Abstract
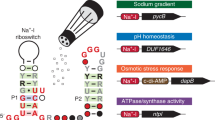
Cyclic di-adenosine monophosphate (c-di-AMP) is a recently discovered bacterial second messenger implicated in the control of cell wall metabolism, osmotic stress responses and sporulation. However, the mechanisms by which c-di-AMP triggers these physiological responses have remained largely unknown. Notably, a candidate riboswitch class called ydaO associates with numerous genes involved in these same processes. Although a representative ydaO motif RNA recently was reported to weakly bind ATP, we report that numerous members of this noncoding RNA class selectively respond to c-di-AMP with subnanomolar affinity. Our findings resolve the mystery regarding the primary ligand for this extremely common riboswitch class and expose a major portion of the super-regulon of genes that are controlled by the widespread bacterial second messenger c-di-AMP.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Römling, U. Great times for small molecules: c-di-AMP, a second messenger candidate in bacteria and archaea. Sci. Signal. 1, pe39 (2008).
Pesavento, C. & Hengge, R. Bacterial nucleotide-based second messengers. Curr. Opin. Microbiol. 12, 170–176 (2009).
Sondermann, H., Shikuma, N.J. & Yildiz, F.H. You've come a long way: c-di-GMP signaling. Curr. Opin. Microbiol. 15, 140–146 (2012).
Krasteva, P.V., Giglio, K.M. & Sondermann, H. Sensing the messenger: the diverse ways that bacteria signal through c-di-GMP. Protein Sci. 21, 929–948 (2012).
Sudarsan, N. et al. Riboswitches in eubacteria sense the second messenger c-di-GMP. Science 321, 411–413 (2008).
Lee, E.R., Baker, J.L., Weinberg, Z., Sudarsan, N. & Breaker, R.R. An allosteric self-splicing ribozyme triggered by a bacterial second messenger. Science 329, 845–848 (2010).
Witte, G., Hartung, S., Buttner, K. & Hopfner, K.P. Structural biochemistry of a bacterial checkpoint protein reveals diadenylate cyclase activity regulated by DNA recombination intermediates. Mol. Cell 30, 167–178 (2008).
Oppenheimer-Shaanan, Y., Wexselblatt, E., Katzhendler, J., Yavin, E. & Ben-Yehuda, S. c-di-AMP reports DNA integrity during sporulation in Bacillus subtilis. EMBO Rep. 12, 594–601 (2011).
Corrigan, R.M., Abbott, J.C., Burhenne, H., Kaever, V. & Gründling, A. c-di-AMP is a new second messenger in Staphylococcus aureus with a role in controlling cell size and envelope stress. PLoS Pathog. 7, e1002217 (2011).
Smith, W.M. et al. Heat resistance and salt hypersensitivity in Lactococcus lactis due to spontaneous mutation of limg_1816 (gdpP) induced by high temperature growth. Appl. Environ. Microbiol. 78, 7753–7759 (2012).
Corrigan, R.M. et al. Systematic identification of conserved bacteria c-di-AMP receptor proteins. Proc. Natl. Acad. Sci. USA 110, 9084–9089 (2013).
Baker, J.L. et al. Widespread genetic switches and toxicity resistance proteins for fluoride. Science 335, 233–235 (2012).
Barrick, J.E. et al. New RNA motifs suggest an expanded scope for riboswitches in bacterial genetic control. Proc. Natl. Acad. Sci. USA 101, 6421–6426 (2004).
Mandal, M. et al. A glycine-dependent riboswitch that uses cooperative binding to control gene expression. Science 306, 275–279 (2004).
Winkler, W.C., Nahvi, A., Roth, A., Collins, J.A. & Breaker, R.R. Control of gene expression by a natural metabolite-responsive ribozyme. Nature 428, 281–286 (2004).
Roth, A. et al. A riboswitch selective for the queuosine precursor preQ1 contains an unusually small aptamer domain. Nat. Struct. Mol. Biol. 14, 308–317 (2007).
Dann, C.E. III et al. Structure and mechanism of a metal-sensing regulatory RNA. Cell 130, 878–892 (2007).
Marchais, A., Duperrier, S., Durand, S., Gautheret, D. & Stragier, P. CsfG, a sporulation-specific, small non-coding RNA highly conserved in endospore formers. RNA Biol. 8, 358–364 (2011).
Block, K.F., Hammond, M.C. & Breaker, R.R. Evidence for widespread gene control function by the ydaO riboswitch candidate. J. Bacteriol. 192, 3983–3989 (2010).
Breaker, R.R. Riboswitches and the RNA world. Cold Spring Harb. Perspect. Biol. 4, a003566 (2012).
Meyer, M.M. et al. Challenges of ligand identification for riboswitch candidates. RNA Biol. 8, 5–10 (2011).
Breaker, R.R. Prospects for riboswitch discovery and analysis. Mol. Cell 43, 867–879 (2011).
Watson, P.Y. & Fedor, M.J. The ydaO motif is an ATP-sensing riboswitch in Bacillus subtilis. Nat. Chem. Biol. 8, 963–965 (2012).
Soukup, G.A. & Breaker, R.R. Relationship between internucleotide linkage geometry and the stability of RNA. RNA 5, 1308–1325 (1999).
Regulski, E.E. & Breaker, R.R. In-line probing analysis of riboswitches. Methods Mol. Biol. 419, 53–67 (2008).
Smith, K.D. et al. Structural basis of ligand binding by a c-di-GMP riboswitch. Nat. Struct. Mol. Biol. 16, 1218–1223 (2009).
Shanahan, C.A., Gaffney, B.L., Jones, R.A. & Strobel, S.A. Identification of c-di-GMP derivatives resistance to an EAL domain phosphodiesterase. Biochemistry 52, 365–377 (2012).
McDaniel, B.A., Grundy, F.J., Artsimovitch, I. & Henkin, T.M. Transcription termination control of the S box system: direct measurement of S-adenosylmethionine by the leader RNA. Proc. Natl. Acad. Sci. USA 100, 3083–3088 (2003).
Epshtein, V., Mironov, A.S. & Nudler, E. The riboswitch-mediated control of sulfur metabolism in bacteria. Proc. Natl. Acad. Sci. USA 100, 5052–5056 (2003).
Sudarsan, N., Wickiser, J.K., Nakamura, S., Ebert, M.S. & Breaker, R.R. An mRNA structure in bacteria that controls gene expression by binding lysine. Genes Dev. 17, 2688–2697 (2003).
Mehne, F.M. et al. Cyclic di-AMP homeostasis in Bacillus subtilis; both lack and high level accumulation of the nucleotide are detrimental for cell growth. J. Biol. Chem. 288, 2004–2017 (2013).
Moriyama, R. et al. Expression of a germination-specific amidase, SleB, of Bacilli in the forespore compartment of sporulating cells and its localization on the exterior side of the cortex in dormant spores. J. Bacteriol. 181, 2373–2378 (1999).
Gilmore, M.E., Bandyopadhyay, D., Dean, A.M., Linnstaedt, S.D. & Popham, D.L. Production of muramic δ-lactam in Bacillus subtilis spore peptidoglycan. J. Bacteriol. 186, 80–89 (2004).
Kana, B.D. et al. The resuscitation-promoting factors of Mycobacterium tuberculosis are required for virulence and resuscitation from dormancy but are collectively dispensable for growth in vitro. Mol. Microbiol. 67, 672–684 (2008).
Kana, B.D. & Mizrahi, V. Resuscitation-promoting factors as lytic enzymes for bacterial growth and signaling. FEMS Immunol. Med. Microbiol. 58, 39–50 (2010).
Klein, D.J., Edwards, T.E. & Ferré-D'Amaré, A.R. Cocrystal structure of a class I preQ1 riboswitch reveals a pseudoknot recognizing an essential hypermodified nucleobase. Nat. Struct. Mol. Biol. 16, 343–344 (2009).
Kang, M., Peterson, R. & Feigon, J. Structural insights into riboswitch control of the biosynthesis of queuosine, a modified nucleotide found in the anticodon of tRNA. Mol. Cell 33, 784–790 (2009).
Ren, A., Rajashankar, K.R. & Patel, D.J. Fluoride ion encapsulation by Mg2+ ions and phosphates in a fluoride riboswitch. Nature 486, 85–89 (2012).
Wickiser, J.K., Winkler, W.C., Breaker, R.R. & Crothers, D.M. The speed of RNA transcription and metabolite binding kinetics operate an FMN riboswitch. Mol. Cell 18, 49–60 (2005).
Wickiser, J.K., Cheah, M.T., Breaker, R.R. & Crothers, D.M. The kinetics of ligand binding by an adenosine-sensing riboswitch. Biochemistry 44, 13404–13414 (2005).
Gilbert, S.D., Stoddard, C.D., Wise, S.J. & Batey, R.T. Thermodynamic and kinetic characterization of ligand binding to the purine riboswitch aptamer domain. J. Mol. Biol. 359, 754–768 (2006).
Lemay, J.-F. et al. Comparative study between transcriptionally- and translationally-acting adenine riboswitches reveals key differences in riboswitch regulatory mechanisms. PLoS Genet. 7, e1001278 (2011).
Haller, A., Soulière, M.F. & Micura, R. The dynamic nature of RNA as key to understanding riboswitch mechanisms. Acc. Chem. Res. 44, 1339–1348 (2011).
Frieda, K.L. & Block, S.M. Direct observation of cotranscriptional folding in an adenine riboswitch. Science 338, 397–400 (2012).
Edwards, A.L., Reyes, F.E., Héroux, A. & Batey, R.T. Structural basis for recognition of S-adenylhomocysteine by riboswitches. RNA 16, 2144–2155 (2010).
Wang, J.X., Lee, E.R., Morales, D.R., Lim, J. & Breaker, R.R. Riboswitches that sense S-adenylhomocysteine and activate genes involved in coenzyme recycling. Mol. Cell 29, 691–702 (2008).
Woodward, J.J., Iavarone, A.T. & Portnoy, D.A. c-di-AMP secreted by intracellular Listeria monocytogenes activates a host type I interferon response. Science 328, 1703–1705 (2010).
Markley, J.L. et al. New bioinformatics resources for metabolomics. Pac. Symp. Biocomput. 2007, 157–168 (2007).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular datasets. Nucleic Acids Res. 40, D109–D114 (2012).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Wach, A. PCR-synthesis of marker cassettes with long flanking homology regions for gene disruptions in S. cerevisiae. Yeast 12, 259–265 (1996).
Vidal-Aroca, F. et al. One-step high-throughput assay for quantitative detection of β-galactosidase activity in intact Gram-negative bacteria, yeast, and mammalian cells. Biotechniques 40, 433–434 (2006).
Barrick, J.E. & Breaker, R.R. The distributions, mechanisms, and structures of metabolite-binding riboswitches. Genome Biol. 8, R239 (2007).
Nawrocki, E.P., Kolbe, D.L. & Eddy, S.R. Infernal 1.0: inference of RNA alignments. Bioinformatics 25, 1335–1337 (2009).
Hofacker, I.L. Curr. Prot. Bioinformatics RNA secondary structure analysis using the Vienna RNA package. 26, 12.2 (2009).
Pruitt, K.D., Tatusova, T. & Maglott, D.R. NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts, and proteins. Nucleic Acids Res. 35, D61–D65 (2007).
Gevers, D. et al. The human microbiome project: a community resource for the healthy human microbiome. PLoS Biol. 10, e1001377 (2012).
Markowitz, V.M. et al. IMG/M: a data management and analysis system for metagenomes. Nucleic Acids Res. 36, D534–D538 (2008).
Meyer, F. et al. The metagenomics RAST server—a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics 9, 386–393 (2008).
Sun, S. et al. Community cyberinfrastructure for advanced microbial ecology research and analysis: the CAMERA resource. Nucleic Acids Res. 39, D546–D551 (2011).
Benson, D.A., Karsch-Mizrachi, I., Lipman, D.J., Ostell, J. & Wheeler, D.L. Genbank. Nucleic Acids Res. 36, D25–D30 (2008).
Roth, A. et al. A novel class of self-cleaving ribozymes is present in many species of bacteria and eukarya. Nat. Chem. Biol. (in the press).
Gerstein, M., Sonnhammer, E.L.L. & Chothia, C. Volume changes on protein evolution. J. Mol. Biol. 236, 1067–1078 (1994).
Acknowledgements
We thank A. Roth and other members of the Breaker laboratory for helpful discussions. We also thank N. Carriero and R. Bjornson for assisting our use of the Yale Life Sciences High Performance Computing Center (NIH grant RR19895-02). K.F. was supported by a Japan Society for the Promotion of Science fellowship for research abroad. This project was supported by US National Institutes of Health grants (GM022778 and DE022340) to R.R.B. Research in the Breaker laboratory is also supported by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
J.X.W. demonstrated the existence of the ligand in B. subtilis extract and K.F. purified the compound. J.W.N. performed MS, in-line probing and reporter assays. N.S. constructed c-di-AMP reporters and genetic knockouts and conducted the transcription termination assays. Z.W. performed bioinformatics. J.W.N., N.S. and R.R.B. designed experiments and interpreted the data. J.W.N., Z.W. and R.R.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–17 and Supplementary Tables 1 and 2. (PDF 2522 kb)
Supplementary Data Set
Locations and gene associations of all known c-di-AMP riboswitches in our database of genomic sequence information (specifically RefSeq and various environmental databases). (PDF 967 kb)
Rights and permissions
About this article
Cite this article
Nelson, J., Sudarsan, N., Furukawa, K. et al. Riboswitches in eubacteria sense the second messenger c-di-AMP. Nat Chem Biol 9, 834–839 (2013). https://doi.org/10.1038/nchembio.1363
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1363
This article is cited by
-
KdpD is a tandem serine histidine kinase that controls K+ pump KdpFABC transcriptionally and post-translationally
Nature Communications (2024)
-
c-di-AMP signaling plays important role in determining antibiotic tolerance phenotypes of Mycobacterium smegmatis
Scientific Reports (2022)
-
Na+ riboswitches regulate genes for diverse physiological processes in bacteria
Nature Chemical Biology (2022)
-
Atypical cyclic di-AMP signaling is essential for Porphyromonas gingivalis growth and regulation of cell envelope homeostasis and virulence
npj Biofilms and Microbiomes (2022)
-
Meta-omics approaches reveal unique small RNAs exhibited by the uncultured microorganisms dwelling deep-sea hydrothermal sediment in Guaymas Basin
Archives of Microbiology (2022)