Abstract
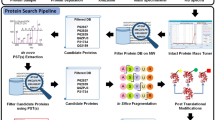
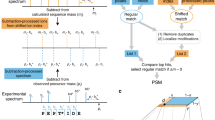
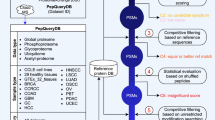
Most tandem mass spectrometry (MS/MS) database search algorithms perform a restrictive search that takes into account only a few types of post-translational modifications (PTMs) and ignores all others. We describe an unrestrictive PTM search algorithm, MS-Alignment, that searches for all types of PTMs at once in a blind mode, that is, without knowing which PTMs exist in nature. Blind PTM identification makes it possible to study the extent and frequency of different types of PTMs, still an open problem in proteomics. Application of this approach to lens proteins resulted in the largest set of PTMs reported in human crystallins so far. Our analysis of various MS/MS data sets implies that the biological phenomenon of modification is much more widespread than previously thought. We also argue that MS-Alignment reveals some uncharacterized modifications that warrant further experimental validation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Shu, H., Chen, S., Bi, Q., Mumby, M. & Brekken, D.L. Identification of phosphoproteins and their phosphorylation sites in the wehi-231 b lymphoma cell line. Mol. Cell. Proteomics 3, 279–286 (2004).
Cantin, G.T. & Yates, J.R. Strategies for shotgun identification of post-translational modifications by mass spectrometry. J. Chromatogr. A. 1053, 7–14 (2004).
Yates, J.R., Eng, J.K. & McCormack, A.L. Mining genomes: correlating tandem mass spectra of modified and unmodified peptides to sequences in nucleotide databases. Anal. Chem. 67, 3202–3210 (1995).
Pevzner, P.A., Dančík, V. & Tang, C.L. Mutation-tolerant protein identification by mass spectrometry. J. Comput. Biol. 7, 777–787 (2000).
Pevzner, P.A., Mulyukov, Z., Dancik, V. & Tang, C.L. Efficiency of database search for identification of mutated and modified proteins via mass spectrometry. Genome Res. 11, 290–299 (2001).
Searle, B.C. et al. High-throughput identification of proteins and unanticipated sequence modifications using a mass-based alignment algorithm for MS/MS de novo sequencing results. Anal. Chem. 76, 2220–2230 (2004).
Han, Y., Ma, B. & Zhang, K. SPIDER: software for protein identification from sequence tags with de novo sequencing error. J. Bioinform. Comput. Biol. 3, 697–716 (2005).
Hansen, B.T., Davey, S.W., Ham, A.J. & Liebler, D.C. P-mod: an algorithm and software to map modifications to peptide sequences using tandem MS data. J. Proteome Res. 4, 358–368 (2005).
Tang, W.H. et al. Discovering known and unanticipated protein modifications using MS/MS database searching. Anal. Chem. 77, 3931–3946 (2005).
Searle, B.S. et al. Identification of protein modifications using MS/MS de novo sequencing and the Opensea alignment algorithm. J. Proteome Res. 4, 546–554 (2005).
MacCoss, M.J., Wu, C.C. & Yates, J.R. Probability-based validation of protein identifications using a modified SEQUEST algorithm. Anal. Chem. 74, 5593–5599 (2002).
Keller, A. et al. Experimental protein mixture for validating tandem mass spectral analysis. OMICS 6, 207–212 (2002).
Tanner, S. et al. Inspect: fast and accurate identification of post-translationally modified peptides from tandem mass spectra. Anal. Chem. 77, 4626–4639 (2005).
Craig, R. & Beavis, R.C. A method for reducing the time required to match protein sequences with tandem mass spectra. Rapid Commun. Mass Spectrom. 17, 2310–2316 (2003).
Yates, J.R., Eng, J.K., McCormack, A.L. & Schieltz, D. Method to correlate tandem mass spectra of modified peptides to amino acid sequences in the protein database. Anal. Chem. 67, 1426–1436 (1995).
Tabb, D.L. et al. Statistical characterization of ion trap tandem mass spectra from doubly charged tryptic peptides. Anal. Chem. 75, 1155–1163 (2003).
Perkins, D.N., Pappin, D.J., Creasy, D.M. & Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Nesvizhskii, A.I., Keller, A., Kolker, E. & Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 75, 4646–4658 (2003).
Razumovskaya, J. et al. A computational method for assessing peptide-identification reliability in tandem mass spectrometry analysis with sequest. Proteomics 4, 961–969 (2004).
Frank, A., Tanner, S.W., Bafna, V. & Pevzner, P.A. Peptide sequence tags for fast database search in mass-spectrometry. J. Proteome Res. 4, 1287–1295 (2005).
Elias, J.E., Gibbons, F.D., King, O.D., Roth, F.P. & Gygi, S.P. Intensity-based protein identification by protein learning from a library of tandem mass spectra. Nat. Biotechnol. 22, 214–219 (2004).
Havilio, M., Haddad, Y. & Smilansky, Z. Intensity-based statistical scorer for tandem mass spectrometry. Anal. Chem. 75, 435–444 (2003).
Anderson, D.C., Li, W., Payan, D.G. & W.S., Noble A new algorithm for the evaluation of shotgun peptide sequencing in proteomics: support vector machine classification of peptide MS/MS spectra and SEQUEST scores. J. Proteome Res. 2, 137–146 (2003).
Geer, L.Y. et al. Open mass spectrometry search algorithm. J. Proteome Res. 3, 958–964 (2004).
Acknowledgements
This project was supported by National Institutes of Health grant NIGMS 1-R01-RR16522. We are grateful to Brian Searle and Larry David for making their lens data set available and to Larry David, Katalin Medzihradszky and Philip Wilmarth for many useful discussions. Production of the lens data set was supported by National Eye Institute grant EY007755. This research was supported in part by the UCSD FWGrid Project, NSF Research Infrastructure Grant Number EIA-0303622. Production of the IKKb data set was supported by NIH grant R01GM65325 and by the Pew Scholars Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Spectra from the ISB spectra were searched against a database containing valid proteins (37) and human nr (90,000). (PDF 32 kb)
Supplementary Fig. 2
Receiver Operating Characteristic (ROC) curve for the SVM score on the ISB data set. (PDF 117 kb)
Supplementary Table 1
Summary of validated modification sites over Lens proteins, compared with results reported by OpenSea (Searle et al, 2005). (PDF 7 kb)
Supplementary Table 2
PTM site count matrix for IKKb dataset (1,072 sites total). (PDF 11 kb)
Supplementary Table 3
Modifications on Lens data-set. (PDF 47 kb)
Rights and permissions
About this article
Cite this article
Tsur, D., Tanner, S., Zandi, E. et al. Identification of post-translational modifications by blind search of mass spectra. Nat Biotechnol 23, 1562–1567 (2005). https://doi.org/10.1038/nbt1168
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1168
This article is cited by
-
Proteome-Wide Analyses Reveal the Diverse Functions of Lysine 2-Hydroxyisobutyrylation in Oryza sativa
Rice (2020)
-
Identification of modified peptides using localization-aware open search
Nature Communications (2020)
-
Increased diversity of peptidic natural products revealed by modification-tolerant database search of mass spectra
Nature Microbiology (2018)
-
Informed-Proteomics: open-source software package for top-down proteomics
Nature Methods (2017)
-
Metabolic regulation of gene expression through histone acylations
Nature Reviews Molecular Cell Biology (2017)