Abstract
High-throughput genetic screens have become essential tools for studying a wide variety of biological processes. Here we experimentally compare systems based on clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated protein 9 (Cas9) or its transcriptionally repressive variant, CRISPR-interference (CRISPRi), with a traditional short hairpin RNA (shRNA)-based system for performing lethality screens. We find that the CRISPR technology performed best, with low noise, minimal off-target effects and consistent activity across reagents.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Bernards, R. Curr. Opin. Genet. Dev. 24, 23–29 (2014).
Amberkar, S., Kiani, N.A., Bartenschlager, R., Alvisi, G. & Kaderali, L. World J. Virol. 2, 18–31 (2013).
Cong, L. et al. Science 339, 819–823 (2013).
Mali, P. et al. Science 339, 823–826 (2013).
Shalem, O. et al. Science 343, 84–87 (2014).
Wang, T., Wei, J.J., Sabatini, D.M. & Lander, E.S. Science 343, 80–84 (2014).
Gilbert, L.A. et al. Cell 154, 442–451 (2013).
Gilbert, L.A. et al. Cell 159, 647–661 (2014).
Fu, Y. et al. Nat. Biotechnol. 31, 822–826 (2013).
Kim, D. et al. Nat. Methods 12, 237–243, 1, 243 (2015).
Hart, T., Brown, K.R., Sircoulomb, F., Rottapel, R. & Moffat, J. Mol. Syst. Biol. 10, 733 (2014).
Li, W. et al. Genome Biol. 15, 554 (2014).
Forbes, S.A. et al. Nucleic Acids Res. 43, D805–D811 (2015).
Sanjana, N.E., Shalem, O. & Zhang, F. Nat. Methods 11, 783–784 (2014).
Chen, B. et al. Cell 155, 1479–1491 (2013).
Rodriguez, J.M. et al. Nucleic Acids Res. 41, D110–D117 (2013).
Acknowledgements
We thank all members of the Bernards and Beijersbergen groups and G. Bounova for useful discussions, and the Netherlands Cancer Institute genomics core facility for next-generation sequencing. The doxycycline-inducible KRAB-dCas9 expression system, consisting of pHR-TRE3G-KRAB-dCas9-P2A-mCherry and pHR-Tet3G, were kind gifts of Luke Gilbert, the Jonathan Weissman laboratories, UCSF. We thank G. Verhaegh, Nijmegen Institute for Molecular Sciences, the Netherlands, and M. Knowles, Leeds Institute of Cancer Studies and Pathology, UK for the cell lines they kindly provided.
Author information
Authors and Affiliations
Contributions
Experiments were designed by B.E., K.J., R.L.B. and R.B., carried out by B.E., K.J., J.P.M.H. and W.G., and analyzed by B.E. The paper was written by B.E., R.L.B. and R.B. This work was supported by the Cancer Genomics Netherlands consortium.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
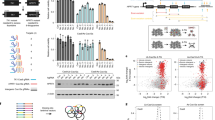
Supplementary Figure 1 Screening technology overview
The shRNA screens were performed in the pLKO.1 puro backbone (Sigma-Aldrich), the CRISPR gRNAs were cloned in a modified version of Lenti-CRISPRv2.0 (see Online Materials and Methods). The CRISPRi system consisted of a Doxycycline inducible KRAB-dCas9 fusion construct co-expressing mCherry, a constitutive tet activator and a plasmid encoding the gRNAs driven from a constitutive U6 promoter.
Supplementary Figure 5 Fraction of functional constructs per gene
Ratios of functional gRNAs to all gRNAs targeting every gene were determined for each screening technology.
Supplementary Figure 6 Comparison of CRISPRi screen with previously published data
Performance of gRNAs that were identical in the libraries used in this publication and the previously published paper by Gilbert et al.7 was plotted as -10Log P-values compared to the performance of the other gRNAs present in the library (A). The growth phenotype upon transduction in K562 cells as reported in this paper was compared to 2Log fold depletion of corresponding gRNAs used in this publication (B).
Supplementary Figure 7 gRNA positions across transcript variants for the CRISPRi library
For every gene, transcripts were plotted from -50 to +300 around the TSS (vertical line) and aligned where possible. gRNAs were plotted yellow when q-values were >0.1 and blue when ≤0.1. For RPL36, two transcripts that were wrongly mapped in hg19 are plotted in gray.
Supplementary Figure 8 Screen results of UM-UC-3 cells
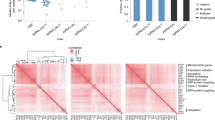
Relative fold-change of averaged normalized read-counts of t=1 vs. t=0 for the shRNA (A) and CRISPR (B) technologies plotted as a function of their initial averaged normalized read-counts at t=0. Orange dots represent constructs targeting essential genes, blue dots represent constructs targeting non-essential genes. Data was averaged over three biological replicate screens.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 3140 kb)
Supplementary Table 1
Data of shRNA screen (XLSX 123 kb)
Supplementary Table 2
Data of CRISPR screen (XLSX 240 kb)
Supplementary Table 3
Data of CRISPRi screen (XLSX 119 kb)
Supplementary Table 4
Primers (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Evers, B., Jastrzebski, K., Heijmans, J. et al. CRISPR knockout screening outperforms shRNA and CRISPRi in identifying essential genes. Nat Biotechnol 34, 631–633 (2016). https://doi.org/10.1038/nbt.3536
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3536
This article is cited by
-
Cas13b-mediated RNA targeted therapy alleviates genetic dilated cardiomyopathy in mice
Cell & Bioscience (2024)
-
To metabolomics and beyond: a technological portfolio to investigate cancer metabolism
Signal Transduction and Targeted Therapy (2023)
-
Dissecting metastasis using preclinical models and methods
Nature Reviews Cancer (2023)
-
The APC/C E3 ligase subunit ANAPC11 mediates FOXO3 protein degradation to promote cell proliferation and lymph node metastasis in urothelial bladder cancer
Cell Death & Disease (2023)
-
Construction of a Set of Novel Transposon Vectors for Efficient Silencing of Protein and lncRNA Genes via CRISPR Interference
Molecular Biotechnology (2023)