Abstract
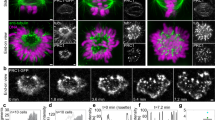
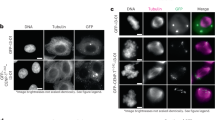
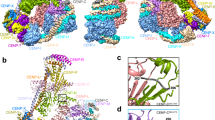
During cell division microtubules capture chromosomes by binding to the kinetochore assembled in the centromeric region of chromosomes. In mitosis sister chromatids are captured by microtubules emanating from both spindle poles, a process called bipolar attachment, whereas in meiosis I sisters are attached to microtubules originating from one spindle pole, called monopolar attachment. For determining chromosome orientation, kinetochore geometry or structure might be an important target of regulation. However, the molecular basis of this regulation has remained elusive. Here we show the link between kinetochore orientation and cohesion within the centromere in fission yeast Schizosaccharomyces pombe by strategies developed to visualize the concealed cohesion within the centromere, and to introduce artificial tethers that can influence kinetochore geometry. Our data imply that cohesion at the core centromere induces the mono-orientation of kinetochores whereas cohesion at the peri-centromeric region promotes bi-orientation. Our study may reveal a general mechanism for the geometric regulation of kinetochores, which collaborates with previously defined tension-dependent reorientation machinery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Nicklas, R. B. How cells get the right chromosomes. Science 275, 632–637 (1997)
Tanaka, T. U. Bi-orienting chromosomes on the mitotic spindle. Curr. Opin. Cell Biol. 14, 365–371 (2002)
Moore, D. P. & Orr-Weaver, T. L. Chromosome segregation during meiosis: building an unambivalent bivalent. Curr. Top. Dev. Biol. 37, 263–299 (1998)
Petronczki, M., Siomos, M. F. & Nasmyth, K. Un menage a quatre: the molecular biology of chromosome segregation in meiosis. Cell 112, 423–440 (2003)
Östergren, G. The mechanism of co-orientation in bivalents and multivalents. Hereditas 37, 85–156 (1951)
Brenner, S., Pepper, D., Berns, M. W., Tan, E. & Brinkley, B. R. Kinetochore structure, duplication, and distribution in mammalian cells: analysis by human autoantibodies from scleroderma patients. J. Cell Biol. 91, 95–102 (1981)
Goldstein, L. S. Kinetochore structure and its role in chromosome orientation during the first meiotic division in male D. melanogaster . Cell 25, 591–602 (1981)
Parra, M. T. et al. Involvement of the cohesin Rad21 and SCP3 in monopolar attachment of sister kinetochores during mouse meiosis I. J. Cell Sci. 117, 1221–1234 (2004)
Lee, J. et al. Specific regulation of CENP-E and kinetochores during meiosis I/meiosis II transition in pig oocytes. Mol. Reprod. Dev. 56, 51–62 (2000)
Hauf, S. & Watanabe, Y. Kinetochore orientation in mitosis and meiosis. Cell 119, 317–327 (2004)
Takahashi, K. et al. A low copy number central sequence with strict symmetry and unusual chromatin structure in fission yeast centromere. Mol. Biol. Cell 3, 819–835 (1992)
Pidoux, A. & Allshire, R. Kinetochore and heterochromatin domains of the fission yeast centromere. Chromosome Res. 12, 521–534 (2004)
Watanabe, Y., Yokobayashi, S., Yamamoto, M. & Nurse, P. Pre-meiotic S phase is linked to reductional chromosome segregation and recombination. Nature 409, 359–363 (2001)
Tomonaga, T. et al. Characterization of fission yeast cohesin: essential anaphase proteolysis of Rad21 phosphorylated in the S phase. Genes Dev. 14, 2757–2770 (2000)
Bernard, P. et al. Requirement of heterochromatin for cohesion at centromeres. Science 294, 2539–2542 (2001)
Nonaka, N. et al. Recruitment of cohesin to heterochromatic regions by Swi6/HP1 in fission yeast. Nature Cell Biol. 4, 89–93 (2002)
Yokobayashi, S. & Watanabe, Y. The kinetochore protein Moa1 enables cohesion-mediated monopolar attachment at meiosis I. Cell 123, 803–817 (2005)
Toth, A. et al. Functional genomics identifies monopolin: a kinetochore protein required for segregation of homologs during meiosis I. Cell 103, 1155–1168 (2000)
Monje-Casas, F., Prabhu, V. R., Lee, B. H., Boselli, M. & Amon, A. Kinetochore orientation during meiosis is controlled by Aurora B and the monopolin complex. Cell 128, 477–490 (2007)
Araki, H. et al. Site-specific recombinase, R, encoded by yeast plasmid pSR1. J. Mol. Biol. 225, 25–37 (1992)
Chang, C. R., Wu, C. S., Hom, Y. & Gartenberg, M. R. Targeting of cohesin by transcriptionally silent chromatin. Genes Dev. 19, 3031–3042 (2005)
Yokobayashi, S., Yamamoto, M. & Watanabe, Y. Cohesins determine the attachment manner of kinetochores to spindle microtubules at meiosis I in fission yeast. Mol. Cell. Biol. 23, 3965–3973 (2003)
Watanabe, Y. & Nurse, P. Cohesin Rec8 is required for reductional chromosome segregation at meiosis. Nature 400, 461–464 (1999)
Goshima, G. & Yanagida, M. Establishing biorientation occurs with precocious separation of sister kinetochores, but not the arms, in the early spindle of budding yeast. Cell 100, 619–633 (2000)
He, X., Asthana, S. & Sorger, P. K. Transient sister chromatid separation and elastic deformation of chromosomes during mitosis in budding yeast. Cell 101, 763–775 (2000)
Ocampo-Hafalla, M. T., Katou, Y., Shirahige, K. & Uhlmann, F. Displacement and re-accumulation of centromeric cohesin during transient pre-anaphase centromere splitting. Chromosoma 116, 531–544 (2007)
Rieder, C. L. & Salmon, E. D. Motile kinetochores and polar ejection forces dictate chromosome position on the vertebrate mitotic spindle. J. Cell Biol. 124, 223–233 (1994)
Kurzbauer, R. et al. Crystal structure of the p14/MP1 scaffolding complex: how a twin couple attaches mitogen-activated protein kinase signaling to late endosomes. Proc. Natl Acad. Sci. USA 101, 10984–10989 (2004)
Gruber, S. et al. Evidence that loading of cohesin onto chromosomes involves opening of its SMC hinge. Cell 127, 523–537 (2006)
Kitajima, T. S., Kawashima, S. A. & Watanabe, Y. The conserved kinetochore protein shugoshin protects centromeric cohesion during meiosis. Nature 427, 510–517 (2004)
Rabitsch, K. P. et al. Two fission yeast homologs of Drosophila Mei-S332 are required for chromosome segregation during meiosis I and II. Curr. Biol. 14, 287–301 (2004)
Yamamoto, A. et al. Spindle checkpoint activation at meiosis I advances anaphase II onset via meiosis-specific APC/C regulation. J. Cell Biol. 182, 277–288 (2008)
Tanaka, T., Fuchs, J., Loidl, J. & Nasmyth, K. Cohesin ensures bipolar attachment of microtubules to sister centromeres and resists their precocious separation. Nature Cell Biol. 2, 492–499 (2000)
Sullivan, B. A. & Karpen, G. H. Centromeric chromatin exhibits a histone modification pattern that is distinct from both euchromatin and heterochromatin. Nature Struct. Mol. Biol. 11, 1076–1083 (2004)
Amor, D. J., Kalitsis, P., Sumer, H. & Choo, K. H. A. Building the centromere: from foundation proteins to 3D organization. Trends Cell Biol. 14, 359–368 (2004)
Dewar, H., Tanaka, K., Nasmyth, K. & Tanaka, T. U. Tension between two kinetochores suffices for their bi-orientation on the mitotic spindle. Nature 428, 93–97 (2004)
Eckert, C. A., Gravdahl, D. J. & Megee, P. C. The enhancement of pericentromeric cohesin association by conserved kinetochore components promotes high-fidelity chromosome segregation and is sensitive to microtubule-based tension. Genes Dev. 21, 278–291 (2007)
Blower, M. D., Sullivan, B. A. & Karpen, G. H. Conserved organization of centromeric chromatin in flies and humans. Dev. Cell 2, 319–330 (2002)
Cleveland, D. W., Mao, Y. & Sullivan, K. F. Centromeres and kinetochores. From epigenetics to mitotic checkpoint signaling. Cell 112, 407–421 (2003)
Tanaka, T. U. et al. Evidence that the Ipl1–Sli15 (Aurora kinase–INCENP) complex promotes chromosome bi-orientation by altering kinetochore–spindle pole connections. Cell 108, 317–329 (2002)
Cimini, D., Wan, X., Hirel, C. B. & Salmon, E. D. Aurora kinase promotes turnover of kinetochore microtubules to reduce chromosome segregation errors. Curr. Biol. 16, 1711–1718 (2006)
Kawashima, S. A. et al. Shugoshin enables tension-generating attachment of kinetochores by loading Aurora to centromeres. Genes Dev. 21, 420–435 (2007)
Winey, M., Morgan, G. P., Straight, P. D., Giddings, T. H. & Mastronarde, D. N. Three-dimensional ultrastructure of Saccharomyces cerevisiae meiotic spindles. Mol. Biol. Cell 16, 1178–1188 (2005)
Yu, H.-G. & Dawe, R. K. Functional redundancy in the maize meiotic kinetochore. J. Cell Biol. 151, 131–141 (2000)
Chelysheva, L. et al. AtREC8 and AtSCC3 are essential to the monopolar orientation of the kinetochores during meiosis. J. Cell Sci. 118, 4621–4632 (2005)
Yamamoto, A. & Hiraoka, Y. Monopolar spindle attachment of sister chromatids is ensured by two distinct mechanisms at the first meiotic division in fission yeast. EMBO J. 22, 2284–2296 (2003)
Nabeshima, K. et al. Dynamics of centromeres during metaphase–anaphase transition in fission yeast: dis1 is implicated in force balance in metaphase bipolar spindle. Mol. Biol. Cell 9, 3211–3225 (1998)
Horie, S. et al. The Schizosaccharomyces pombe mei4+ gene encodes a meiosis-specific transcription factor containing a forkhead DNA-binding domain. Mol. Cell. Biol. 18, 2118–2129 (1998)
Izawa, D., Goto, M., Yamashita, A., Yamano, H. & Yamamoto, M. Fission yeast Mes1p ensures the onset of meiosis II by blocking degradation of cyclin Cdc13p. Nature 434, 529–533 (2005)
Allshire, R. C., Nimmo, E. R., Ekwall, K., Javerzat, J. P. & Cranston, G. Mutations derepressing silent centromeric domains in fission yeast disrupt chromosome segregation. Genes Dev. 9, 218–233 (1995)
Acknowledgements
We thank S. Hauf for critically reading the manuscript. We thank H. Matsuzaki, A. Yamamoto, R. Allshire, K. Nasmyth and the Yeast Genetic Resource Center (YGRC) for yeast strains. We also thank all the members of our laboratory for their support and discussion, especially S. Yokobayashi for materials and assistance in the initial stage of this project. This work was supported in part by Special Coordination Funds for Promoting Science and Technology (to T.S.), the Global COE Program (Integrative Life Science Based on the Study of Biosignaling Mechanisms), MEXT, Japan, and a Grant-in-Aid for Specially Promoted Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan (to Y.W.).
Author Contributions T.S. and Y.W. conceived and designed the experiments. T.S. performed all experiments. K.T. set up the TetR–tdTomato system in fission yeast. Y.W. planned research and wrote the manuscript with input from T.S.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Table 1, Supplementary Figures 1-13 with Legends and Supplementary References. (PDF 3130 kb)
Supplementary Movie 1
This movie shows wild-type prophase I zygote classified as "No exc." in Fig.1b. (MOV 122 kb)
Supplementary Movie 2
This movie shows wild-type prophase I zygote classified as "One exc." in Fig.1b. (MOV 126 kb)
Supplementary Movie 3
This movie shows wild-type prophase I zygote classified as "Cohered" in Fig.1b. (MOV 91 kb)
Supplementary Movie 4
This movie shows wild-type prophase I zygote classified as "Separated" in Fig.1b. (MOV 112 kb)
Rights and permissions
About this article
Cite this article
Sakuno, T., Tada, K. & Watanabe, Y. Kinetochore geometry defined by cohesion within the centromere. Nature 458, 852–858 (2009). https://doi.org/10.1038/nature07876
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature07876
This article is cited by
-
A TILLING by sequencing approach to identify induced mutations in sunflower genes
Scientific Reports (2021)
-
Pomegranate: 2D segmentation and 3D reconstruction for fission yeast and other radially symmetric cells
Scientific Reports (2020)
-
Untimely expression of gametogenic genes in vegetative cells causes uniparental disomy
Nature (2017)
-
The distribution of α-kleisin during meiosis in the holocentromeric plant Luzula elegans
Chromosome Research (2016)
-
No longer a nuisance: long non-coding RNAs join CENP-A in epigenetic centromere regulation
Cellular and Molecular Life Sciences (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.