Abstract
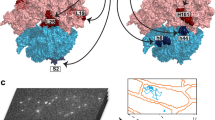
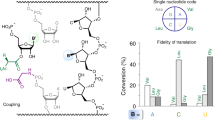
Translation by the ribosome occurs by a complex mechanism involving the coordinated interaction of multiple nucleic acid and protein ligands. Here we use zero-mode waveguides (ZMWs) and sophisticated detection instrumentation to allow real-time observation of translation at physiologically relevant micromolar ligand concentrations. Translation at each codon is monitored by stable binding of transfer RNAs (tRNAs)—labelled with distinct fluorophores—to translating ribosomes, which allows direct detection of the identity of tRNA molecules bound to the ribosome and therefore the underlying messenger RNA (mRNA) sequence. We observe the transit of tRNAs on single translating ribosomes and determine the number of tRNA molecules simultaneously bound to the ribosome, at each codon of an mRNA molecule. Our results show that ribosomes are only briefly occupied by two tRNA molecules and that release of deacylated tRNA from the exit (E) site is uncoupled from binding of aminoacyl-tRNA site (A-site) tRNA and occurs rapidly after translocation. The methods outlined here have broad application to the study of mRNA sequences, and the mechanism and regulation of translation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Green, R. & Noller, H. F. Ribosomes and translation. Annu. Rev. Biochem. 66, 679–716 (1997)
Moazed, D. & Noller, H. F. Intermediate states in the movement of transfer RNA in the ribosome. Nature 342, 142–148 (1989)
Hausner, T. P., Geigenmüller, U. & Nierhaus, K. H. The allosteric three-site model for the ribosomal elongation cycle. New insights into the inhibition mechanism of aminoglycosides, thiostrepton, and viomycin. J. Biol. Chem. 263, 13103–13111 (1988)
Rodnina, M. W. & Wintermeyer, W. Fidelity of aminoacyl-tRNA selection on the ribosome: kinetic and structural mechanisms. Annu. Rev. Biochem. 70, 415–435 (2001)
Agirrezabala, X. et al. Visualization of the hybrid state of tRNA binding promoted by spontaneous racheting of the ribosome. Mol. Cell 32, 190–197 (2008)
Marshall, R. A., Aitken, C. E., Dorywalska, M. & Puglisi, J. D. Translation at the single-molecule level. Annu. Rev. Biochem. 77, 177–203 (2008)
Levene, M. J. et al. Zero-mode waveguides for single-molecule analysis at high concentrations. Science 299, 682–686 (2003)
Foquet, M. et al. Improved fabrication of zero-mode waveguides for single-molecule detection. J. Appl. Phys. 103, 034301 (2008)
Korlach, J. et al. Selective aluminum passivation for targeted immobilization of single DNA polymerase molecules in zero-mode waveguide nanostructures. Proc. Natl Acad. Sci. USA 105, 1176–1181 (2008)
Lundquist, P. M. et al. Parallel confocal detection of single molecules in real time. Opt. Lett. 33, 1026–1028 (2008)
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009)
Blanchard, S. C., Gonzalez, R. L., Kim, H. D., Chu, S. & Puglisi, J. D. tRNA selection and kinetic proofreading in translation. Nature Struct. Mol. Biol. 11, 1008–1014 (2004)
Blanchard, S. C., Kim, H. D., Gonzalez, R. L., Puglisi, J. D. & Chu, S. tRNA dynamics on the ribosome during translation. Proc. Natl Acad. Sci. USA 101, 12893–12898 (2004)
Chan, V., Graves, D. J. & McKenzie, S. E. The biophysics of DNA hybridization with immobilized oligonucleotide probes. Biophys. J. 69, 2243–2255 (1995)
Schlunzen, F. et al. Structural basis for the interaction of antibiotics with the peptidyl transferase centre in eubacteria. Nature 413, 814–821 (2001)
Tenson, T., Lovmar, M. & Ehrenberg, M. The mechanism of action of macrolides, lincosamides and streptogramin B reveals the nascent peptide exit path in the ribosome. J. Mol. Biol. 330, 1005–1014 (2003)
Bodley, J. W. & Godtfredsen, W. O. Studies on translocation. XI. Structure-function relationships of the fusidane-type antibiotics. Biochem. Biophys. Res. Commun. 46, 871–877 (1972)
Underwood, K. A., Swartz, J. R. & Puglisi, J. D. Quantitative polysome analysis identifies limitations in bacterial cell-free protein synthesis. Biotechnol. Bioeng. 91, 425–435 (2005)
Marshall, R. A., Aitken, C. E. & Puglisi, J. D. GTP hydrolysis by IF2 guides progression of the ribosome into elongation. Mol. Cell 35, 37–47 (2009)
Gao, Y. G. et al. The structure of the ribosome with elongation factor G trapped in the posttranslocational state. Science 326, 694–699 (2009)
Lill, R. & Wintermeyer, W. Destabilization of codon-anticodon interaction in the ribosomal exit site. J. Mol. Biol. 196, 137–148 (1987)
Semenkov, Y. P., Rodnina, M. V. & Wintermeyer, W. The “allosteric three-site model” of elongation cannot be confirmed in a well-defined ribosome system from Escherichia coli . Proc. Natl Acad. Sci. USA 93, 12183–12188 (1996)
Cornish, P. V. et al. Following movement of the L1 stalk between three functional states in single ribosomes. Proc. Natl Acad. Sci. USA 106, 2571–2576 (2009)
Fei, J., Kosuri, P., MacDougall, D. D. & Gonzalez, R. L. Coupling of ribosomal L1 stalk and tRNA dynamics during translation elongation. Mol. Cell 30, 348–359 (2008)
Fei, J. et al. Allosteric collaboration between elongation factor G and the ribosomal L1 stalk directs tRNA movements during translation. Proc. Natl Acad. Sci. USA 106, 15702–15707 (2009)
Sanders, C. L. & Curran, J. F. Genetic analysis of the E site during RF2 programmed frameshifting. RNA 13, 1483–1491 (2007)
Aitken, C. E., Marshall, R. A. & Puglisi, J. D. An oxygen scavenging system for improvement of dye stability in single-molecule fluorescence experiments. Biophys. J. 94, 1826–1835 (2008)
Aitken, C. E. & Puglisi, J. D. Following the intersubunit conformation of the ribosome during translation in real time. Nat. Struct. Mol. Biol. (in the press)
Acknowledgements
This work was supported by National Institutes of Health grant GM51266 (to J.D.P.). We thank T. Funatsu, A. Tsai and A. Petrov for encouragement and discussions. We thank J. Gray for performing ellipsometry experiments.
Author Contributions S.U. performed all experiments and data analysis. S.U., C.E.A. and J.D.P. discussed results and wrote the manuscript. S.W.T., J.K. and B.A.F. provided technical expertise with instrumentation and data processing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
S.W.T., B.A.F. and J.K. are employees and stock option holders, and J.D.P. a consultant, of Pacific Biosciences, a company commercializing sequencing technologies.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-7 with legends and Supplementary References. (PDF 3336 kb)
Rights and permissions
About this article
Cite this article
Uemura, S., Aitken, C., Korlach, J. et al. Real-time tRNA transit on single translating ribosomes at codon resolution. Nature 464, 1012–1017 (2010). https://doi.org/10.1038/nature08925
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08925
This article is cited by
-
Visualizing translation dynamics at atomic detail inside a bacterial cell
Nature (2022)
-
Third-generation sequencing and metabolome analysis reveal candidate genes and metabolites with altered levels in albino jackfruit seedlings
BMC Genomics (2021)
-
cAMP binding to closed pacemaker ion channels is non-cooperative
Nature (2021)
-
Development of a sequencing system for spatial decoding of DNA barcode molecules at single-molecule resolution
Communications Biology (2020)
-
Algorithms for ribosome traffic engineering and their potential in improving host cells' titer and growth rate
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.