Abstract
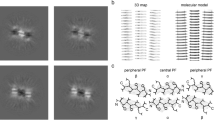
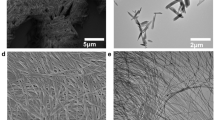
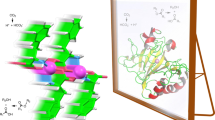
Enzymes fold into unique three-dimensional structures, which underlie their remarkable catalytic properties. The requirement to adopt a stable, folded conformation is likely to contribute to their relatively large size (>10,000 Da). However, much shorter peptides can achieve well-defined conformations through the formation of amyloid fibrils. To test whether short amyloid-forming peptides might in fact be capable of enzyme-like catalysis, we designed a series of seven-residue peptides that act as Zn2+-dependent esterases. Zn2+ helps stabilize the fibril formation, while also acting as a cofactor to catalyse acyl ester hydrolysis. These results indicate that prion-like fibrils are able to not only catalyse their own formation, but they can also catalyse chemical reactions. Thus, they might have served as intermediates in the evolution of modern-day enzymes. These results also have implications for the design of self-assembling nanostructured catalysts including ones containing a variety of biological and non-biological metal ions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Greenwald, J. & Riek, R. On the possible amyloid origin of protein folds. J. Mol. Biol. 421, 417–426 (2012).
Carny, O. & Gazit, E. A model for the role of short self-assembled peptides in the very early stages of the origin of life. FASEB J. 19, 1051–1055 (2005).
DeGrado, W. F. & Lear, J. D. Induction of peptide conformation at apolar/water interfaces: a study with model peptides of defined hydrophobic periodicity. J. Am. Chem. Soc. 107, 7684–7689 (1985).
Eisenberg, D. & Jucker, M. The amyloid state of proteins in human diseases. Cell 148, 1188–1203 (2012).
Colletier, J. P. et al. Molecular basis for amyloid-beta polymorphism. Proc. Natl Acad. Sci. USA 108, 16938–16943 (2011).
Nowick, J. S. Exploring β-sheet structure and interactions with chemical model systems. Acc. Chem. Res. 41, 1319–1330 (2008).
Wachtershauser, G. Before enzymes and templates: theory of surface metabolism. Microbiol. Rev. 52, 452–484 (1988).
Rode, B. M., Son, H. L., Suwannachot, Y. & Bujdak, J. The combination of salt induced peptide formation reaction and clay catalysis: a way to higher peptides under primitive earth conditions. Orig. Life Evol. Biosph. 29, 273–286 (1999).
Christianson, D. W. & Fierke, C. A. Carbonic anhydrase: evolution of a zinc binding site by nature and by design. Acc. Chem. Res. 29, 331–339 (1996).
Iverson, T. M., Alber, B. E., Kisker, C., Ferry, J. G. & Rees, D. C. A closer look at the active site of gamma-class carbonic anhydrases: high-resolution crystallographic studies of the carbonic anhydrase from Methanosarcina thermophila. Biochemistry 39, 9222–9231 (2000).
West, M. W. et al. De novo amyloid proteins from designed combinatorial libraries. Proc. Natl Acad. Sci. USA 96, 11211–11216 (1999).
DeGrado, W. F., Wasserman, Z. R. & Lear, J. D. Protein design, a minimalist approach. Science 243, 622–628 (1989).
Schneider, J. P. et al. Responsive hydrogels from the intramolecular folding and self-assembly of a designed peptide. J. Am. Chem. Soc. 124, 15030–15037 (2002).
Zhang, S., Marini, D. M., Hwang, W. & Santoso, S. Design of nanostructured biological materials through self-assembly of peptides and proteins. Curr. Opin. Chem. Biol. 6, 865–871 (2002).
Minor, D. L. & Kim, P. S. Measurement of the β-sheet-forming propensities of amino acids. Nature 367, 660–663 (1994).
Smith, C. K., Withka, J. M. & Regan, L. A thermodynamic scale for the β-sheet forming tendencies of the amino acids. Biochemistry 33, 5510–5517 (1994).
Krattiger, P., Kovasy, R., Revell, J. D., Ivan, S. & Wennemers, H. Increased structural complexity leads to higher activity: peptides as efficient and versatile catalysts for asymmetric aldol reactions. Org. Lett. 7, 1101–1103 (2005).
Tang, Z. et al. Small peptides catalyze highly enantioselective direct aldol reactions of aldehydes with hydroxyacetone: unprecedented regiocontrol in aqueous media. Org. Lett. 6, 2285–2287 (2004).
Kodaka, M. Hydrolysis of p-nitrophenyl esters by histidine-containing linear and cyclic peptides. Bull. Chem. Soc. Jpn 56, 3857–3858 (1983).
Bolon, D. N. & Mayo, S. Enzyme-like proteins by computational design. Proc. Natl Acad. Sci. USA 98, 14272–14279 (2001).
Wei, Y. & Hecht, M. H. Enzyme-like proteins from an unselected library of designed amino acid sequences. Protein Eng. Des. Sel. 17, 67–75 (2004).
Patel, S. C., Bradley, L. H., Jinadasa, S. P. & Hecht, M. H. Cofactor binding and enzymatic activity in an unevolved superfamily of de novo designed 4-helix bundle proteins. Protein Sci. 18, 1388–1400 (2009).
Broo, K. S., Brive, L., Ahlberg, P. & Baltzer, L. Catalysis of hydrolysis and transesterification reactions of p-nitrophenyl esters by a designed helix-loop-helix dimer. J. Am. Chem. Soc. 119, 11362–11372 (1997).
Zastrow, M. L., Peacock, A. F. A., Stuckey, J. A. & Pecoraro, V. L. Hydrolytic catalysis and structural stabilization in a designed metalloprotein. Nature Chem. 4, 118–123 (2012).
Der, B. S., Edwards, D. R. & Kuhlman, B. Catalysis by a de novo zinc-mediated protein interface: implications for natural enzyme evolution and rational enzyme engineering. Biochemistry 51, 3933–3940 (2012).
Verpoort, J. A., Mehta, S. & Edsall, J. T. Esterase activities of human carbonic anhydrases B and C. J. Biol. Chem. 242, 4221–4229 (1967).
Calero, M. & Gasset, M. Fourier transform infrared and circular dichroism spectroscopies for amyloid studies. Methods Mol. Biol. 299, 129–151 (2005).
Yamaguchi, K., Kamatari, Y. O., Fukuoka, M., Miyaji, R. & Kuwata, K. Nearly reversible conformational change of amyloid fibrils as revealed by pH-jump experiments. Biochemistry 52, 6797–6806 (2013).
Yamaguchi, K., Matsumoto, T. & Kuwata, K. Critical region for amyloid fibril formation of mouse prion protein: unusual amyloidogenic properties of the helix 2 peptide. Biochemistry 47, 13242–13251 (2008).
Coleman, J. E. & Coleman, R. V. Magnetic circular dichroism of Co(II) carbonic anhydrase. J. Biol. Chem. 247, 4718–4728 (1972).
Spevacek, A. R. et al. Zinc drives a tertiary fold in the prion protein with familial disease mutation sites at the interface. Structure 21, 236–246 (2013).
Walter, E. D., Stevens, D. J., Visconte, M. P. & Millhauser, G. L. The prion protein is a combined zinc and copper binding protein: Zn2+ alters the distribution of Cu2+ coordination modes. J. Am. Chem. Soc. 129, 15440–15441 (2007).
Donnelly, P. S., Xiao, Z. & Wedd, A. G. Copper and Alzheimer's disease. Curr. Opin. Chem. Biol. 11, 128–133 (2007).
Chassaing, S. et al. Copper and heme-mediated Aβ toxicity: redox chemistry, Aβ oxidations and anti-ROS compounds. Curr. Top. Med. Chem. 12, 2573–2595 (2012).
Cassagnes, L-E. et al. The catalytically active copper-amyloid-beta state: coordination site responsible for reactive oxygen species production. Angew. Chem. Int. Ed. 52, 11110–11113 (2013).
Korendovych, I. V. et al. Computational design of a self-assembling β-peptide oligomer. Org. Lett. 12, 5142–5145 (2010).
Kuipers, B. J. H. & Gruppen, H. Prediction of molar extinction coefficients of proteins and peptides using UV absorption of the constituent amino acids at 214 nm to enable quantitative reverse phase high-performance liquid chromatography–mass spectrometry analysis. J. Agricult. Food Chem. 55, 5445–5451 (2007).
Acknowledgements
The authors thank K. L. Mack, G. G. Ariotti, R. P. Doyle, M. M. Maye and R. P. Smith for technical assistance and discussions. This work was in part supported by grant no. GM54616 from the National Institutes of Health (to W.F.D.), grant no. 1332349 from the National Science Foundation (NSF) Emerging Frontiers in Research and Innovation (EFRI) program, and an Oak Ridge Associated Universities Ralph E. Powe Junior Faculty Enhancement award to I.V.K. The authors also acknowledge support from the Materials Research Science and Engineering Center programme of the NSF, grant DMR-1120901.
Author information
Authors and Affiliations
Contributions
C.M.R., Y.S.M., O.V.M., J.S., T.A.S. and X.H. performed the experiments and analysed the data. W.F.D. and I.V.K. conceived and designed the experiments and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 7531 kb)
Rights and permissions
About this article
Cite this article
Rufo, C., Moroz, Y., Moroz, O. et al. Short peptides self-assemble to produce catalytic amyloids. Nature Chem 6, 303–309 (2014). https://doi.org/10.1038/nchem.1894
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1894
This article is cited by
-
Enzymatic self-assembly/disassembly turns “ON”/“OFF” the mimetic hydrolytic activity of histidine nanofibers
Science China Chemistry (2024)
-
Cryo-EM structure and polymorphic maturation of a viral transduction enhancing amyloid fibril
Nature Communications (2023)
-
A supramolecular metalloenzyme possessing robust oxidase-mimetic catalytic function
Nature Communications (2023)
-
Highly stable and tunable peptoid/hemin enzymatic mimetics with natural peroxidase-like activities
Nature Communications (2022)
-
Machine learning overcomes human bias in the discovery of self-assembling peptides
Nature Chemistry (2022)