Abstract
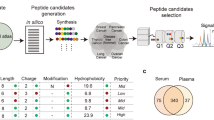
High-throughput technologies can now identify hundreds of candidate protein biomarkers for any disease with relative ease. However, because there are no assays for the majority of proteins and de novo immunoassay development is prohibitively expensive, few candidate biomarkers are tested in clinical studies. We tested whether the analytical performance of a biomarker identification pipeline based on targeted mass spectrometry would be sufficient for data-dependent prioritization of candidate biomarkers, de novo development of assays and multiplexed biomarker verification. We used a data-dependent triage process to prioritize a subset of putative plasma biomarkers from >1,000 candidates previously identified using a mouse model of breast cancer. Eighty-eight novel quantitative assays based on selected reaction monitoring mass spectrometry were developed, multiplexed and evaluated in 80 plasma samples. Thirty-six proteins were verified as being elevated in the plasma of tumor-bearing animals. The analytical performance of this pipeline suggests that it should support the use of an analogous approach with human samples.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Anderson, N.L. The clinical plasma proteome: a survey of clinical assays for proteins in plasma and serum. Clin. Chem. 56, 177–185 (2010).
Anderson, N.L. et al. A human proteome detection and quantitation project. Mol. Cell. Proteomics 8, 883–886 (2009).
Bordeaux, J. et al. Antibody validation. Biotechniques 48, 197–209 (2010).
Haab, B.B. et al. A reagent resource to identify proteins and peptides of interest for the cancer community: a workshop report. Mol. Cell. Proteomics 5, 1996–2007 (2006).
Stoevesandt, O. & Taussig, M.J. Affinity reagent resources for human proteome detection: initiatives and perspectives. Proteomics 7, 2738–2750 (2007).
Jaffe, J.D. et al. Accurate inclusion mass screening: a bridge from unbiased discovery to targeted assay development for biomarker verification. Mol. Cell. Proteomics 7, 1952–1962 (2008).
Kuhn, E. et al. Quantification of C-reactive protein in the serum of patients with rheumatoid arthritis using multiple reaction monitoring mass spectrometry and 13C-labeled peptide standards. Proteomics 4, 1175–1186 (2004).
Gerber, S.A., Rush, J., Stemman, O., Kirschner, M.W. & Gygi, S.P. Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc. Natl. Acad. Sci. USA 100, 6940–6945 (2003).
Barr, J. et al. Isotope-dilution mass spectrometric quantification of specific proteins: model application with apolipoprotein A-1. Clin. Chem. 42, 1676–1682 (1996).
Wu, S. et al. Targeted proteomics of low-level proteins in human plasma by LC/MSn: using human growth hormone as a model system. J. Proteome Res. 1, 459–465 (2002).
Barnidge, D., Goodmanson, M., Klee, G. & Muddiman, D. Absolute quantification of the model biomarker prostate-specific antigen in serum by LC-MS/MS using protein cleavage and isotope dilution MS. J. Proteome Res. 3, 644–652 (2004).
Wang, P., Whiteaker, J.R. & Paulovich, A.G. The evolving role of mass spectrometry in cancer biomarker discovery. Cancer Biol. Ther. 8, 1083–1094 (2009).
Rifai, N., Gillette, M.A. & Carr, S.A. Protein biomarker discovery and validation: the long and uncertain path to clinical utility. Nat. Biotechnol. 24, 971–983 (2006).
Anderson, N.L. The roles of multiple proteomic platforms in a pipeline for new diagnostics. Mol. Cell. Proteomics 4, 1441–1444 (2005).
Makawita, S. & Diamandis, E.P. The bottleneck in the cancer biomarker pipeline and protein quantification through mass spectrometry-based approaches: current strategies for candidate verification. Clin. Chem. 56, 212–222 (2010).
Lange, V., Picotti, P., Domon, B. & Aebersold, R. Selected reaction monitoring for quantitative proteomics: a tutorial. Mol. Syst. Biol. 4, 222 (2008).
Pan, S. et al. Mass spectrometry based targeted protein quantification: methods and applications. J. Proteome Res. 8, 787–797 (2009).
Carr, S.A. & Anderson, L. Protein quantitation through targeted mass spectrometry: the way out of biomarker purgatory? Clin. Chem. 54, 1749–1752 (2008).
Boja, E. et al. Restructuring proteomics through verification. Biomark Med 4, 799–803 (2010).
Paulovich, A.G., Whiteaker, J.R., Hoofnagle, A.N. & Wang, P. The interface between biomarker discovery and clinical validation: the tar pit of the protein biomarker pipeline. Proteomics Clin. Appl. 2, 1386–1402 (2008).
Whiteaker, J.R. et al. Integrated pipeline for mass spectrometry-based discovery and confirmation of biomarkers demonstrated in a mouse model of breast cancer. J. Proteome Res. 6, 3962–3975 (2007).
Schoenherr, R.M. et al. Proteome and transcriptome profiles of a Her2/Neu-driven mouse model of breast cancer. Proteomics Clin. Appl. 5, 179–188 (2011).
Moody, S.E. et al. Conditional activation of Neu in the mammary epithelium of transgenic mice results in reversible pulmonary metastasis. Cancer Cell 2, 451–461 (2002).
Omenn, G.S., Yocum, A.K. & Menon, R. Alternative splice variants, a new class of protein cancer biomarker candidates: findings in pancreatic cancer and breast cancer with systems biology implications. Dis. Markers 28, 241–251 (2010).
Zhang, H. et al. Methods for peptide and protein quantitation by liquid chromatography-multiple reaction monitoring mass spectrometry. Mol. Cell. Proteomics 10, M110.006593 (2011).
Anderson, N.L. et al. Mass spectrometric quantitation of peptides and proteins using Stable Isotope Standards and Capture by Anti-Peptide Antibodies (SISCAPA). J. Proteome Res. 3, 235–244 (2004).
Hoofnagle, A.N., Becker, J.O., Wener, M.H. & Heinecke, J.W. Quantification of thyroglobulin, a low-abundance serum protein, by immunoaffinity peptide enrichment and tandem mass spectrometry. Clin. Chem. 54, 1796–1804 (2008).
Kuhn, E. et al. Developing multiplexed assays for troponin I and interleukin-33 in plasma by peptide immunoaffinity enrichment and targeted mass spectrometry. Clin. Chem. 55, 1108–1117 (2009).
Whiteaker, J.R., Zhao, L., Anderson, L. & Paulovich, A.G. An automated and multiplexed method for high throughput peptide immunoaffinity enrichment and multiple reaction monitoring mass spectrometry-based quantification of protein biomarkers. Mol. Cell. Proteomics 9, 184–196 (2010).
Want, E.J., Cravatt, B.F. & Siuzdak, G. The expanding role of mass spectrometry in metabolite profiling and characterization. ChemBioChem 6, 1941–1951 (2005).
Picotti, P. et al. High-throughput generation of selected reaction-monitoring assays for proteins and proteomes. Nat. Methods 7, 43–46 (2010).
Kiyonami, R. et al. Increased selectivity, analytical precision, and throughput in targeted proteomics. Mol. Cell. Proteomics 10, M110.002931 (2011).
Whiteaker, J.R. et al. Evaluation of large scale quantitative proteomic assay development using peptide affinity-based mass spectrometry. Mol. Cell. Proteomics 10, M110.005645 (2011).
Fraser, C.G. Biological Variation: from Principles to Practice (AACC Press, Washington, DC; 2001).
Hoofnagle, A.N. & Wener, M.H. The fundamental flaws of immunoassays and potential solutions using tandem mass spectrometry. J. Immunol. Methods 347, 3–11 (2009).
Keshishian, H. et al. Quantification of cardiovascular biomarkers in patient plasma by targeted mass spectrometry and stable isotope dilution. Mol. Cell. Proteomics 8, 2339–2349 (2009).
Stahl-Zeng, J. et al. High sensitivity detection of plasma proteins by multiple reaction monitoring of N-glycosites. Mol. Cell. Proteomics 6, 1809–1817 (2007).
Keshishian, H., Addona, T., Burgess, M., Kuhn, E. & Carr, S.A. Quantitative, multiplexed assays for low abundance proteins in plasma by targeted mass spectrometry and stable isotope dilution. Mol. Cell. Proteomics 6, 2212–2229 (2007).
Anderson, L. & Hunter, C.L. Quantitative mass spectrometric multiple reaction monitoring assays for major plasma proteins. Mol. Cell. Proteomics 5, 573–588 (2006).
Ekins, R. & Edwards, P. On the meaning of “sensitivity”. Clin. Chem. 43, 1824–1831 (1997).
Ekins, R. & Edwards, P. On the meaning of “sensitivity”: a rejoinder. Clin. Chem. 44, 1773–1778 (1998).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Schoenherr, R.M. et al. Automated screening of monoclonal antibodies for SISCAPA assays using a magnetic bead processor and liquid chromatography-selected reaction monitoring-mass spectrometry. J. Immunol. Methods 353, 49–61 (2010).
Acknowledgements
This work was funded by a grant from The Paul G. Allen Family Foundation and the National Institutes of Health grant U24 CA126476 from the National Cancer Institute Clinical Proteomic Technology Assessment Center (CPTAC). We thank our Biomarker Project advisory board members for advice and input: R. Aebersold, L. Anderson, E. Diamandis, R. Smith, F. Appelbaum, D. Gottschling, M. Groudine, J. Roberts, V. Vasioukhin and N. Urban. We thank all members of the Biomarker Project team for input: L. Hartwell, S.M. Hanash, S.J. Pitteri, H. Wong, K.E. Gurley, D. Liggitt, D.B. Martin, T. Whitmore, A. Peterson, R. Prueitt, M. Fitzgibbon, J.K. Eng, D. May, T. Holzman, Y. Zhang, A. Stimmel, S.L. Zriny, R. Dumpit, I. Coleman and T.D. Lorentzen.
Author information
Authors and Affiliations
Contributions
J.R.W. oversaw all proteomic experiments, guided the data analysis and assisted in writing the manuscript. C.L. performed or oversaw all data analysis and assisted in writing the manuscript. J.K. performed all verification studies using the 57-plex SRM assay. L.H. performed all SQ-SRM studies, assisted with the AIMS studies and assisted with configuration of the multiplex SRM assays. M.T. performed plasma sample processing throughout the project. I.S. led all verification studies using the 31-plex immuno-SRM assay and performed all ELISA analyses. P.Y. performed analysis of all response curve SRM data. R.M.S. assisted with the 31-plex immuno-SRM assay characterization and verification studies, identification of candidate biomarkers, depletion of plasma samples for verification studies and editing of the manuscript. L.Z. assisted with the 31-plex immuno-SRM assay characterization and verification studies. U.J.V. performed all western blot analyses. L.A.C. provided the Her2 mouse model. K.S.K.-S. and C.J.K. helped design the mouse experiments, performed all mouse husbandry and provided the required plasma samples. A.K. assisted in selection of candidate biomarkers, assembly of the inclusion lists for the AIMS analyses and analysis of the AIMS data. P.R.G., J.M.H. and L.A.J. performed the AIMS analyses. P.S.N. managed the overall consortium, contributed data for candidate selection, made intellectual contributions to the design of the study and assisted with editing the manuscript. P.W., M.W.M. and L.A. provided statistical input for the design of the experiments and the analysis of the data and assisted with editing the manuscript. A.G.P. designed experiments, coordinated execution of the project, coordinated analysis/interpretation of the data and assumed primary responsibility for writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results Sections 1–6, Supplementary Appendices A–C and Supplementary Methods (PDF 7024 kb)
Supplementary Data 1
Skyline data file name: “mouseQSRM_targets.sky” (XML 122 kb)
Supplementary Data 2
Skyline data file name: “mouseImmunoSRM_targets.sky” (XML 66 kb)
Rights and permissions
About this article
Cite this article
Whiteaker, J., Lin, C., Kennedy, J. et al. A targeted proteomics–based pipeline for verification of biomarkers in plasma. Nat Biotechnol 29, 625–634 (2011). https://doi.org/10.1038/nbt.1900
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1900
This article is cited by
-
A combination of molecular and clinical parameters provides a new strategy for high-grade serous ovarian cancer patient management
Journal of Translational Medicine (2022)
-
Plasma protein biomarker model for screening Alzheimer disease using multiple reaction monitoring-mass spectrometry
Scientific Reports (2022)
-
Light contamination in stable isotope-labelled internal peptide standards is frequent and a potential source of false discovery and quantitation error in proteomics
Analytical and Bioanalytical Chemistry (2022)
-
Clinical potential of mass spectrometry-based proteogenomics
Nature Reviews Clinical Oncology (2019)
-
Method Validation by CPTAC Guidelines for Multi-protein Marker Assays Using Multiple Reaction Monitoring-mass Spectrometry
Biotechnology and Bioprocess Engineering (2019)