Abstract
In mammalian development, dynamic epigenetic reprogramming occurs in pre-implantation embryos and primordial germ cells and plays a critical role in conferring pluripotency on embryonic cells. Pluripotent stem cells, such as embryonic stem cells and induced pluripotent stem cells, have been derived and maintained in vitro under culture conditions that include stimulators and inhibitors of extrinsic signaling. Recent advances in stem cell cultivation have opened the possibility of capturing naive pluripotency, which is reminiscent of the pluripotency of inner cell mass cells, in vitro. However, emerging evidence has revealed complexity of epigenetic regulation in pluripotent stem cells in vitro that reflects the developmental stage, gender, and species. In this review, we describe the developmental potential and epigenetic regulation of pluripotent stem cells in rodents and humans in vitro and discuss unsolved issues in developing strategies to capture in vivo pluripotency in vitro.
Similar content being viewed by others
Main
In mammals, cell fate specification is governed by highly coordinated and sequential gene regulatory mechanisms. Transcription factors (TFs) and epigenetic modifications, including DNA methylation and histone modifications, play a central role in these programs.1, 2, 3 Once cells differentiate into a particular cell lineage, the committed cell fate is stably maintained by epigenetic control of the gene regulation.1, 4, 5 However, during the mammalian life cycle, epigenetic reprogramming occurs in primordial germ cells (PGCs) and pre-implantation embryos, which resets the cellular commitment and confers totipotency on the zygote3, 6, 7, 8 (Figure 1a). In PGCs, epigenetic memory, particularly DNA methylation, is erased before establishing novel epigenetic regulation during gametogenesis, which includes genomic imprinting with sperm-/oocyte-specific de novo DNA methylation9, 10 (Figures 1a and b). Further epigenetic reorganization occurs after fertilization, which is essential for the acquisition of totipotency and subsequent pluripotency.
Epigenetic reprogramming and programming in mouse development. (a) DNA methylation levels in mammalian development. Global DNA demethylation occurs in pre-implantation embryos and PGCs. DNA methylation in the paternal genome reaches the level in the maternal genome after fertilization, and then both genomes are further demethylated until a zygote develops into blastocyst. After blastocyst development, post-implantation epiblast gains de novo global methylation during gastrulation and differentiates into three germ layers. PGCs emerged from the epiblast also undergo global demethylation. Germ cells (sperm or oocyte) gain de novo methylation during their maturation in a sex-dependent manner to establish genomic imprints. E indicates the embryonic day. (b) Genomic imprints in mammalian development. Imprints are established in germline (oocyte or sperm) in a sex-dependent manner (blue: paternal imprints, red: maternal imprints). The established imprints are strictly maintained in ICM, embryo, and somatic cells throughout life. Parental imprints are erased only in PGCs to re-establishment imprints in the next generation.
Pluripotency is defined as the potential of a cell to give rise to all three germ layers, including germ cells, and is linked with unique epigenetic control of gene regulation. Indeed, after epigenetic reprogramming, inner cell mass (ICM) of blastocysts as well as PGCs display global DNA hypomethylation and characteristic transcriptional profiling6, 7, 11, 12 (Figure 1a). Pre-implantation embryos undergo dynamic reorganization of the epigenetic regulation. Importantly, however, DNA methylation at imprinting control regions (ICRs) that are inherited from either the oocyte or sperm as ‘parental memory’ are resistant to demethylation in ICM cells (Figure 1b). The maintenance of genomic imprints in ICM, embryonic, and adult cells is thought to be important, since aberrant imprints are often linked with various diseases including behavior anomalies and tumorigenesis.13, 14, 15 After ICM development, post-implantation epiblasts undergo epigenetic programming including genome-wide de novo DNA methylation and differentiate into all three germ layers5, 16 (Figure 1a).
Recapitulation of in vivo pluripotency in vitro culture has been a major goal of developmental biologists for decades. There exist several types of pluripotent stem cells. In mice, embryonic stem cells (ES cells)17, 18 and epiblast stem cells (EpiS cells)19, 20 are derived from ICM and post-implantation epiblast, respectively, and both are typical pluripotent cell lines isolated from early embryos (Figure 2). Although both ES cells and EpiS cells have pluripotency, they exhibit distinct transcriptional and epigenetic signatures.21, 22 Intriguingly, human ES cells are transcriptionally and epigenetically distinct from mouse ES cells and resemble mouse EpiS cells, despite being derived from developing pre-implantation blastocysts, suggesting different regulatory mechanisms of pluripotency between rodents and humans.21 The derivation of induced pluripotent stem cells (iPS cells) has enabled the creation of pluripotent stem cells from somatic cells without the need for embryos. Because ES/iPS cells have the ability of self-renewal and pluripotency, they could serve clinical applications such as regenerative medicine and drug discovery.23 For these purposes, however, it is important to understand the common and distinctive molecular and functional features of different pluripotent stem cells. In this review, we discuss current understandings and unresolved key issues in the developmental potential and epigenetic regulation of pluripotent states in vivo and in vitro.
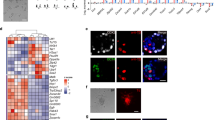
Effects of genomic imprints on stem cell potential. (a) The expression of Gtl2, an imprinted gene, is frequently silenced in iPS cells (OSKM; Oct3/4, Sox2, Klf4, and c-Myc and OSKM+AA; Oct3/4, Sox2, Klf4, and c-Myc with ascorbic acid (AA)). Gtl2-OFF iPS cell clones tend to poorly contribute to chimeric mice and fail to produce all-iPS-cell mice by 4n complementation. AA treatment prevents the imprinted region from aberrant methylation and yields iPS cells with Gtl2 expression. Gtl2-ON iPS cell clones are able to efficiently contribute to chimeras and produce all-iPS-cell mice. (b) Gtl2 is expressed only from the maternal allele, which is regulated by paternal allele-specific DNA methylation at the Dlk1-Dio3 imprinted cluster. The imprinted cluster is often aberrantly hypermethylated in Gtl2-OFF iPS cells, whereas it is methylated only at the paternal allele in Gtl2-ON iPS cells.
Epigenetic reprogramming and programming in mouse early development in vivo
DNA methylation patterns are dynamically altered during early embryogenesis in mice.5, 7 Paternal and maternal genomes are demethylated asymmetrically after fertilization.24 The paternal genome is immediately demethylated in the zygote, while the maternal genome undergoes demethylation during the cleavage stage. Interestingly, the oocyte already exhibits DNA hypomethylation relative to sperm, particularly at transposable elements.7 However, upon fertilization, sperm methylation rapidly decreases to levels comparable of the oocyte, indicating that the oocyte methylation pattern is a better predictor of epigenetic reprogramming in the zygote. Continuous genome-wide DNA demethylation occurs in pre-implantation embryo until the ICM stage, when the DNA methylation status reaches to a lower level7, 12 (Figure 1a). Mechanistically, both paternal and maternal genomes undergo passive demethylation during the pre-implantation stage. In agreement with this property, DNA methyltransferases are mainly localized outside of the nuclei from the two-cell embryo to blastocyst stage.25 Despite dynamic epigenetic reprogramming in pre-implantation embryos, unique sequences in the genome are resistant to DNA demethylation. Such representative genetic elements include ICRs and intracisternal A particle transposons.26 After specification of the ICM, genome-wide remethylation in the epiblast is essential for gastrulation and proper cellular differentiation into the three germ layers including germ line5, 27 (Figure 1a). X chromosome inactivation is one of the essential epigenetic phenomena. In female pre-implantation mouse embryos, paternal X chromosome is reactivated only in ICM and not in extraembryonic cells. Then, one X chromosome is randomly inactivated for dosage compensation after exit from the ICM state.28, 29, 30 This dynamic epigenetic regulation plays an important role in normal mammalian development.
Establishment and maintenance of genomic imprinting in mice
Because imprinted genes often directly regulate fetal growth, genomic imprinting is an essential epigenetic phenomenon for mammalian development. Mammalian cells are normally diploid, with the two sets of genes inherited from both parents. Most genes are expressed from both alleles, but a subset is expressed from only one. Monoallelic expression is regulated by ICR methylation that is specifically established during germ cell differentiation (oocyte or sperm) (Figure 1b). Mechanistically, the establishment of imprints in oocyte/sperm is dependent on the de novo methyltransferase Dnmt3a and stimulatory factor Dnmt3l. In fact, genetic deletion of either fails to establish imprints in mice.31, 32, 33, 34
Parent-specific ICR methylation is maintained in embryos and adults by Dnmt1 (Figure 1b). The functional significance of the original parental expression for mammalian development can be explained by the lethal phenotype of parthenogenetic and androgenetic mouse embryos as well as uniparental disomies.35, 36 It was previously demonstrated that Dnmt1 knockout causes embryonic lethal and deletion of Dnmt1 in mouse ES cells results in global DNA hypomethylation including imprinted loci.13, 37, 38 Notably, once imprints are erased in mouse ES cells, they remain unmethylated even after differentiation, indicating that somatic cells lack the ability to re-establish imprints.13, 38 Given that aberrant imprinting persists in somatic cells throughout the lifetime, the epigenetic stability of imprints should contribute to the developmental potential of pluripotent stem cells.
Derivation of mouse pluripotent stem cells from somatic cells
Cellular specification and commitment are orchestrated by regulatory mechanisms that mediate spatiotemporal control of gene expression. Crosstalk between TFs and epigenetic regulations governs gene regulatory programs that drive precise control of gene expression patterns during development. Given that the epigenetic control of gene regulatory programs is a largely irreversible process, it was thought that somatic cells are unlikely reprogrammable to the pluripotent state once cells differentiate. However, that idea has since been refuted, as it was shown that somatic differentiated cells are reprogrammable to ES cell-like pluripotent cells by somatic cell nuclear transfer39, 40 and the ectopic expression of four TFs, Oct3/4, Sox2, Klf4, and c-Myc (hereafter referred to as OSKM).41 The iPS cells were successfully generated from mouse fibroblasts in 200641 and from human fibroblasts in 2007.42, 43 Since then, many laboratories have succeeded in establishing iPS cells with various methods (eg, virus-based, integration-free, and piggyBac transposon system, among others).44, 45, 46, 47, 48, 49, 50 Furthermore, a recent study showed that Sall4, Nanog, Esrrb, and Lin28 could replace OSKM to induce somatic cell reprogramming into iPS cells in mice.51 Consistent with reprogramming to the pluripotent state, the epigenetic signatures of iPS cells are similar with those of ES cells. Moreover, mouse iPS cells are competent for tetraploid complementation assay,52 demonstrating that iPS cells are indeed pluripotent and are functionally equivalent to ES cells.53 Collectively, either ex vivo culture of early embryos or somatic cell reprogramming can obtain pluripotent cells in vitro.
Aberrant genomic imprinting in mouse pluripotent stem cells
Aberrant imprints have been implicated in various diseases. Notably, mouse ES cell lines have been shown to occasionally exhibit aberrant ICR methylation.54 Previous studies also reported that several imprinted genes within the Dlk1-Dio3 cluster are aberrantly silenced in most mouse iPS cell lines.55 The aberrant silencing is accompanied by increased methylation at these ICRs (Figures 2a and b). These epigenetic alterations are not observed in most ES cells but only some iPS cell lines, suggesting that iPS cells exhibit considerable epigenetic variation when compared with ES cells. Together, it appears that genomic imprinting is vulnerable in mouse pluripotent stem cells in vitro. Given that genomic imprints are essential epigenetic regulation mechanisms in mammalian development, a key issue is whether the imprint status in stem cells affects stem cell functionality. Notably, mouse iPS cells harboring aberrant imprints at the Dlk1-Dio3 cluster poorly contribute to chimeras and fail to produce all-iPS-cell mice by tetraploid embryo (4n) complementation, which is one of the most stringent tests for developmental assays (Figure 2a). Of note, histone deacetylase inhibitor and ascorbic acid reactivated the silenced loci in iPS cells and eventually rescued the defect permitting all-iPS-cell mice.55, 56 These findings indicate that the imprint status in pluripotent states significantly affects the developmental potential of iPS cells (Figure 2a). Collectively, the imprint status is key epigenetic regulation for the propagation of ICM-like naive pluripotency in vitro.
Naive and primed mouse pluripotent stem cells in ex vivo culture
Since ICM cells can contribute to all cell lineages of the body, they are functionally described as having ‘naive pluripotency.’22 Mouse naive ES cells derived from ICM of blastocysts have historically been maintained in serum and leukemia inhibitory factor (LIF) on feeder cells, and they indefinitely proliferate on culture dish and differentiate into all three germ layers including germ cell line when they are injected into blastocysts57, 58, 59 (Figure 3). After post implantation, mouse epiblast cells form an egg cylinder structure at around E5.5–E6.5. EpiS cells are derived from post-implantation epiblast under media containing basic fibroblast growth factor and activin and defined as having ‘primed pluripotency’19, 20, 22 (Figure 3). EpiS cells can differentiate into various cell types in vitro and form teratomas, but they lack an ability to contribute to chimeras when injected into blastocysts.19 While the maintenance of mouse ES cells is dependent on the LIF/Stat3 signaling pathway, the maintenance of EpiS cells is dependent on the FGF/ERK pathway.
Transcriptional and epigenetic signatures in naive and primed mouse pluripotent stem cells. In mice, ES cells derived from ICM of blastocysts are defined as having naive pluripotency, while EpiS cells derived from post-implantation epiblasts are defined as having primed pluripotency. The X chromosome status is XaXa in naive state and XaXi in primed state (Xa; active X chromosome and Xi; inactive X chromosome). Serum/LIF-cultured ES cells display global DNA hypermethylation and heterogeneous expression patterns of naive pluripotent marker genes. 2i/LIF media enables ES cells to maintain homogenous ground-state pluripotency and global DNA hypomethylation.
Consistent with the in vivo epigenetic property of female developing embryos, mouse ES cells have two active X chromosomes (XaXa), whereas one copy of X chromosomes is inactive (XaXi) in EpiS cells (Figure 3). Transcriptionally, EpiS cells express core pluripotent marker genes including Nanog and Oct3/4. However, Klf4 and Zfp42 (Rex1) are downregulated in primed EpiS cells compared to naive ES cells.60 Thus, there are marked functional and molecular differences between the naive and primed states.
Capturing ground state in vitro by modulating extrinsic signaling pathways in mice
Although mouse pluripotent stem cells maintained in vitro are able to contribute to all somatic cell lineages, they contain heterogeneous populations in terms of morphology and transcriptional patterns of naive marker genes, suggesting that ES/iPS cells in conventional culture condition fluctuate between the naive ICM-like state and primed epiblast-like state (Figure 3). Moreover, ES/iPS cells exhibit global DNA hypermethylation, whereas ICM cells exhibit global hypomethylation, suggesting that ES/iPS cells in vitro do not faithfully capture the naive ICM-like state22, 61 (Figure 3). In 2008, Smith and colleagues discovered key signaling pathways to overcome such metastable characteristics.62 Based on the fact that Fgf4−/− ES cells are compromised in their differentiation to neural and mesodermal lineages,63 the FGF/ERK signal was identified as a key upstream pathway for cellular differentiation. Another key pathway is Wnt signaling, which enhances the self-renewing activity of ES cells.64 Notably, dual inhibition of the FGF/ERK and GSK3 (MEK1/2 inhibitor; PD0325901 and Gsk3β inhibitor; CHIR99021), respectively (2i), makes it possible to propagate ground-state mouse ES cells by shielding cells from differentiation and reinforcing the core naive pluripotency circuit65, 66 (Figure 3). Consistent with the naive ICM-like state, exposing ES cells to 2i/LIF induces global DNA hypomethylation to an extent similar to ICM cells in vivo67, 68, 69, 70 (Figure 3). Moreover, 2i/LIF culture enhances the derivation of ES cells even from non-permissive mouse strains, which are refractory to ES cell propagation, and other species such as rats.71, 72, 73 Collectively, the 2i/LIF culture system enhances stem cell propagation while reinforcing transcriptional and epigenetic properties of the ICM-like naive state. However, it has not been fully elucidated whether genetic and epigenetic stabilities of 2i-treated ES cells are stable. Recent studies demonstrated that derivation of ES cells directly from blastocysts in 2i/LIF and prolonged culture of ES cells in 2i/LIF result in loss of DNA methylation at imprinted loci74, 75 (Figure 4). Surprisingly, such imprint eroded ES cells compromise autonomous developmental potential by tetraploid embryo complementation and nuclear transfer74, 75 (Figure 4). Mechanistically, the inhibition of MEK1/2 is responsible for these opposing effects. In addition, replacement of the MEK1/2 inhibitor with an Src inhibitor (a2i includes Src inhibitor; CGP77675 and CHIR99021) preserves genetic and epigenetic stability as well as autonomous developmental potential74, 75 (Figure 4). Given that many laboratories have implemented 2i culture condition as standard practice since the discovery of 2i, these findings should be taken into account for future experiments using 2i-treated ES cells.
2i affects genetic and epigenetic stability. Mouse ES cells directly derived from blastocyst in 2i and exposed for a prolonged period in 2i display genetic and epigenetic instability (eg, loss of imprints and karyotypic abnormality), which affects full-term autonomous developmental potential by tetraploid embryo complementation. a2i condition (including Src inhibitor and Gsk3β inhibitor) preserves genomic imprints, chromosomal stability, and developmental potential.
Epigenetic and genetic instability in female mouse es cells
Gender differences are thought to be another important aspect of mouse naive ES/iPS cells in vitro. Female ICM cells retain XaXa only for a transient period in vivo, while female ES/iPS cells sustain XaXa during cultivation in vitro. Recent accumulating studies showed that female mouse ES cells unexpectedly display global DNA hypomethylation including reduced methylation at imprinted loci.76 This property is attributable to XaXa, since XO cells and XY cells exhibit similarly higher levels of DNA methylation.76, 77 Mechanistically, the expression levels of Dnmt3a and Dnmt3b, but not Dnmt1, are markedly lower in XX cells than in XO and XY cells,76, 77 suggesting that lower activity of de novo methyltransferases due to XaXa is responsible for the global hypomethylation. Indeed, ectopic expression of Dnmt3a or Dnm3b can rescue the reduced methylation levels at particular regions in female ES cells.76 More recent studies showed that female mouse ES cells retaining XaXa displayed lower protein level of Uhrf1 than male ES cells, although mRNA level of Uhrf1 is comparable between male and female.74, 78 Moreover, reduced Uhrf1 protein level is linked with XaXa state since XO ES cells and XX MEFs exhibiting XaXi maintained Uhrf1 protein level.74, 78 These results suggest that female-specific hypomethylation is in part caused by impaired maintenance of DNA methylation due to two active X chromosomes. It is also interesting that female ES cells tend to lose all or part of one X chromosome during their propagation, while male ES cells occasionally lose Y chromosome, indicating that female ES cells are genetically unstable.76 Collectively, female mouse ES cells often display epigenetic and genetic variations that are associated with XaXa. Considering persistent XaXa in vitro, such variations should be considered for capturing naive ICM cells in vitro.
Epigenetic properties in human naive pluripotent stem cells
Human ES/iPS cells display primed pluripotency features, which correspond to an advanced stage of pluripotent cells that resembles post-implantation epiblast. A number of studies have sought to acquire naive pluripotency in human cells in vitro.79, 80, 81, 82, 83 Based on recent transcriptome and epigenetic analyses, it seems that two methods, (i) transgene (NANOG and KLF2)-dependent system in conjunction with 2i/LIF (t2i/L+PKCi) media82, 83 and (ii) chemical conditions with 5i/L/A or 4i/L/A,80 succeeded the transition of primed pluripotent stem cells to the naive state84, 85 (Figure 5). Naive human ES cells were similarly established directly from human blastocysts.83 Consistent with ICM-like pluripotency, human ES cells in these conditions display global DNA hypomethylation and the gene expression patterns of naive markers and transposons, which are reminiscent of pre-implantation human embryos84, 85, 86 (Figure 5). Of particularly note, inactive X chromosome becomes active in naive-like female cells in vitro, suggesting that reactivation of X chromosome occurs during naive conversion84, 87, 88 (Figure 5). Therefore, it is expected that human naive cells serve as a powerful tool for studying human early embryogenesis and human diseases.
Molecular dynamics during naive conversion of human pluripotent stem cells in vitro. Recent studies revealed that human naive-like ES cells can be generated from primed ES cells or directly from blastocysts. Human naive-like ES cells acquire transcriptional profiling reminiscent of ICM cells and display global DNA hypomethylation. Notably, erosion of X chromosome (Xe)89, 90 in primed cells is canceled by naive conversion. However, genomic imprints are lost in human naive-like cells and non-random X chromosome inactivation occurs after differentiation, both of which are not observed in normal early development in humans. Collectively, current human naive-like cells do not fully recapitulate human naive pluripotency in vivo.
However, more recent studies demonstrated that naive human ES cells tend to exhibit non-random X chromosome inactivation upon their differentiation87 (Figure 5). This phenomenon does not mimic human development in vivo. Similarly, several studies reported that ICR methylation in human naive cells was markedly decreased although the original primed pluripotent stem cells retained the methylation84, 86 (Figure 5). Such aberrant imprinting in naive state was inherited even after re-priming, which is consistent with the fact that cells outside of germline lack the ability to re-establish imprints in mice. Taken together, current human naive cells fail to faithfully recapitulate ICM cells in vivo in terms of epigenetic aspects. Considering the critical role of genomic imprints in developmental potential, there remain notable differences between in vitro naive cells and in vivo ICM cells in humans.
Perspective
Pre-implantation embryos undergo dynamic epigenetic reprogramming, which is essential for the mammalian life cycle. Proper reprogramming confers pluripotency on early embryos. Pluripotent stem cells in vitro capture many aspects of pluripotency in vivo and therefore provide a powerful experimental platform to study early embryogenesis. Accumulating evidence has suggested that 2i/LIF culture system maintains transcriptional and epigenetic signatures reminiscent of naive ICM-like cells in mice.65 Indeed, 2i/LIF-cultured mouse ES cells display a number of shared characteristics with ICM cells.66, 67, 68 Moreover, previous studies have provided insightful clues for naive transition of primed human pluripotent stem cells using culture conditions containing 2i. However, recent studies indicated that MEK1/2 inhibitor may cause genetic and epigenetic instability of mouse ES cells, which is associated with impaired developmental potential. Indeed, current human naive-like cells harbor karyotypic abnormalities and epigenetic abnormalities, such as a loss of imprints and distinct X chromosome regulation. Notably, a2i culture condition, in which a MEK1/2 inhibitor is replaced with an Src inhibitor, can be used for mouse ES cell maintenance. Moreover, a2i-cultured ES cells maintain genetic and epigenetic stability as well as autonomous developmental potential. These findings may help for generating human naive pluripotent state, which retains genetic and epigenetic stability. Given that pluripotent stem cells offer hope not only for understanding of human embryogenesis, but also for cell transplantation therapy as well as drug discovery, it will be necessary to integrate the complexity of epigenetic regulation in pluripotent stem cells in vitro into standard approaches that faithfully capture in vivo ICM cells on culture dish.
References
Surani MA, Hayashi K, Hajkova P . Genetic and epigenetic regulators of pluripotency. Cell 2007;128:747–762.
Apostolou E, Hochedlinger K . Chromatin dynamics during cellular reprogramming. Nature 2013;502:462–471.
Lee HJ, Hore TA, Reik W . Reprogramming the methylome: erasing memory and creating diversity. Cell Stem Cell 2014;14:710–719.
Reik W . Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 2007;447:425–432.
Smith ZD, Meissner A . DNA methylation: roles in mammalian development. Nat Rev Genet 2013;14:204–220.
Seisenberger S, Andrews S, Krueger F et al. The dynamics of genome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol Cell 2012;48:849–862.
Smith ZD, Chan MM, Mikkelsen TS et al. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 2012;484:339–344.
Popp C, Dean W, Feng S et al. Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature 2010;463:1101–1105.
Hajkova P, Ancelin K, Waldmann T et al. Chromatin dynamics during epigenetic reprogramming in the mouse germ line. Nature 2008;452:877–881.
Hackett JA, Sengupta R, Zylicz JJ et al. Germline DNA demethylation dynamics and imprint erasure through 5-hydroxymethylcytosine. Science 2013;339:448–452.
Boroviak T, Loos R, Bertone P et al. The ability of inner-cell-mass cells to self-renew as embryonic stem cells is acquired following epiblast specification. Nat Cell Biol 2014;16:516–528.
Kobayashi H, Sakurai T, Imai M et al. Contribution of intragenic DNA methylation in mouse gametic DNA methylomes to establish oocyte-specific heritable marks. PLoS Genet 2012;8:e1002440.
Holm TM, Jackson-Grusby L, Brambrink T et al. Global loss of imprinting leads to widespread tumorigenesis in adult mice. Cancer Cell 2005;8:275–285.
Ohnishi K, Semi K, Yamamoto T et al. Premature termination of reprogramming in vivo leads to cancer development through altered epigenetic regulation. Cell 2014;156:663–677.
Peters J . The role of genomic imprinting in biology and disease: an expanding view. Nat Rev Genet 2014;15:517–530.
Hon GC, Rajagopal N, Shen Y et al. Epigenetic memory at embryonic enhancers identified in DNA methylation maps from adult mouse tissues. Nat Genet 2013;45:1198–1206.
Martin GR . Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc Natl Acad Sci USA 1981;78:7634–7638.
Evans MJ, Kaufman MH . Establishment in culture of pluripotential cells from mouse embryos. Nature 1981;292:154–156.
Tesar PJ, Chenoweth JG, Brook FA et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 2007;448:196–199.
Brons IG, Smithers LE, Trotter MW et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 2007;448:191–195.
Weinberger L, Ayyash M, Novershtern N et al. Dynamic stem cell states: naive to primed pluripotency in rodents and humans. Nat Rev Mol Cell Biol 2016;17:155–169.
Nichols J, Smith A . Naive and primed pluripotent states. Cell Stem Cell 2009;4:487–492.
Shi Y, Inoue H, Wu JC et al. Induced pluripotent stem cell technology: a decade of progress. Nat Rev Drug Discov 2016;16:115–130.
Nakamura T, Arai Y, Umehara H et al. PGC7/Stella protects against DNA demethylation in early embryogenesis. Nat Cell Biol 2007;9:64–71.
Ko YG, Nishino K, Hattori N et al. Stage-by-stage change in DNA methylation status of Dnmt1 locus during mouse early development. J Biol Chem 2005;280:9627–9634.
Bartolomei MS, Ferguson-Smith AC . Mammalian genomic imprinting. Cold Spring Harb Perspect Biol 2011;3:a002592.
Smallwood SA, Tomizawa S, Krueger F et al. Dynamic CpG island methylation landscape in oocytes and preimplantation embryos. Nat Genet 2011;43:811–814.
Heard E . Recent advances in X-chromosome inactivation. Curr Opin Cell Biol 2004;16:247–255.
Pasque V, Plath K . X chromosome reactivation in reprogramming and in development. Curr Opin Cell Biol 2015;37:75–83.
Schulz EG, Heard E . Role and control of X chromosome dosage in mammalian development. Curr Opin Genet Dev 2013;23:109–115.
Bourc'his D, Xu GL, Lin CS et al. Dnmt3L and the establishment of maternal genomic imprints. Science 2001;294:2536–2539.
Kaneda M, Okano M, Hata K et al. Essential role for de novo DNA methyltransferase Dnmt3a in paternal and maternal imprinting. Nature 2004;429:900–903.
Hata K, Okano M, Lei H et al. Dnmt3L cooperates with the Dnmt3 family of de novo DNA methyltransferases to establish maternal imprints in mice. Development 2002;129:1983–1993.
Okano M, Bell DW, Haber DA et al. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 1999;99:247–257.
Surani MA, Barton SC, Norris ML . Development of reconstituted mouse eggs suggests imprinting of the genome during gametogenesis. Nature 1984;308:548–550.
McGrath J, Solter D . Completion of mouse embryogenesis requires both the maternal and paternal genomes. Cell 1984;37:179–183.
Li E, Bestor TH, Jaenisch R . Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 1992;69:915–926.
Tucker KL, Beard C, Dausmann J et al. Germ-line passage is required for establishment of methylation and expression patterns of imprinted but not of nonimprinted genes. Genes Dev 1996;10:1008–1020.
Wilmut I, Schnieke AE, McWhir J et al. Viable offspring derived from fetal and adult mammalian cells. Nature 1997;385:810–813.
Wakayama T, Perry AC, Zuccotti M et al. Full-term development of mice from enucleated oocytes injected with cumulus cell nuclei. Nature 1998;394:369–374.
Takahashi K, Yamanaka S . Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006;126:663–676.
Takahashi K, Tanabe K, Ohnuki M et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 2007;131:861–872.
Yu J, Vodyanik MA, Smuga-Otto K et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 2007;318:1917–1920.
Wernig M, Meissner A, Foreman R et al. In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature 2007;448:318–324.
Woltjen K, Michael IP, Mohseni P et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 2009;458:766–770.
Kaji K, Norrby K, Paca A et al. Virus-free induction of pluripotency and subsequent excision of reprogramming factors. Nature 2009;458:771–775.
Yusa K, Rad R, Takeda J et al. Generation of transgene-free induced pluripotent mouse stem cells by the piggyBac transposon. Nat Methods 2009;6:363–369.
Yu J, Hu K, Smuga-Otto K et al. Human induced pluripotent stem cells free of vector and transgene sequences. Science 2009;324:797–801.
Stadtfeld M, Nagaya M, Utikal J et al. Induced pluripotent stem cells generated without viral integration. Science 2008;322:945–949.
Okita K, Nakagawa M, Hyenjong H et al. Generation of mouse induced pluripotent stem cells without viral vectors. Science 2008;322:949–953.
Buganim Y, Markoulaki S, van Wietmarschen N et al. The developmental potential of iPSCs is greatly influenced by reprogramming factor selection. Cell Stem Cell 2014;15:295–309.
Zhao XY, Li W, Lv Z et al. iPS cells produce viable mice through tetraploid complementation. Nature 2009;461:86–90.
Choi J, Lee S, Mallard W et al. A comparison of genetically matched cell lines reveals the equivalence of human iPSCs and ESCs. Nat Biotechnol 2015;33:1173–1181.
Humpherys D, Eggan K, Akutsu H et al. Epigenetic instability in ES cells and cloned mice. Science 2001;293:95–97.
Stadtfeld M, Apostolou E, Akutsu H et al. Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. Nature 2010;465:175–181.
Stadtfeld M, Apostolou E, Ferrari F et al. Ascorbic acid prevents loss of Dlk1-Dio3 imprinting and facilitates generation of all-iPS cell mice from terminally differentiated B cells. Nat Genet 2012;44:S1–S2.
Martin GR, Evans MJ . Differentiation of clonal lines of teratocarcinoma cells: formation of embryoid bodies in vitro. Proc Natl Acad Sci USA 1975;72:1441–1445.
Smith AG, Heath JK, Donaldson DD et al. Inhibition of pluripotential embryonic stem cell differentiation by purified polypeptides. Nature 1988;336:688–690.
Williams RL, Hilton DJ, Pease S et al. Myeloid leukaemia inhibitory factor maintains the developmental potential of embryonic stem cells. Nature 1988;336:684–687.
Festuccia N, Osorno R, Halbritter F et al. Esrrb is a direct Nanog target gene that can substitute for Nanog function in pluripotent cells. Cell Stem Cell 2012;11:477–490.
Hackett JA, Surani MA . Regulatory principles of pluripotency: from the ground state up. Cell Stem Cell 2014;15:416–430.
Martello G, Smith A . The nature of embryonic stem cells. Annu Rev Cell Dev Biol 2014;30:647–675.
Kunath T, Saba-El-Leil MK, Almousailleakh M et al. stimulation of the Erk1/2 signalling cascade triggers transition of pluripotent embryonic stem cells from self-renewal to lineage commitment. Development 2007;134:2895–2902.
Sato N, Meijer L, Skaltsounis L et al. Maintenance of pluripotency in human and mouse embryonic stem cells through activation of Wnt signaling by a pharmacological GSK-3-specific inhibitor. Nat Med 2004;10:55–63.
Ying QL, Wray J, Nichols J et al. The ground state of embryonic stem cell self-renewal. Nature 2008;453:519–523.
Marks H, Kalkan T, Menafra R et al. The transcriptional and epigenomic foundations of ground state pluripotency. Cell 2012;149:590–604.
Ficz G, Hore TA, Santos F et al. FGF signaling inhibition in ESCs drives rapid genome-wide demethylation to the epigenetic ground state of pluripotency. Cell Stem Cell 2013;13:351–359.
Habibi E, Brinkman AB, Arand J et al. Whole-genome bisulfite sequencing of two distinct interconvertible DNA methylomes of mouse embryonic stem cells. Cell Stem Cell 2013;13:360–369.
Hackett JA, Dietmann S, Murakami K et al. Synergistic mechanisms of DNA demethylation during transition to ground-state pluripotency. Stem Cell Rep 2013;1:518–531.
Leitch HG, McEwen KR, Turp A et al. Naive pluripotency is associated with global DNA hypomethylation. Nat Struct Mol Biol 2013;20:311–316.
Hanna J, Markoulaki S, Mitalipova M et al. Metastable pluripotent states in NOD-mouse-derived ESCs. Cell Stem Cell 2009;4:513–524.
Buehr M, Meek S, Blair K et al. Capture of authentic embryonic stem cells from rat blastocysts. Cell 2008;135:1287–1298.
Czechanski A, Byers C, Greenstein I et al. Derivation and characterization of mouse embryonic stem cells from permissive and nonpermissive strains. Nat Protoc 2014;9:559–574.
Yagi M, Kishigami S, Tanaka A et al. Derivation of ground-state female ES cells maintaining gamete-derived DNA methylation. Nature 2017;548:224–227.
Choi J, Huebner AJ, Clement K et al. Prolonged Mek1/2 suppression impairs the developmental potential of embryonic stem cells. Nature 2017;548:219–223.
Zvetkova I, Apedaile A, Ramsahoye B et al. Global hypomethylation of the genome in XX embryonic stem cells. Nat Genet 2005;37:1274–1279.
Schulz EG, Meisig J, Nakamura T et al. The two active X chromosomes in female ESCs block exit from the pluripotent state by modulating the ESC signaling network. Cell Stem Cell 2014;14:203–216.
Choi J, Clement K, Huebner AJ et al. DUSP9 modulates DNA hypomethylation in female mouse pluripotent stem cells. Cell Stem Cell 2017;20:706–719.
Gafni O, Weinberger L, Mansour AA et al. Derivation of novel human ground state naive pluripotent stem cells. Nature 2013;504:282–286.
Theunissen TW, Powell BE, Wang H et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 2014;15:471–487.
Chan YS, Goke J, Ng JH et al. Induction of a human pluripotent state with distinct regulatory circuitry that resembles preimplantation epiblast. Cell Stem Cell 2013;13:663–675.
Takashima Y, Guo G, Loos R et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 2014;158:1254–1269.
Guo G, von Meyenn F, Santos F et al. Naive pluripotent stem cells derived directly from isolated cells of the human inner cell mass. Stem Cell Rep 2016;6:437–446.
Theunissen TW, Friedli M, He Y et al. Molecular criteria for defining the naive human pluripotent state. Cell Stem Cell 2016;19:502–515.
Nakamura T, Okamoto I, Sasaki K et al. A developmental coordinate of pluripotency among mice, monkeys and humans. Nature 2016;537:57–62.
Pastor WA, Chen D, Liu W et al. Naive human pluripotent cells feature a methylation landscape devoid of blastocyst or germline memory. Cell Stem Cell 2016;18:323–329.
Sahakyan A, Kim R, Chronis C et al. Human naive pluripotent stem cells model X chromosome dampening and X inactivation. Cell Stem Cell 2017;20:87–101.
Vallot C, Patrat C, Collier AJ et al. XACT noncoding RNA competes with XIST in the control of X chromosome activity during human early development. Cell Stem Cell 2016;20:102–111.
Vallot C, Ouimette JF, Makhlouf M et al. Erosion of X chromosome inactivation in human pluripotent cells initiates with XACT coating and depends on a specific heterochromatin landscape. Cell Stem Cell 2015;16:533–546.
Mekhoubad S, Bock C, de Boer AS et al. Erosion of dosage compensation impacts human iPSC disease modeling. Cell Stem Cell 2012;10:595–609.
Acknowledgements
We are grateful to P Karagiannis for critical reading of this manuscript. YY was supported in part by P-CREATE, Japan Agency for Medical Research and Development (AMED); SICORP, AMED; Research Center Network for Realization of Regenerative Medicine, AMED; JSPS KAKENHI Grant Number 15H04721; the Princess Takamatsu Cancer Research Fund; the Takeda Science Foundation; and the Naito Foundation. MY was supported by JSPS KAKENHI Grant Number 15J05792.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest. S.Y. is a scientific advisor of iPS Academia Japan without salary.
Additional information
In this review, the authors describe the functionality and epigenetic regulations of pluripotent stem cells in rodents and humans in vitro and discuss unsolved issues in developing strategies to capture in vivo pluripotency in vitro.
Rights and permissions
About this article
Cite this article
Yagi, M., Yamanaka, S. & Yamada, Y. Epigenetic foundations of pluripotent stem cells that recapitulate in vivo pluripotency. Lab Invest 97, 1133–1141 (2017). https://doi.org/10.1038/labinvest.2017.87
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2017.87
This article is cited by
-
Anti-fibrotic effect of a selective estrogen receptor modulator in systemic sclerosis
Stem Cell Research & Therapy (2022)
-
Identification of microRNAs related with neural germ layer lineage-specific progenitors during reprogramming
Journal of Molecular Histology (2022)
-
Prevention of tumor risk associated with the reprogramming of human pluripotent stem cells
Journal of Experimental & Clinical Cancer Research (2020)
-
Immortalization of human hepatocytes from biliary atresia with CDK4R24C, cyclin D1, and TERT for cytochrome P450 induction testing
Scientific Reports (2020)
-
Establishment of a Gorlin syndrome model from induced neural progenitor cells exhibiting constitutive GLI1 expression and high sensitivity to inhibition by smoothened (SMO)
Laboratory Investigation (2020)