Abstract
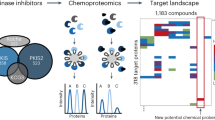
We describe a chemical proteomics approach to profile the interaction of small molecules with hundreds of endogenously expressed protein kinases and purine-binding proteins. This subproteome is captured by immobilized nonselective kinase inhibitors (kinobeads), and the bound proteins are quantified in parallel by mass spectrometry using isobaric tags for relative and absolute quantification (iTRAQ). By measuring the competition with the affinity matrix, we assess the binding of drugs to their targets in cell lysates and in cells. By mapping drug-induced changes in the phosphorylation state of the captured proteome, we also analyze signaling pathways downstream of target kinases. Quantitative profiling of the drugs imatinib (Gleevec), dasatinib (Sprycel) and bosutinib in K562 cells confirms known targets including ABL and SRC family kinases and identifies the receptor tyrosine kinase DDR1 and the oxidoreductase NQO2 as novel targets of imatinib. The data suggest that our approach is a valuable tool for drug discovery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Hall, S.E. Chemoproteomics-driven drug discovery: addressing high attrition rates. Drug Discov. Today 11, 495–502 (2006).
Morphy, R., Kay, C. & Rankovic, Z. From magic bullets to designed multiple ligands. Drug Discov. Today 9, 641–651 (2004).
Cohen, P. Protein kinases–the major drug targets of the twenty-first century? Nat. Rev. Drug Discov. 1, 309–315 (2002).
Liu, Y. & Gray, N.S. Rational design of inhibitors that bind to inactive kinase conformations. Nat. Chem. Biol. 2, 358–364 (2006).
Daub, H., Specht, K. & Ullrich, A. Strategies to overcome resistance to targeted protein kinase inhibitors. Nat. Rev. Drug Discov. 3, 1001–1010 (2004).
Mol, C.D., Fabbro, D. & Hosfield, D.J. Structural insights into the conformational selectivity of STI-571 and related kinase inhibitors. Curr. Opin. Drug Discov. Devel. 7, 639–648 (2004).
Fabbro, D. & Garcia-Echeverria, C. Targeting protein kinases in cancer therapy. Curr. Opin. Drug Discov. Devel. 5, 701–712 (2002).
Manning, G., Whyte, D.B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Haystead, T.A. The purinome, a complex mix of drug and toxicity targets. Curr. Top. Med. Chem. 6, 1117–1127 (2006).
Fabian, M.A. et al. A small molecule-kinase interaction map for clinical kinase inhibitors. Nat. Biotechnol. 23, 329–336 (2005).
Fliri, A.F., Loging, W.T., Thadeio, P.F. & Volkmann, R.A. Analysis of drug-induced effect patterns to link structure and side effects of medicines. Nat. Chem. Biol. 1, 389–397 (2005).
Szardenings, K., Li, B., Ma, L. & Wu, M. Fishing for targets: novel approaches using small molecule baits. Drug Discov. Today: Technologies 3, 9–15 (2004).
Godl, K. et al. An efficient proteomics method to identify the cellular targets of protein kinase inhibitors. Proc. Natl. Acad. Sci. USA 100, 15434–15439 (2003).
Knockaert, M. et al. Intracellular targets of cyclin-dependent kinase inhibitors: identification by affinity chromatography using immobilised inhibitors. Chem. Biol. 7, 411–422 (2000).
Ross, P.L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Hantschel, O. & Superti-Furga, G. Regulation of the c-Abl and Bcr-Abl tyrosine kinases. Nat. Rev. Mol. Cell Biol. 5, 33–44 (2004).
Capdeville, R., Buchdunger, E., Zimmermann, J. & Matter, A. Glivec (STI571, imatinib), a rationally developed, targeted anticancer drug. Nat. Rev. Drug Discov. 1, 493–502 (2002).
Weisberg, E., Manley, P.W., Cowan-Jacob, S.W., Hochhaus, A. & Griffin, J.D. Second generation inhibitors of BCR-ABL for the treatment of imatinib-resistant chronic myeloid leukaemia. Nat. Rev. Cancer 7, 345–356 (2007).
Ding, S. et al. Synthetic small molecules that control stem cell fate. Proc. Natl. Acad. Sci. USA 100, 7632–7637 (2003).
Rappsilber, J., Ryder, U., Lamond, A.I. & Mann, M. Large-scale proteomic analysis of the human spliceosome. Genome Res. 12, 1231–1245 (2002).
Wissing, J. et al. Chemical proteomic analysis reveals alternative modes of action for pyrido[2,3-d]pyrimidine kinase inhibitors. Mol. Cell. Proteomics 3, 1181–1193 (2004).
Graves, P.R. et al. Discovery of novel targets of quinoline drugs in the human purine binding proteome. Mol. Pharmacol. 62, 1364–1372 (2002).
Golas, J.M. et al. SKI-606, a 4-anilino-3-quinolinecarbonitrile dual inhibitor of Src and Abl kinases, is a potent antiproliferative agent against chronic myelogenous leukemia cells in culture and causes regression of K562 xenografts in nude mice. Cancer Res. 63, 375–381 (2003).
Kantarjian, H., Jabbour, E., Grimley, J. & Kirkpatrick, P. Dasatinib. Nat. Rev. Drug Discov. 5, 717–718 (2006).
Druker, B.J. et al. Effects of a selective inhibitor of the Abl tyrosine kinase on the growth of Bcr-Abl positive cells. Nat. Med. 2, 561–566 (1996).
Carter, T.A. et al. Inhibition of drug-resistant mutants of ABL, KIT, and EGF receptor kinases. Proc. Natl. Acad. Sci. USA 102, 11011–11016 (2005).
Kowanetz, K. et al. Suppressors of T-cell receptor signaling Sts-1 and Sts-2 bind to Cbl and inhibit endocytosis of receptor tyrosine kinases. J. Biol. Chem. 279, 32786–32795 (2004).
Tuveson, D.A. et al. STI571 inactivation of the gastrointestinal stromal tumor c-KIT oncoprotein: biological and clinical implications. Oncogene 20, 5054–5058 (2001).
Su, A.I. et al. Large-scale analysis of the human and mouse transcriptomes. Proc. Natl. Acad. Sci. USA 99, 4465–4470 (2002).
L'hote, C.G., Thomas, P.H. & Ganesan, T.S. Functional analysis of discoidin domain receptor 1: effect of adhesion on DDR1 phosphorylation. FASEB J. 16, 234–236 (2002).
Vella, F., Ferry, G., Delagrange, P. & Boutin, J.A. NRH:quinone reductase 2: an enzyme of surprises and mysteries. Biochem. Pharmacol. 71, 1–12 (2005).
Knox, R.J. et al. Bioactivation of 5-(aziridin-1-yl)-2,4-dinitrobenzamide (CB 1954) by human NAD(P)H quinone oxidoreductase 2: a novel co-substrate-mediated antitumor prodrug therapy. Cancer Res. 60, 4179–4186 (2000).
Patricelli, M.P. et al. Functional interrogation of the kinome using nucleotide acyl phosphates. Biochemistry 46, 350–358 (2007).
Knight, Z.A. & Shokat, K.M. Features of selective kinase inhibitors. Chem. Biol. 12, 621–637 (2005).
Buchdunger, E. et al. Inhibition of the Abl protein-tyrosine kinase in vitro and in vivo by a 2-phenylaminopyrimidine derivative. Cancer Res. 56, 100–104 (1996).
Lombardo, L.J. et al. Discovery of N-(2-chloro-6-methyl-phenyl)-2-(6-(4-(2-hydroxyethyl)-piperazin-1-yl)-2-methylpyrimidin-4-ylamino)thiazole-5-carboxamide (BMS-354825), a dual Src/Abl kinase inhibitor with potent antitumor activity in preclinical assays. J. Med. Chem. 47, 6658–6661 (2004).
Lowe, C.R., Harvey, M.J., Craven, D.B. & Dean, P.D. Some parameters relevant to affinity chromatography on immobilized nucleotides. Biochem. J. 133, 499–506 (1973).
Vogel, W.F., Abdulhussein, R. & Ford, C.E. Sensing extracellular matrix: an update on discoidin domain receptor function. Cell. Signal. 18, 1108–1116 (2006).
Ongusaha, P.P. et al. p53 induction and activation of DDR1 kinase counteract p53-mediated apoptosis and influence p53 regulation through a positive feedback loop. EMBO J. 22, 1289–1301 (2003).
Rossler, J., Zambrzycka, I., Lagodny, J., Kontny, U. & Niemeyer, C.M. Effect of STI-571 (imatinib mesylate) in combination with retinoic acid and gamma-irradiation on viability of neuroblastoma cells. Biochem. Biophys. Res. Commun. 342, 1405–1412 (2006).
Avivi-Green, C., Singal, M. & Vogel, W.F. Discoidin domain receptor 1-deficient mice are resistant to bleomycin-induced lung fibrosis. Am. J. Respir. Crit. Care Med. 174, 420–427 (2006).
Daniels, C.E. et al. Imatinib mesylate inhibits the profibrogenic activity of TGF-beta and prevents bleomycin-mediated lung fibrosis. J. Clin. Invest. 114, 1308–1316 (2004).
Kneidinger, M. et al. Dasatinib (BMS354825) inhibits IgE-dependent activation and histamine release in human blood basophils. Blood 108 Abs. 1365 (2006).
Shah, N.P. et al. Dasatinib (BMS-354825) inhibits KITD816V, an imatinib-resistant activating mutation that triggers neoplastic growth in most patients with systemic mastocytosis. Blood 108, 286–291 (2006).
Yang, T.T., Xiong, Q., Graef, I.A., Crabtree, G.R. & Chow, C.W. Recruitment of the extracellular signal-regulated kinase/ribosomal S6 kinase signaling pathway to the NFATc4 transcription activation complex. Mol. Cell. Biol. 25, 907–920 (2005).
Drewes, G. et al. Process for the identification of novel enzyme interacting compounds. Patent WO 2006/134056 A1 (2006).
Boschelli, D.H. et al. 7-Alkoxy-4-phenylamino-3-quinolinecar-bonitriles as dual inhibitors of Src and Abl kinases. J. Med. Chem. 47, 1599–1601 (2004).
Das, J. et al. 2-aminothiazole as a novel kinase inhibitor template. Structure-activity relationship studies toward the discovery of N-(2-chloro-6-methylphenyl)-2-[[6-[4-(2-hydroxyethyl)-1-piperazinyl)]-2-methyl-4-pyrimidinyl]amino)]-1,3-thiazole-5-carboxamide (dasatinib, BMS-354825) as a potent pan-Src kinase inhibitor. J. Med. Chem. 49, 6819–6832 (2006).
Shevchenko, A., Wilm, M., Vorm, O. & Mann, M. Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal. Chem. 68, 850–858 (1996).
Pozuelo, R.M., Campbell, D.G., Morrice, N.A. & Mackintosh, C. Phosphodiesterase 3A binds to 14–3-3 proteins in response to PMA-induced phosphorylation of Ser428. Biochem. J. 392, 163–172 (2005).
Acknowledgements
This work was partially supported by a grant from the German Bundesministerium fuer Bildung und Forschung (BMBF BioChancePLUS grant 0313335A). We would like to thank Charles Cohen, Tim Edwards, David Middlemiss, Markus Schirle, David Simmons and Francis Wilson for helpful discussions and support. We are grateful to Birgit Duempelfeld, Eva-Maria Kashammer-Lorenz, Jana Krause, Anja Podszuweit, Tatjana Rudi and Thilo Werner for expert technical assistance, to Svenja Burckhardt and Cyrille Boussard for the synthesis of compounds, to Michael Rinner, Judith Schlegl and Marianna Stabilini for database and IT tool development, and to Frank Weisbrodt for help with the figures. We would also like to thank Oliver Hantschel and Giulio Superti-Furga for stimulating discussions and valuable comments on the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors are employees of Cellzome AG or Cellzome, UK, Ltd. These companies funded the work.
Supplementary information
Supplementary Text and Figures
Figure 1–5, Tables 1–11, Methods, Data (PDF 2257 kb)
Supplementary Table 10
Peptide identification data collected in the experiments shown in Fig. 1 and Supplementary Fig. 1 (TXT 5210 kb)
Supplementary Table 11
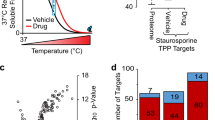
Peptide identification data collected in the quantitative experiments shown in Fig. 3b and Fig. 3c. (TXT 16370 kb)
Rights and permissions
About this article
Cite this article
Bantscheff, M., Eberhard, D., Abraham, Y. et al. Quantitative chemical proteomics reveals mechanisms of action of clinical ABL kinase inhibitors. Nat Biotechnol 25, 1035–1044 (2007). https://doi.org/10.1038/nbt1328
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1328
This article is cited by
-
Kinase inhibitor pulldown assay (KiP) for clinical proteomics
Clinical Proteomics (2024)
-
Development of a novel AAK1 inhibitor via Kinobeads-based screening
Scientific Reports (2024)
-
Profiling the interactome of oligonucleotide drugs by proximity biotinylation
Nature Chemical Biology (2024)
-
Dihydroartemisinin suppresses renal fibrosis in mice by inhibiting DNA-methyltransferase 1 and increasing Klotho
Acta Pharmacologica Sinica (2022)
-
The emerging role of mass spectrometry-based proteomics in drug discovery
Nature Reviews Drug Discovery (2022)