 6, 75–80 (2014).
Agarwala, P., Pandey, S. & Maiti, S. Org. Biomol. Chem. 13, 5570–5585 (2015).
Millevoi, S., Moine, H. & Vagner, S. WIREs RNA 3, 495–507 (2012).
Kumari, S., Bugaut, A., Huppert, J.L. & Balasubramanian, S. Nat. Chem. Biol. 3, 218–221 (2007).
Darnell, J.C. et al. Cell 107, 489–499 (2001).
Decorsière, A., Cayrel, A., Vagner, S. & Millevoi, S. Genes Dev. 25, 220–225 (2011).
Wolfe, A.L. et al. Nature 513, 65–70 (2014).
Marcel, V. et al. Carcinogenesis 32, 271–278 (2011).
Ribeiro, M.M. et al. Hum. Genet. 134, 37–44 (2015).
Smith, L.D. et al. Cell Rep. 9, 193–205 (2014).
Zizza, P. et al. Nucleic Acids Res. 44, 1579–1590 (2016).
Munroe, S.H., Morales, C.H., Duyck, T.H. & Waters, P.D. PLoS One 10, e0137893 (2015).
Murchie, A.I. & Lilley, D.M. Nucleic Acids Res. 20, 49–53 (1992).
Boise, L.H. et al. Cell 74, 597–608 (1993).
Garneau, D., Revil, T., Fisette, J.-F. & Chabot, B. J. Biol. Chem. 280, 22641–22650 (2005).
Kikin, O., D'Antonio, L. & Bagga, P.S. Nucleic Acids Res. 34, W676–W682 (2006).
Zuker, M. Nucleic Acids Res. 31, 3406–3415 (2003).
Merino, E.J., Wilkinson, K.A., Coughlan, J.L. & Weeks, K.M. J. Am. Chem. Soc. 127, 4223–4231 (2005).
Kwok, C.K., Ding, Y., Tang, Y., Assmann, S.M. & Bevilacqua, P.C. Nat. Commun. 4, 2971 (2013).
Phan, A.T. et al. Nat. Struct. Mol. Biol. 18, 796–804 (2011).
Huang, H. et al. Nat. Chem. Biol. 10, 686–691 (2014).
Warner, K.D. et al. Nat. Struct. Mol. Biol. 21, 658–663 (2014).
Acknowledgements
This work was supported by a Medical Research Council Career Development Award (G1000526) to C.D. and a Sir Dudley Spurling Post Graduate Scholarship from the Bank of Butterfield Foundation in Bermuda to C.W. This work was also funded by CNRS and Lorraine University (UMR 7365 and previously 7214) and the European Alternative Splicing Network of Excellence (EURASNET, FP6 life sciences, genomics and biotechnology for health; LSHG-CT-2005-518238).
Author information
Authors and Affiliations
Contributions
C.W. performed all experiments under the guidance of I.C.E. and C.D. Footprinting experiments and structural model calculation were performed under the guidance of I.B.-A. and C.B., G.A.B. and L.H.H. contributed to the development and the validation of the strategy. C.W., I.C.E. and C.D. interpreted the results and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Figures 1 – 11. (PDF 13506 kb)
Supplementary Data set 1
SAFA quantification of Bcl-x-681 native footprinting. (XLSX 136 kb)
Supplementary Data set 2
Comparison of footprinting on Bcl-x-681 native and 7-deaza-G-substituted transcripts. (XLSX 122 kb)
Supplementary Data set 3
Comparison of footprinting on Bcl-x-681 in either Potassium- or lithium-containing buffers. (XLSX 131 kb)
Rights and permissions
About this article
Cite this article
Weldon, C., Behm-Ansmant, I., Hurley, L. et al. Identification of G-quadruplexes in long functional RNAs using 7-deazaguanine RNA. Nat Chem Biol 13, 18–20 (2017). https://doi.org/10.1038/nchembio.2228
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2228