Abstract
Epithelioid fibrous histiocytoma is a rare and distinctive cutaneous neoplasm. Most cases harbor ALK rearrangement and show ALK overexpression, which distinguish this neoplasm from conventional cutaneous fibrous histiocytoma and variants. SQSTM1 and VCL have previously been shown to partner with ALK in one case each of epithelioid fibrous histiocytoma. The purpose of this study was to examine a large cohort of epithelioid fibrous histiocytomas by next-generation sequencing to characterize the nature and prevalence of ALK fusion partners. A retrospective archival review was performed to identify cases of epithelioid fibrous histiocytoma (2012–2016). Immunohistochemistry was performed to confirm ALK expression. Targeted next-generation sequencing was applied on RNA extracted from formalin-fixed paraffin-embedded tissue to identify the fusion partners. Twenty-three cases fulfilled inclusion criteria. The mean patient age was 39 years (range, 8–74), there was no sex predilection, and >75% of cases involved the lower extremities. The most common gene fusions were SQSTM1-ALK (N=12; 52%) and VCL-ALK (N=7; 30%); the other four cases harbored novel fusion partners (DCTN1, ETV6, PPFIBP1, and SPECC1L). The pattern of ALK immunoreactivity was usually granular cytoplasmic (N=12; 52%) or granular cytoplasmic and nuclear (N=10; 43%); the case containing an ETV6 fusion partner showed nuclear staining alone. There was no apparent relationship between tumor morphology and the ALK fusion partner. In summary, SQSTM1 and VCL are the most common ALK fusion partners in epithelioid fibrous histiocytoma; DCTN1, ETV6, PPFIBP1, and SPECC1L represent rare fusion partners. The proteins encoded by these genes play diverse roles in scaffolding, cell adhesion, signaling, and transcription (among others) without clear commonalities. These findings expand the oncogenic promiscuity of many of these ALK fusion genes, which drive neoplasia in tumors of diverse lineages with widely varied clinical behavior. This is the first documented account of ETV6-ALK and SPECC1L-ALK translocations in neoplasms.
Similar content being viewed by others
Main
Epithelioid fibrous histiocytoma, also known as epithelioid cell histiocytoma, is a rare and enigmatic cutaneous neoplasm.1, 2 Clinically, lesions often resemble pyogenic granuloma.1, 2 Tumors tend to be small and solitary; they occur across a broad age-range, predominating between the third and seventh decades; and the extremities represent the most frequent sites of involvement.1, 2 Morphologically, epithelioid fibrous histiocytoma is frequently exophytic, with an epidermal collarette.2 Centered in the dermis, tumors are comprised of plump epithelioid cells occasionally with a spindle cell component.1, 2 The nuclei are round-ovoid, with minimal atypia, vesicular chromatin and scattered bi- and tri-nucleation; mitotic activity is typically scant.
Originally considered a variant of fibrous histiocytoma,1 subsequent studies emphasizing morphologic and immunohistochemical differences challenged this notion.3 The discovery that epithelioid fibrous histiocytoma harbors canonical ALK translocations in a majority of tumors ultimately confirmed a biologic distinction.4 Next-generation sequencing of two cases recently identified VCL and SQSTM1 as distinct ALK fusion partners.5 VCL-ALK rearrangements have previously been reported in a distinctive group of pediatric renal cell carcinomas,6 while SQSTM1-ALK has been identified in ALK-positive large B-cell lymphoma.7 A diverse array of neoplasms has been reported to contain ALK fusion products, including inflammatory myofibroblastic tumor,8 lymphoma (eg, ALK-positive anaplastic large cell lymphoma, and ALK-positive large B-cell lymphoma),9, 10, 11 lung adenocarcinoma,12 renal cell carcinoma,13 thyroid carcinoma,14 and Spitz neoplasms.15 ALK has the potential to fuse with numerous partner genes, and disparate tumor types (with widely variable clinical behavior) may possess the exact same fusion partner. In view of the tremendous potential for overlap among ALK fusion partners, the purpose of this study was to interrogate a large cohort of cases of epithelioid fibrous histiocytoma to characterize the nature and prevalence of ALK products in this neoplasm.
Materials and methods
Following institutional Research Ethics Board approval, we performed a retrospective review of our archives for cases of epithelioid fibrous histiocytoma diagnosed between 2012 and 2016. Each case was re-reviewed to confirm the diagnosis. Immunohistochemistry for ALK was performed as previously reported to confirm ALK expression.4 The intent of this study was to characterize ALK partner genes; therefore, cases of epithelioid fibrous histiocytoma lacking ALK immunoexpression were not included. The pattern (i.e., subcellular localization) of ALK staining was recorded.
Each case was interrogated by a targeted next-generation sequencing fusion panel. Briefly, for each case 4–5 unstained slides cut from formalin-fixed paraffin-embedded tissue (4 microns) were scraped into Eppendorf tubes. RNA was extracted using the ExpressArt FFPE Clear RNA Ready kit (Amsbio, Cambridge, MA). RNA-seq libraries were prepared using 20–100 ng total RNA with the TruSight RNA Fusion Panel (Illumina, San Diego, CA), an enrichment-based assay that targets 507 known fusion-associated genes. Each sample was sequenced with 76 base-pair paired-end reads on an Illumina MiSeq at 8 samples per flow cell (~3 million reads per sample). The results were analyzed using the STAR aligner and Manta fusion caller as well as the JAFFA fusion caller utilizing BOWTIE2 aligner.16, 17
Each fusion product was subsequently independently confirmed by reverse transcriptase polymerase chain reaction (RT-PCR). Briefly, RT-PCR was performed on the extracted RNA using random primers and Moloney Murine Leukemia Virus (M-MLV) reverse transcriptase according to manufacturer’s instructions (Invitrogen, Burlington, Ontario, Canada). Polymerase chain reaction (PCR) was subsequently performed using primers specific for the fusion breakpoints (Supplementary Table 1). Forty cycles of PCR were performed at 94 °C for 30 s, 55 °C for one minute, and 72 °C for one minute using HotStarTaq DNA Polymerase (Qiagen, Toronto, Ontario, Canada). PCR products were sequenced on the Applied Biosystems 3730 DNA Analyzer using the Big Dye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Burlington, Ontario, Canada).
Results
A total of 24 cases fulfilled the inclusion criteria. Adequate RNA was obtained from all but one case, leaving 23 cases for molecular evaluation. Clinically, the mean patient age was 39 years (range: 8–74 years), and there was a similar number of females and males. The majority of tumors arose in the lower extremities (78%), with the thigh (30%) representing the most common site of involvement (Table 1).
Next-generation sequencing confirmed the presence of ALK rearrangement in all tumors (Table 1). In 12 cases (52%), SQSTM1 was the ALK fusion partner; 7 cases (30%) had VCL-ALK as the fusion product. Four cases contained novel fusion partners that have not been reported previously in epithelioid fibrous histiocytoma: DCTN1, ETV6, PPFIBP1, and SPECC1L (Figure 1). All fusion breakpoints involved exon 20 of the ALK gene (NCBI Reference Sequence: NM_004304.4). The fusion partner exons were: SQSTM1 exon 5 (NM_003900.4), VCL exon 16 (NM_014000.2), DCTN1 exon 21 (NM_023019.3), ETV6 exon 5 (NM_001987.4), PPFIBP1 exon 9 (NM_003622.3), and SPECC1L exon 9 (NM_015330.4). Each of the fusion products was independently confirmed by RT-PCR followed by Sanger sequencing (Figure 2).
The patterns of ALK expression by immunohistochemistry were as follows: granular cytoplasmic staining (N=12; 52%), granular cytoplasmic staining combined with nuclear staining (N=10; 43%), and nuclear staining alone (N=1; 4%). The SQSTM1-ALK fusions tended to be more often associated with granular cytoplasmic and nuclear ALK staining (75%), whereas VCL-ALK fusions generally had granular cytoplasmic immunoreactivity without nuclear staining (86%). Of the cases with novel fusion partners, the tumor with ETV6-ALK fusion showed nuclear staining alone, while the remaining cases showing granular cytoplasmic staining patterns.
There was no apparent association between morphology and the nature of the underlying ALK fusion partner. Tumors with SQSTM1-ALK and VCL-ALK fusions were morphologically and immunohistochemically similar to previous reports (Figure 3).1, 2, 4 Each of the four cases with novel fusion partners were polypoid; cases 20, 22 and 23 contained an epidermal collarette. Case 20 (with DCTN1-ALK) was ulcerated with mixed epithelioid and spindle cell morphology (Figure 4). Immunohistochemistry for ALK showed diffuse granular cytoplasmic staining. Case 21 (with ETV6-ALK) was separated from the epidermis by a Grenz zone, and had a somewhat irregular peripheral tumor margin; this was the only case to show a strictly nuclear pattern of ALK staining (Figure 4f). Case 22 (with PPFIBP1-ALK) and Case 23 (with SPECC1L-ALK) both had granular cytoplasmic ALK expression (Figure 5). Morphologically, the latter tumor was somewhat less cellular than the other tumors, with collagenous stroma interspersed among the tumor cells (Figures 5d and e).
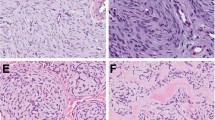
Representative photomicrographs of cases of epithelioid fibrous histiocytoma containing previously reported fusion products: (a–c) SQSTM1-ALK and (d–f) VCL-ALK fusion products. Note: granular cytoplasmic ALK staining in both tumors. (a and d) (hematoxylin and eosin, × 100); (b and e) (hematoxylin and eosin, × 200); (c and f) (immunohistochemistry for ALK, × 200).
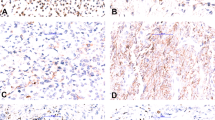
Representative photomicrographs of epithelioid fibrous histiocytoma with (a–c) DCTN1-ALK and (d–f) ETV6-ALK fusion products. Note: nuclear pattern of immunohistochemical staining with ETV6-ALK translocation. (a and d) (hematoxylin and eosin, × 100); (b and e) (hematoxylin and eosin, × 200); (c and f) (immunohistochemistry for ALK, × 200).
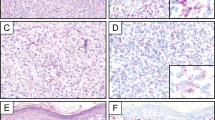
Representative photomicrographs of epithelioid fibrous histiocytoma with (a–c) PPFIBP1-ALK, and (d–f) SPECC1L-ALK fusion products. A delicate peripheral vasculature was a common finding among most tumors. Note: areas of interspersed collagenous stroma with SPECC1L-ALK translocation. (a and d) (hematoxylin and eosin, × 100); (b and e) (hematoxylin and eosin, × 200); (c and f) (immunohistochemistry for ALK, × 200).
Discussion
Epithelioid fibrous histiocytoma is a benign cutaneous neoplasm with a characteristic morphology and immunophenotype.1, 2, 3 The presence of ALK rearrangement, and concomitant ALK protein overexpression, previously confirmed this neoplasm to be biologically distinct from conventional fibrous histiocytoma and variants;4, 5 however, owing to its rarity, few cases have undergone rigorous molecular characterization. The goal of this study was to examine the nature and breadth of ALK fusion products in epithelioid fibrous histiocytoma. We show that SQSTM1 and VCL are the most prevalent ALK fusion partners. In addition, we report four novel ALK partners in this rare neoplasm: DCTN1, ETV6, PPFIBP1, and SPECC1L. Moreover, this is the first documented report of neoplasms containing disease-defining ETV6-ALK and SPECC1L-ALK fusion genes.
ALK (anaplastic lymphoma kinase; CD246) encodes a transmembrane receptor tyrosine kinase within the insulin receptor superfamily. Alterations in this gene–including translocations, mutations, copy number changes and dysregulated expression–have been implicated in numerous tumor types (reviewed in detail elsewhere 18, 19, 20). A heterogeneous group of neoplasms–spanning mesenchymal, epithelial, hematolymphoid and melanocytic lines of differentiation–have been reported to contain translocations involving ALK. Despite seemingly disparate ontogenies, many ALK fusion products are shared among different tumors. For example, EML4-ALK translocations occur in lung adenocarcinoma,21 inflammatory myofibroblastic tumor,22 ALK-positive large B-cell lymphoma23 and renal cell carcinoma.24, 25 There is tremendous variability among potential ALK fusion partners; further plasticity arises from the finding that other tyrosine kinase receptor genes may substitute for ALK, thereby yielding a potentially staggering number of fusion gene permutations. For example, in inflammatory myofibroblastic tumor TFG has been reported to partner with both ALK26 and ROS1.27
This study identified SQSTM1-ALK as the most common fusion product in epithelioid fibrous histiocytoma. SQSTM1 encodes sequestosome 1 (p62), a scaffolding protein with multiple roles including NFκB signaling and ubiquitin-mediated autophagy.28 Mutations of this gene have been implicated in multiple diseases, including Paget disease of bone and frontotemporal lobar degeneration. In this study, the SQSTM1-ALK fusion was present in slightly over half of cases. In addition to epithelioid fibrous histiocytoma,5 SQSTM1-ALK fusions have been reported previously in ALK-positive large B-cell lymphoma7 and lung adenocarcinoma,29 as well as inflammatory myofibroblastic tumor (Dickson, unpublished observation). Non-ALK fusion partners of SQSTM1 have also been reported, including NTRK3 in papillary thyroid carcinoma30 and FGFR1 in acute myelomonocytic leukemia.31
Slightly less than a third of cases in this study were characterized by VCL-ALK fusion products. VCL encodes vinculin, an adhesion protein linking the extracellular matrix to the actomyosin cytoskeleton. Vinculin is believed to play a role in cell migration, cell-matrix adhesion and cell-cell junctions.32, 33 Mutations in this gene have been implicated in dilated cardiomyopathy.34 In addition to epithelioid fibrous histiocytoma,5 VCL-ALK fusions have also been reported in some pediatric renal cell carcinomas associated with sickle cell trait.6, 13
This is the first report of DCTN1-ALK in epithelioid fibrous histiocytoma. DCTN1 encodes dynactin subunit 1, the largest subunit of the dynactin complex, which binds microtubules and cytoplasmic dynein. Dynactin is a microtubule-based biologic motor with diverse cellular functions, including cell division and intracytoplasmic vesicular and organelle movement.35 DCTN1-ALK translocations have previously been identified in Spitz tumor,36 pancreatic ductal carcinoma,37 inflammatory myofibroblastic tumor,38 and lung adenocarcinoma.29
ETV6 (ETS Variant 6; TEL) encodes an E-Twenty-Six (ETS) family transcription factor involved in DNA and protein binding, with important roles in development, differentiation and cell proliferation.39 Mutations in this gene have been linked to forms of thrombocytopenia and hematologic malignancy.40 This gene is promiscuous, forming fusion partners with numerous other genes in neoplasia. In addition to hematologic neoplasms, ETV6-NTRK3 fusions have been reported in congenital/infantile fibrosarcoma,41 congenital mesoblastic nephroma,42 gastrointestinal stromal tumor,43 infantile NTRK-associated mesenchymal tumors,44 inflammatory myofibroblastic tumor,45, 46 mammary analog secretory carcinoma, secretory breast carcinoma,47 sinonasal low-grade intestinal-type adenocarcinoma,48 Spitz tumor,49 and radiation-associated thyroid carcinoma.50 ETV6 fusions are not restricted to NTRK3; indeed, there are roughly 30 different ETV6 fusion partners among hematologic malignancies alone (reviewed in 39). To our knowledge, this is the first documented account of a neoplasm bearing an ETV6-ALK fusion product. Interestingly, this fusion was associated with a nuclear pattern of ALK immunoreactivity, whereas the other tumors in this study showed either granular cytoplasmic or a combination of granular cytoplasmic and nuclear staining. Different patterns of ALK staining have been associated with specific fusion partners;51 it remains to be determined whether this pattern is unique–and/or universal–to cases with an ETV6-ALK fusion product in epithelioid fibrous histiocytoma.
PPFIBP1 encodes PPFIA Binding Protein 1 (liprin beta 1), a leukocyte common antigen-related (LAR) protein-tyrosine phosphatase-interacting protein. This protein is believed to be involved in cell adhesion, migration and development.52 Translocations involving PPFIBP1-ALK have previously been reported in inflammatory myofibroblastic tumor;53 this is the first report of PPFIBP1 rearrangement in epithelioid fibrous histiocytoma. Non-ALK fusion partners of PPFIBP1 have also been identified, including ROS1 in Spitz nevus.54 SPECC1L (sperm antigen with calponin homology and coiled-coil domains 1) encodes a cytoskeletal cross-linking protein, with a purported role in cell adhesion, movement and division.55, 56 Mutations in this gene have been linked to abnormalities in craniofacial development.55 This is the first report of SPECC1L forming a somatic gene fusion with ALK in disease. We have previously encountered a single case involving a SPECC1L-NTRK2 fusion product; this was an undifferentiated spindle cell neoplasm with a prominent myxoid stroma (Dickson, unpublished observation).
Approximately 88% of epithelioid fibrous histiocytomas have evidence of ALK rearrangement.4 Presumably, in some cases other genes, such as ROS1, may substitute for ALK. Mechanistically it has not been fully resolved why ALK is so readily capable of translocating with other partner genes in neoplasia. In some translocation-associated sarcomas, such as synovial sarcoma, multiple partners occur as a result of sequence homology within the partner gene; however, this does not appear to account for the breadth of ALK partners in epithelioid fibrous histiocytoma. Further molecular characterization of epithelioid fibrous histiocytoma, and other “ALKomas”, is warranted.
In summary, these results suggest that SQSTM1 is the most common ALK fusion partner, followed by VCL, in epithelioid fibrous histiocytoma. In addition, we report four novel ALK partners in these tumors: DCTN1, ETV6, PPFIBP1, and SPECC1L. To our knowledge, this is the first documented account of ETV6-ALK and SPECC1L-ALK translocations occurring in neoplasms. It is presumed that additional ALK fusion partners will be identified in the future; furthermore, since other genes, such as ROS1, may substitute for ALK, it is likely some of the aforementioned genes may be capable of generating non-ALK partners in epithelioid fibrous histiocytoma and other tumors. These data provide additional support for the separation of epithelioid fibrous histiocytoma from other types of cutaneous fibrous histiocytoma, although we have no good alternative nomenclature, at least for now.
References
Jones EW, Cerio R, Smith NP . Epithelioid cell histiocytoma: a new entity. Br J Dermatol 1989;120:185–195.
Singh Gomez C, Calonje E, Fletcher CD . Epithelioid benign fibrous histiocytoma of skin: clinico-pathological analysis of 20 cases of a poorly known variant. Histopathology 1994;24:123–129.
Doyle LA, Fletcher CD . EMA positivity in epithelioid fibrous histiocytoma: a potential diagnostic pitfall. J Cutan Pathol 2011;38:697–703.
Doyle LA, Marino-Enriquez A, Fletcher CD et al. ALK rearrangement and overexpression in epithelioid fibrous histiocytoma. Mod Pathol 2015;28:904–912.
Jedrych J, Nikiforova M, Kennedy TF et al. Epithelioid cell histiocytoma of the skin with clonal ALK gene rearrangement resulting in VCL-ALK and SQSTM1-ALK gene fusions. Br J Dermatol 2015;172:1427–1429.
Debelenko LV, Raimondi SC, Daw N et al. Renal cell carcinoma with novel VCL-ALK fusion: new representative of ALK-associated tumor spectrum. Mod Pathol 2011;24:430–442.
Takeuchi K, Soda M, Togashi Y et al. Identification of a novel fusion, SQSTM1-ALK, in ALK-positive large B-cell lymphoma. Haematologica 2011;96:464–467.
Bridge JA, Kanamori M, Ma Z et al. Fusion of the ALK gene to the clathrin heavy chain gene, CLTC, in inflammatory myofibroblastic tumor. Am J Pathol 2001;159:411–415.
Morris SW, Kirstein MN, Valentine MB et al. Fusion of a kinase gene, ALK, to a nucleolar protein gene, NPM, in non-Hodgkin's lymphoma. Science 1994;263:1281–1284.
Pan Z, Hu S, Li M et al. ALK-positive large B-cell lymphoma: a clinicopathologic study of 26 cases with review of additional 108 cases in the literature. Am J Surg Pathol 2017;41:25–38.
Onciu M, Behm FG, Downing JR et al. ALK-positive plasmablastic B-cell lymphoma with expression of the NPM-ALK fusion transcript: report of 2 cases. Blood 2003;102:2642–2644.
Mano H . Non-solid oncogenes in solid tumors: EML4-ALK fusion genes in lung cancer. Cancer Sci 2008;99:2349–2355.
Marino-Enriquez A, Ou WB, Weldon CB et al. ALK rearrangement in sickle cell trait-associated renal medullary carcinoma. Genes Chromosomes Cancer 2011;50:146–153.
Chou A, Fraser S, Toon CW et al. A detailed clinicopathologic study of ALK-translocated papillary thyroid carcinoma. Am J Surg Pathol 2015;39:652–659.
Amin SM, Haugh AM, Lee CY et al. A comparison of morphologic and molecular features of BRAF, ALK, and NTRK1 fusion spitzoid neoplasms. Am J Surg Pathol 2017;41:491–498.
Chen X, Schulz-Trieglaff O, Shaw R et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics 2016;32:1220–1222.
Liu S, Tsai WH, Ding Y et al. Comprehensive evaluation of fusion transcript detection algorithms and a meta-caller to combine top performing methods in paired-end RNA-seq data. Nucleic Acids Res 2016;44:e47.
Duyster J, Bai RY, Morris SW . Translocations involving anaplastic lymphoma kinase (ALK). Oncogene 2001;20:5623–5637.
Chiarle R, Voena C, Ambrogio C et al. The anaplastic lymphoma kinase in the pathogenesis of cancer. Nat Rev Cancer 2008;8:11–23.
Hallberg B, Palmer RH . Mechanistic insight into ALK receptor tyrosine kinase in human cancer biology. Nat Rev Cancer 2013;13:685–700.
Soda M, Choi YL, Enomoto M et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 2007;448:561–566.
Antonescu CR, Suurmeijer AJ, Zhang L et al. Molecular characterization of inflammatory myofibroblastic tumors with frequent ALK and ROS1 gene fusions and rare novel RET rearrangement. Am J Surg Pathol 2015;39:957–967.
Sakamoto K, Nakasone H, Togashi Y et al. ALK-positive large B-cell lymphoma: identification of EML4-ALK and a review of the literature focusing on the ALK immunohistochemical staining pattern. Int J Hematol 2016;103:399–408.
Sugawara E, Togashi Y, Kuroda N et al. Identification of anaplastic lymphoma kinase fusions in renal cancer: large-scale immunohistochemical screening by the intercalated antibody-enhanced polymer method. Cancer 2012;118:4427–4436.
Yu W, Wang Y, Jiang Y et al. Genetic analysis and clinicopathological features of ALK-rearranged renal cell carcinoma in a large series of resected Chinese renal cell carcinoma patients and literature review. Histopathology 2017;71:53–62.
Lee JC, Li CF, Huang HY et al. ALK oncoproteins in atypical inflammatory myofibroblastic tumours: novel RRBP1-ALK fusions in epithelioid inflammatory myofibroblastic sarcoma. J Pathol 2017;241:316–323.
Lovly CM, Gupta A, Lipson D et al. Inflammatory myofibroblastic tumors harbor multiple potentially actionable kinase fusions. Cancer Discov 2014;4:889–895.
Rea SL, Majcher V, Searle MS et al. SQSTM1 mutations—bridging Paget disease of bone and ALS/FTLD. Exp Cell Res 2014;325:27–37.
Iyevleva AG, Raskin GA, Tiurin VI et al. Novel ALK fusion partners in lung cancer. Cancer Lett 2015;362:116–121.
Iyama K, Matsuse M, Mitsutake N et al. Identification of three novel fusion oncogenes, SQSTM1/NTRK3, AFAP1L2/RET, and PPFIBP2/RET, in thyroid cancers of young patients in Fukushima. Thyroid 2017;27:811–818.
Nakamura Y, Ito Y, Wakimoto N et al. A novel fusion of SQSTM1 and FGFR1 in a patient with acute myelomonocytic leukemia with t(5;8)(q35;p11) translocation. Blood Cancer J 2014;4:e265.
Bays JL, DeMali KA . Vinculin in cell-cell and cell-matrix adhesions. Cell Mol Life Sci 2017;74:2999–3009.
Han SP, Yap AS . An alpha-catenin deja vu. Nat Cell Biol 2013;15:238–239.
Maeda M, Holder E, Lowes B et al. Dilated cardiomyopathy associated with deficiency of the cytoskeletal protein metavinculin. Circulation 1997;95:17–20.
Kardon JR, Vale RD . Regulators of the cytoplasmic dynein motor. Nat Rev Mol Cell Biol 2009;10:854–865.
Busam KJ, Kutzner H, Cerroni L et al. Clinical and pathologic findings of Spitz nevi and atypical Spitz tumors with ALK fusions. Am J Surg Pathol 2014;38:925–933.
Shimada Y, Kohno T, Ueno H et al. An oncogenic ALK fusion and an RRAS mutation in KRAS mutation-negative pancreatic ductal adenocarcinoma. Oncologist 2017;22:158–164.
Subbiah V, McMahon C, Patel S et al. STUMP un"stumped": anti-tumor response to anaplastic lymphoma kinase (ALK) inhibitor based targeted therapy in uterine inflammatory myofibroblastic tumor with myxoid features harboring DCTN1-ALK fusion. J. Hematol Oncol 2015;8:66.
De Braekeleer E, Douet-Guilbert N, Morel F et al. ETV6 fusion genes in hematological malignancies: a review. Leuk Res 2012;36:945–961.
Zhang MY, Churpek JE, Keel SB et al. Germline ETV6 mutations in familial thrombocytopenia and hematologic malignancy. Nat Genet 2015;47:180–185.
Knezevich SR, McFadden DE, Tao W et al. A novel ETV6-NTRK3 gene fusion in congenital fibrosarcoma. Nat Genet 1998;18:184–187.
Rubin BP, Chen CJ, Morgan TW et al. Congenital mesoblastic nephroma t(12;15) is associated with ETV6-NTRK3 gene fusion: cytogenetic and molecular relationship to congenital (infantile) fibrosarcoma. Am J Pathol 1998;153:1451–1458.
Brenca M, Rossi S, Polano M et al. Transcriptome sequencing identifies ETV6-NTRK3 as a gene fusion involved in GIST. J Pathol 2016;238:543–549.
Davis JL, Lockwood CM, Albert CM et al. Infantile NTRK-associated mesenchymal tumors. Pediatr Dev Pathol [e-pub ahead of print, 1 January 2017].
Yamamoto H, Yoshida A, Taguchi K et al. ALK, ROS1 and NTRK3 gene rearrangements in inflammatory myofibroblastic tumours. Histopathology 2016;69:72–83.
Alassiri AH, Ali RH, Shen Y et al. ETV6-NTRK3 is expressed in a subset of ALK-negative inflammatory myofibroblastic tumors. Am J Surg Pathol 2016;40:1051–1061.
Tognon C, Knezevich SR, Huntsman D et al. Expression of the ETV6-NTRK3 gene fusion as a primary event in human secretory breast carcinoma. Cancer Cell 2002;2:367–376.
Andreasen S, Skalova A, Agaimy A et al. ETV6 gene rearrangements characterize a morphologically distinct subset of sinonasal low-grade non-intestinal-type adenocarcinoma: a novel translocation-associated carcinoma restricted to the sinonasal tract. Am J Surg Pathol 2017;41:1552–1560.
Yeh I, Tee MK, Botton T et al. NTRK3 kinase fusions in Spitz tumours. J Pathol 2016;240:282–290.
Leeman-Neill RJ, Kelly LM, Liu P et al. ETV6-NTRK3 is a common chromosomal rearrangement in radiation-associated thyroid cancer. Cancer 2014;120:799–807.
Marino-Enriquez A, Wang WL, Roy A et al. Epithelioid inflammatory myofibroblastic sarcoma: an aggressive intra-abdominal variant of inflammatory myofibroblastic tumor with nuclear membrane or perinuclear ALK. Am J Surg Pathol 2011;35:135–144.
Wei Z, Zheng S, Spangler SA et al. Liprin-mediated large signaling complex organization revealed by the liprin-alpha/CASK and liprin-alpha/liprin-beta complex structures. Mol Cell 2011;43:586–598.
Takeuchi K, Soda M, Togashi Y et al. Pulmonary inflammatory myofibroblastic tumor expressing a novel fusion, PPFIBP1-ALK: reappraisal of anti-ALK immunohistochemistry as a tool for novel ALK fusion identification. Clin Cancer Res 2011;17:3341–3348.
Wiesner T, He J, Yelensky R et al. Kinase fusions are frequent in Spitz tumours and spitzoid melanomas. Nat Commun 2014;5:3116.
Saadi I, Alkuraya FS, Gisselbrecht SS et al. Deficiency of the cytoskeletal protein SPECC1L leads to oblique facial clefting. Am J Hum Genet 2011;89:44–55.
Gfrerer L, Shubinets V, Hoyos T et al. Functional analysis of SPECC1L in craniofacial development and oblique facial cleft pathogenesis. Plast Reconstr Surg 2014;134:748–759.
Acknowledgements
The authors are indebted to Dr Martin E Blackstein for providing funding support for this study. In addition, they are grateful to Mr Andrew Wong for bioinformatic assistance, and Ms Eva Agro and Ms Sharon Crafter for providing molecular support.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Supplementary information
Rights and permissions
About this article
Cite this article
Dickson, B., Swanson, D., Charames, G. et al. Epithelioid fibrous histiocytoma: molecular characterization of ALK fusion partners in 23 cases. Mod Pathol 31, 753–762 (2018). https://doi.org/10.1038/modpathol.2017.191
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2017.191
This article is cited by
-
Detection of sarcoma fusions by a next-generation sequencing based–ligation-dependent multiplex RT-PCR assay
Modern Pathology (2022)
-
Epithelioid inflammatory myofibroblastic sarcoma with VCL–ALK fusion of central nervous system: case report and brief review of the literature
Brain Tumor Pathology (2022)
-
Epithelioid fibrous histiocytoma: three diagnostically challenging cases with novel ALK gene fusions, unusual storiform growth pattern, and a prominent spindled morphology
Virchows Archiv (2022)
-
Broadening the spectrum of NTRK rearranged mesenchymal tumors and usefulness of pan-TRK immunohistochemistry for identification of NTRK fusions
Modern Pathology (2021)
-
Superficial ALK-rearranged myxoid spindle cell neoplasm: a cutaneous soft tissue tumor with distinctive morphology and immunophenotypic profile
Modern Pathology (2021)