Abstract
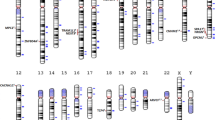
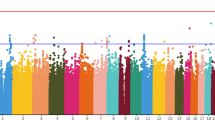
Little is known for certain about the genetics of schizophrenia. The advent of genomewide association has been widely anticipated as a promising means to identify reproducible DNA sequence variation associated with this important and debilitating disorder. A total of 738 cases with DSM-IV schizophrenia (all participants in the CATIE study) and 733 group-matched controls were genotyped for 492 900 single-nucleotide polymorphisms (SNPs) using the Affymetrix 500K two-chip genotyping platform plus a custom 164K fill-in chip. Following multiple quality control steps for both subjects and SNPs, logistic regression analyses were used to assess the evidence for association of all SNPs with schizophrenia. We identified a number of promising SNPs for follow-up studies, although no SNP or multimarker combination of SNPs achieved genomewide statistical significance. Although a few signals coincided with genomic regions previously implicated in schizophrenia, chance could not be excluded. These data do not provide evidence for the involvement of any genomic region with schizophrenia detectable with moderate sample size. However, a planned genomewide association study for response phenotypes and inclusion of individual phenotype and genotype data from this study in meta-analyses hold promise for eventual identification of susceptibility and protective variants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Saha S, Chant D, Welham J, McGrath J . A systematic review of the prevalence of schizophrenia. PLoS Med 2005; 2: e141.
Lichtenstein P, Bjork C, Hultman CM, Scolnick EM, Sklar P, Sullivan PF . Recurrence risks for schizophrenia in a Swedish national cohort. Psychol Med 2006; 36: 1417–1426.
Gottesman II, Shields J . Schizophrenia: The Epigenetic Puzzle. Cambridge University Press: Cambridge, UK, 1982.
Sullivan PF, Kendler KS, Neale MC . Schizophrenia as a complex trait: evidence from a meta-analysis of twin studies. Arch Gen Psychiatry 2003; 60: 1187–1192.
Falconer DS, Mackay TFC . Introduction to Quantitative Genetics, 4th edn Longman Group Ltd: London, 1996.
Risch N . Linkage strategies for genetically complex traits: I. Multilocus models. Am J Hum Genet 1990; 46: 222–228.
Neale B, Ferreira M, Medland S, Posthuma D . Statistical Genetics: Gene Mapping through Linkage and Association. Taylor and Francis: London, 2007.
Sullivan PF . The genetics of schizophrenia. PLoS Med 2005; 2: 614–618.
Stefansson H, Sigurdsson E, Steinthorsdottir V, Bjornsdottir S, Sigmundsson T, Ghosh S et al. Neuregulin 1 and susceptibility to schizophrenia. Am J Hum Genet 2002; 71: 877–892.
Li D, Collier DA, He L . Meta-analysis shows strong positive association of the neuregulin 1 (NRG1) gene with schizophrenia. Hum Mol Genet 2006; 15: 1995–2002.
Straub RE, Jiang Y, MacLean CJ, Ma Y, Webb BT, Myakishev MV et al. Genetic variation in the 6p22.3 gene DTNBP1, the human ortholog of the mouse Dysbindin gene, is associated with schizophrenia. Am J Hum Genet 2002; 71: 337–348. Erratum in American Journal of Human Genetics 2002 Oct; 72(4): 1007.
Mutsuddi M, Morris DW, Waggoner SG, Daly MJ, Scolnick EM, Sklar P . Analysis of high-resolution HapMap of DTNBP1 (Dysbindin) suggests no consistency between reported common variant associations and schizophrenia. Am J Hum Genet 2006; 79: 903–909.
Millar JK, Wilson-Annan JC, Anderson S, Christie S, Taylor MS, Semple CA et al. Disruption of two novel genes by a translocation co-segregating with schizophrenia. Hum Mol Genet 2000; 9: 1415–1423.
Sullivan PF . Spurious genetic associations. Biol Psychiatry 2007; 61: 1121–1126.
Frayling TM, Timpson NJ, Weedon MN, Zeggini E, Freathy RM, Lindgren CM et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 2007; 316: 889–894.
Klein RJ, Zeiss C, Chew EY, Tsai JY, Sackler RS, Haynes C et al. Complement factor H polymorphism in age-related macular degeneration. Science 2005; 308: 385–389.
Duerr RH, Taylor KD, Brant SR, Rioux JD, Silverberg MS, Daly MJ et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science 2006; 314: 1461–1463.
WTCCC. Genome-wide association study of 14 000 cases of seven common diseases and 3000 shared controls. Nature 2007; 447: 661–678.
Saxena R, Voight BF, Lyssenko V, Burtt NP, de Bakker PI, Chen H et al. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 2007; 316: 1331–1336.
Scott LJ, Mohlke KL, Bonnycastle LL, Willer CJ, Li Y, Duren WL et al. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science 2007; 316: 1341–1345.
Steinthorsdottir V, Thorleifsson G, Reynisdottir I, Benediktsson R, Jonsdottir T, Walters GB et al. A variant in CDKAL1 influences insulin response and risk of type 2 diabetes. Nat Genet 2007; 39: 770–775.
Zeggini E, Weedon MN, Lindgren CM, Frayling TM, Elliott KS, Lango H et al. Replication of genome-wide association signals in U.K. samples reveals risk loci for type 2 diabetes. Science 2007; 316: 1336–1341.
Lencz T, Morgan TV, Athanasiou M, Dain B, Reed CR, Kane JM et al. Converging evidence for a pseudoautosomal cytokine receptor gene locus in schizophrenia. Mol Psychiatry 2007; 12: 572–580.
Stroup TS, McEvoy JP, Swartz MS, Byerly MJ, Glick ID, Canive JM et al. The National Institute of Mental Health Clinical Antipsychotic Trials of Intervention Effectiveness (CATIE) project: schizophrenia trial design and protocol development. Schizophr Bull 2003; 29: 15–31.
Lieberman JA . Comparative effectiveness of antipsychotic drugs. A commentary on: Cost Utility Of The Latest Antipsychotic Drugs In Schizophrenia Study (CUtLASS 1) and Clinical Antipsychotic Trials Of Intervention Effectiveness (CATIE). Arch Gen Psychiatry 2006; 63: 1069–1072.
Chanock SJ, Manolio T, Boehnke M, Boerwinkle E, Hunter DJ, Thomas G et al. Replicating genotype–phenotype associations. Nature 2007; 447: 655–660.
Lieberman JA, Stroup TS, McEvoy JP, Swartz MS, Rosenheck RA, Perkins DO et al. Effectiveness of antipsychotic drugs in patients with chronic schizophrenia. N Engl J Med 2005; 353: 1209–1223.
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders, 4th edn. American Psychiatric Association: Washington, DC, 1994.
First M, Spitzer R, Gibbon M, Williams J . Structured Clinical Interview for DSM-IV Axis I Disorders––Administration Booklet. American Psychiatric Press Inc.: Washington DC, 1994.
Affymetrix. GeneChip® Mapping 500K Assay Manual. Santa Clara, CA, 2006.
Bierut LJ, Madden PA, Breslau N, Johnson EO, Hatsukami D, Pomerleau OF et al. Novel genes identified in a high-density genome wide association study for nicotine dependence. Hum Mol Genet 2007; 16: 24–35.
Hinds DA, Stuve LL, Nilsen GB, Halperin E, Eskin E, Ballinger DG et al. Whole-genome patterns of common DNA variation in three human populations. Science 2005; 307: 1072–1079.
Affymetrix. BRLMM: an Improved Genotype Calling Method for the GeneChip® Human Mapping 500 K Array Set. Santa Clara, CA, 2006, 4/14/2006.
GAIN Collaborative Research Group. New models of collaboration in genome-wide association studies: the Genetic Association Information Network (GAIN). Nat Genet 2007; 39: 1045–1051.
Sullivan PF, Purcell S . Analyzing genomewide association study data: a tutorial. In: Neale B, Ferreira M, Medland S, Posthuma D (eds). Statistical Genetics: Gene Mapping through Linkage and Association. London: Taylor and Francis, 2007 p. 355–394.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira M, Bender D et al. PLINK: a toolset for whole-genome association and population-based linkage analysis. Am J Hum Genet 2007; 81: 559–575.
Wigginton JE, Cutler DJ, Abecasis GR . A note on exact tests of Hardy–Weinberg equilibrium. Am J Hum Genet 2005; 76: 887–893.
Wittke-Thompson JK, Pluzhnikov A, Cox NJ . Rational inferences about departures from Hardy–Weinberg equilibrium. Am J Hum Genet 2005; 76: 967–986.
Marchini J, Cardon LR, Phillips MS, Donnelly P . The effects of human population structure on large genetic association studies. Nat Genet 2004; 36: 512–517.
Heiman GA, Hodge SE, Gorroochurn P, Zhang J, Greenberg DA . Effect of population stratification on case–control association studies. I. Elevation in false positive rates and comparison to confounding risk ratios (a simulation study). Hum Hered 2004; 58: 30–39.
Shields AE, Fortun M, Hammonds EM, King PA, Lerman C, Rapp R et al. The use of race variables in genetic studies of complex traits and the goal of reducing health disparities. Am Psychol 2005; 60: 77–103.
Devlin B, Roeder K . Genomic control for association studies. Biometrics 1999; 55: 997–1004.
Pritchard JK, Stephens M, Rosenberg NA, Donnelley P . Association mapping in structured populations. Am J Hum Genet 2000; 67: 170–181.
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D . Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 2006; 38: 904–909.
Epstein MP, Allen AS, Satten GA . A simple and improved correction for population stratification in case–control studies. Am J Hum Genet 2007; 80: 921–930.
Lee S, Sullivan PF, Zou F, Wright FA . Comment on a simple and improved correction of population stratification. Am J Hum Genet 2008; 82: 524–526.
Patterson N, Price AL, Reich D . Population structure and eigenanalysis. PLoS Genet 2006; 22: e190.
Lettre G, Lange C, Hirschhorn JN . Genetic model testing and statistical power in population-based association studies of quantitative traits. Genet Epidemiol 2007; 31: 358–362.
Storey JD . The positive false discovery rate: a Bayesian interpretation and the q-value. Ann Stat 2003; 31: 2013–2035.
Storey JD, Tibshirani R . Statistical significance for genomewide studies. Proc Natl Acad Sci USA 2003; 100: 9440–9445.
van den Oord EJ, Sullivan PF . False discoveries and models for gene discovery. Trends Genet 2003; 19: 537–542.
van den Oord EJCG . Controlling false discoveries in genetic studies. Am J Med Genet B Neuropsychiatr Genet, (in press).
Benjamini Y, Hochberg Y . Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B 1995; 57: 289–300.
Brown BW, Russell K . Methods of correcting for multiple testing: operating characteristics. Stat Med 1997; 16: 2511–2528.
Fernando RL, Nettleton D, Southey BR, Dekkers JC, Rothschild MF, Soller M . Controlling the proportion of false positives in multiple dependent tests. Genetics 2004; 166: 611–619.
van den Oord EJ, Sullivan PF . A framework for controlling false discovery rates and minimizing the amount of genotyping in the search for disease mutations. Hum Hered 2003; 56: 188–199.
Tsai CA, Hsueh HM, Chen JJ . Estimation of false discovery rates in multiple testing: application to gene microarray data. Biometrics 2003; 59: 1071–1081.
van den Oord EJ . Controlling false discoveries in candidate gene studies. Mol Psychiatry 2005; 10: 230–231.
Sabatti C, Service S, Freimer N . False discovery rate in linkage and association genome screens for complex disorders. Genetics 2003; 164: 829–833.
Meinhausen N, Rice J . Estimating the proportion of false null hypotheses among a large number of independently tested hypotheses. Annals of Statistics 2006; 34: 373–393.
Schaid DJ, Rowland CM, Tines DE, Jacobson RM, Poland GA . Score tests for association between traits and haplotypes when linkage phase is ambiguous. Am J Hum Genet 2002; 70: 425–434.
R Development Core Team. R: A Language And Environment For Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria, 2005.
Gabriel SB, Schaffner SF, Nguyen H, Moore JM, Roy J, Blumenstiel B et al. The structure of haplotype blocks in the human genome. Science 2002; 296: 2225–2229.
Barrett JC, Fry B, Maller J, Daly MJ . Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 2005; 21: 263–265.
Altshuler D, Brooks LD, Chakravarti A, Collins FS, Daly MJ, Donnelly P . A haplotype map of the human genome. Nature 2005; 437: 1299–1320.
Gauderman WJ . Sample size requirements for association studies of gene-gene interaction. Am J Epidemiol 2002; 155: 478–484.
Gauderman WJ . Sample size requirements for matched case-control studies of gene-environment interaction. Stat Med 2002; 21: 35–50.
SAS Institute Inc. SAS/STAT® Software: Version 9. SAS Institute, Inc.: Cary, NC, 2004.
SAS Institute Inc. JMP User's Guide (Version 6). SAS Institute, Inc.: Cary, NC, 2005.
Hinrichs AS, Karolchik D, Baertsch R, Barber GP, Bejerano G, Clawson H et al. The UCSC Genome Browser Database: update 2006. Nucleic Acids Res 2006; 34 (Database issue): D590–D598.
Wheeler DL, Barrett T, Benson DA, Bryant SH, Canese K, Chetvernin V et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 2006; 34 (Database issue): D173–D180.
Blaschke RJ, Rappold G . The pseudoautosomal regions, SHOX and disease. Curr Opin Genet Dev 2006; 16: 233–239.
Hemminger BM, Saelim B, Sullivan PF . TAMAL: An integrated approach to choosing SNPs for genetic studies of human complex traits. Bioinformatics 2006; 22: 626–627.
Fisher RA . Statistical Methods for Reesearch Workers (11th edition). Oliver and Boyd: London, 1950.
Kay SR, Fiszbein A, Opler LA . The positive and negative syndrome scale (PANSS) for schizophrenia. Schizophr Bull 1987; 13: 261–276.
Sullivan PF, Keefe RSE, Lange LA, Lange EM, Stroup TS, Lieberman JA et al. NCAM1 and neurocognition in schizophrenia. Biological Psychiatry 2007; 61: 902–910.
Balding DJ . A tutorial on statistical methods for population association studies. Nat Rev Genet 2006; 7: 781–791.
Sachse C, Ruschen S, Dettling M, Schley J, Bauer S, Muller-Oerlinghausen B et al. Flavin monooxygenase 3 (FMO3) polymorphism in a white population: allele frequencies, mutation linkage, and functional effects on clozapine and caffeine metabolism. Clin Pharmacol Ther 1999; 66: 431–438.
Farrer LA, Cupples LA, Haines JL, Hyman B, Kukull WA, Mayeux R et al. Effects of age, sex, and ethnicity on the association between apolipoprotein E genotype and Alzheimer disease. A meta-analysis. APOE and Alzheimer Disease Meta Analysis Consortium. Jama 1997; 278: 1349–1356.
Hsu F, Kent WJ, Clawson H, Kuhn RM, Diekhans M, Haussler D . The UCSC Known Genes. Bioinformatics 2006; 22: 1036–1046.
Mah S, Nelson MR, Delisi LE, Reneland RH, Markward N, James MR et al. Identification of the semaphorin receptor PLXNA2 as a candidate for susceptibility to schizophrenia. Mol Psychiatry 2006; 11: 471–478.
Hennah W, Varilo T, Kestila M, Paunio T, Arajarvi R, Haukka J et al. Haplotype transmission analysis provides evidence of association for DISC1 to schizophrenia and suggests sex-dependent effects. Hum Mol Genet 2003; 12: 3151–3159.
Cannon TD, Hennah W, van Erp TG, Thompson PM, Lonnqvist J, Huttunen M et al. Association of DISC1/TRAX haplotypes with schizophrenia, reduced prefrontal gray matter, and impaired short- and long-term memory. Arch Gen Psychiatry 2005; 62: 1205–1213.
Law AJ, Lipska BK, Weickert CS, Hyde TM, Straub RE, Hashimoto R et al. Neuregulin 1 transcripts are differentially expressed in schizophrenia and regulated by 5′ SNPs associated with the disease. Proc Natl Acad Sci USA 2006; 103: 6747–6752.
Hall J, Whalley HC, Job DE, Baig BJ, McIntosh AM, Evans KL et al. A neuregulin 1 variant associated with abnormal cortical function and psychotic symptoms. Nat Neurosci 2006; 9: 1477–1478.
Crowley JJ, Keefe RS, Perkins DO, Stroup TS, Lieberman JA, Sullivan PF . The neuregulin 1 promoter polymorphism rs6994992 is not associated with chronic schizophrenia or neurocognition. Am J Med Genet B Neuropsychiatr Genet 2008; e-pub ahead of print 19 February 2008. PMID: 18286587.
Shifman S, Bronstein M, Sternfeld M, Pisante-Shalom A, Lev-Lehman E, Weizman A et al. A highly significant association a COMT haplotype and schizophrenia. Am J Hum Genet 2002; 71: 1296–1302.
Skol AD, Scott LJ, Abecasis GR, Boehnke M . Joint analysis is more efficient than replication-based analysis for two-stage genome-wide association studies. Nat Genet 2006; 38: 209–213.
Sladek R, Rocheleau G, Rung J, Dina C, Shen L, Serre D et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 2007; 445: 881–885.
Frayling TM . Genome-wide association studies provide new insights into type 2 diabetes aetiology. Nat Rev Genet 2007; 8: 657–662.
Manolio TA, Rodriguez LL, Brooks L, Abecasis G, Ballinger D, Daly M et al. New models of collaboration in genome-wide association studies: the Genetic Association Information Network. Nat Genet 2007; 39: 1045–1051.
Barrett JC, Cardon LR . Evaluating coverage of genome-wide association studies. Nat Genet 2006; 38: 659–662.
Miller D, McEvoy JP, Davis SM, Caroff SN, Saltz BL, Chakos MH et al. Clinical correlates of tardive dyskinesia in schizophrenia: baseline data from the CATIE schizophrenia trial. Schizophr Res 2005; 80: 33–43.
Keefe RS, Bilder RM, Harvey PD, Davis SM, Palmer BW, Gold JM et al. Baseline neurocognitive deficits in the CATIE schizophrenia trial. Neuropsychopharmacology 2006; 31: 2033–2046.
World Health Organization. The ICD-10 Classification of Mental and Behavioural Disorders: Diagnostic Criteria for Research. World Health Organization: Geneva, 1993.
McClellan JM, Susser E, King MC . Schizophrenia: a common disease caused by multiple rare alleles. Br J Psychiatry 2007; 190: 194–199.
Acknowledgements
Dr Sullivan was supported by R01s MH074027 and MH077139, Dr Zou by GM074175 and Dr Wright by P30 HD003110. The CATIE project was funded by NIMH contract N01 MH90001. We thank Dr Nick Patterson for assistance with the EigenSoft program. We are indebted to the ‘Molecular Genetics of Schizophrenia II’ (MGS-2) collaboration for their exceptional collegiality in making the control samples available to the research community. Control subjects were from the National Institute of Mental Health Schizophrenia Genetics Initiative, and phenotypes and DNA samples were collected by the MGS-2 collaboration whose investigators and co-investigators are Evanston Northwestern Healthcare (Northwestern University, Evanston, IL, USA; MH059571), Pablo V Gejman, MD (Collaboration Coordinator; PI), Alan R Sanders, MD (Emory University School of Medicine, Atlanta, GA, USA; MH59587), Farooq Amin, MD (PI) (Louisiana State University Health Sciences Center, New Orleans, LA, USA; MH067257), Nancy Buccola APRN, BC, MSN (PI) (University of California-Irvine, Irvine, CA, USA; MH60870), William Byerley, MD (PI) (Washington University, St Louis, MO, USA; U01, MH060879), C Robert Cloninger, MD (PI) (University of Iowa, Iowa, IA, USA; MH59566), Raymond Crowe, MD (PI), Donald Black, MD (University of Colorado, Denver, CO, USA; MH059565), Robert Freedman, MD (PI) (University of Pennsylvania, Philadelphia, PA, USA; MH061675), Douglas Levinson MD (PI) (University of Queensland, QLD, Australia; MH059588), Bryan Mowry, MD (PI) (Mt Sinai School of Medicine, New York, NY, USA; MH59586), Jeremy Silverman, PhD (PI).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Financial disclosures
Eli Lilly funded the GWAS genotyping done at Perlegen Sciences. Dr Sullivan reports receiving research funding from Eli Lilly in connection with this project. Dr Stroup reports having received research funding from Eli Lilly and consulting fees from Janssen Pharmaceutica, GlaxoSmithKline and Bristol-Myers Squibb. Dr Lieberman reports having received research funding from AstraZeneca Pharmaceuticals, Bristol-Myers Squibb, GlaxoSmithKline, Janssen Pharmaceutica and Pfizer, and consulting and educational fees from AstraZeneca Pharmaceuticals, Bristol-Myers Squibb, Eli Lilly, Forest Pharmaceuticals, GlaxoSmithKline, Janssen Pharmaceutica, Novartis, Pfizer and Solvay.
Supplementary Information accompanies the paper on the Molecular Psychiatry website (http://www.nature.com/mp)
Supplementary information
Rights and permissions
About this article
Cite this article
Sullivan, P., Lin, D., Tzeng, JY. et al. Genomewide association for schizophrenia in the CATIE study: results of stage 1. Mol Psychiatry 13, 570–584 (2008). https://doi.org/10.1038/mp.2008.25
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2008.25
Keywords
This article is cited by
-
Genome-wide association study identified six loci associated with adverse drug reactions to aripiprazole in schizophrenia patients
Schizophrenia (2023)
-
Genome wide study of tardive dyskinesia in schizophrenia
Translational Psychiatry (2021)
-
Variants of GRM7 as risk factor and response to antipsychotic therapy in schizophrenia
Translational Psychiatry (2020)
-
A Powerful Test for SNP Effects on Multivariate Binary Outcomes Using Kernel Machine Regression
Statistics in Biosciences (2018)
-
Prediction of Schizophrenia Diagnosis by Integration of Genetically Correlated Conditions and Traits
Journal of Neuroimmune Pharmacology (2018)