Abstract
Bitter taste perception, mediated by receptors encoded by the TAS2R loci, has important roles in human health and nutrition. Prior studies have demonstrated that nonsynonymous variation at site 516 in the coding exon of TAS2R16, a bitter taste receptor gene on chromosome 7, has been subject to positive selection and is strongly correlated with differences in sensitivity to salicin, a bitter anti-inflammatory compound, in human populations. However, a recent study suggested that the derived G-allele at rs702424 in the TAS2R16 promoter has also been the target of recent selection and may have an additional effect on the levels of salicin bitter taste perception. Here, we examined alleles at rs702424 for signatures of selection using Extended Haplotype Homozygosity (EHH) and FST statistics in diverse populations from West Central, Central and East Africa. We also performed a genotype–phenotype analysis of salicin sensitivity in a subset of 135 individuals from East Africa. Based on our data, we did not find evidence for positive selection at rs702424 in African populations, suggesting that nucleotide position 516 is likely the site under selection at TAS2R16. Moreover, we did not detect a significant association between rs702424 alleles and salicin bitter taste recognition, implying that this site does not contribute to salicin phenotypic variance. Overall, this study of African diversity provides further information regarding the genetic architecture and evolutionary history of a biologically-relevant trait in humans.
Similar content being viewed by others
Main
Bitter taste receptors, encoded by the TAS2R genes, are involved in a number of biological processes, including oral sensory perception1, 2, 3 and immune response,4, 5, 6 adding a level of complexity to our understanding of these receptors’ function. Prior genetic analyses have reported strong signatures of selection at TAS2R loci on chromosome 7,7, 8, 9, 10, 11 suggesting these genes have been functionally important during human evolution. For example, the derived T-allele at site 516 within the coding exon of TAS2R16 has been identified as a target of positive selection, and has been correlated with increased cell surface expression of the TAS2R16 receptor and heightened sensitivity to salicin bitterness.8, 10 A recent genetic study of global populations (including six African populations) also detected long-range linkage disequilibrium (LD) on chromosomes with both the derived T-allele at site 516 and the derived G-allele at rs702424 in the TAS2R16 promoter, and inferred recent selection for haplotypes carrying these two mutations.9 To test for signatures of selection within the TAS2R16 promoter, we analyzed allelic variation at rs702424 together with sequence data from the TAS2R16 coding exon in a large set of diverse populations from West Central, Central and East Africa. We also tested for an association between genetic variability at rs702424 and threshold levels of salicin taste recognition in individuals from East Africa.
We examined allelic variation (G/A) at rs702424 from the 1M Illumina Duo array genotyped in 697 individuals originating from 44 populations in West Central, Central and East Africa (Supplementary Table 1). African populations with shared genetic similarity, as well as cultural and/or linguistic properties (for example, Khoesan-speaking hunter-gatherers, Niger-Kordofanian speakers and Afroasiatic speakers) were pooled together for subsequent analyses as described by Campbell et al.8 and Tishkoff et al.12. We found that the derived G-allele at rs702424 ranged in frequency from 4.2% in the Bulala population from Chad (Central Africa) to 32.5% in the Luo Nilo-Saharan speakers from Kenya (East Africa) (Supplementary Table 2). To investigate signals of selection based on allele frequency differences, we estimated average FST among populations at rs70242413 and compared this observed estimate with a genome-wide distribution of FST values derived from (a) 1069 randomly sampled single-nucleotide polymorphisms (SNPs) from the 1M Illumina Duo array and (b) HapMap Phase III SNP data in African populations. These analyses indicated that our observed FST estimate (FST=0.01) at rs702424 was not unusual compared with the empirical distribution (occurring at the 27th percentile for both the 1M Illumina and HapMap SNP data), implying the absence of extensive population divergence at this site. Furthermore, our observed FST at rs702424 was not significantly different from mean FST of the 1069 SNPs from the 1M Illumina array based on the one-sample t-test (t=22.8, df=1068, P>0.05). By contrast, however, in a prior analysis of TAS2R16 in a similar set of diverse African populations, we found that observed FST at polymorphic site 516 (FST=0.076) in the coding exon was an outlier (>98th percentile) in the genome-wide distribution of FST values derived from the 1M Illumina data, consistent with a model of local adaptation at this locus.8
To increase power to accurately reconstruct haplotypes, we integrated genotype data at rs702424 with previously collected TAS2R16 coding exon sequences for the same individuals (N=165), and we inferred nine distinct haplotypes. We then constructed a median-joining network (Figure 1), which showed that haplotype diversity grouped into two main clusters: (1) ‘A-G’ haplotypes (defined by the ancestral A-allele at rs702424 and the ancestral G-allele at site 516 within TAS2R16) and (2) ‘A-T’ haplotypes (defined by the ancestral A-allele at rs702424 and the derived T-allele at site 516) and ‘G-T’ haplotypes (defined by the derived G-allele at rs702424 and the derived T-allele at site 516) (Table 1 and Figure 1). Our genealogy also revealed that ‘G-T’ haplotypes were present in divergent African populations with different diets and living in different geographic regions (Table 1; Figure 1), suggesting that these variants are quite old. However, given that the ‘G-T’ haplotypes are positioned furthest from the root of the tree (Figure 1), we argue that the derived G-allele at rs702424 is likely younger than the T-allele at site 516, which was previously estimated to be ∼1 million years old.8 In addition, the reticulation among the ‘A-T’ and ‘G-T’ haplotypes in the ‘high-sensitivity’ clade in Figure 1 indicates that recombination has occurred between variation at rs702424 and the T-allele at site 516 in African populations.
Genealogy of haplotype relationships in African populations. Here the circles represent haplotypes, and the size of the circles indicates the number of chromosomes in our samples with that particular haplotype. The colors within each circle also represent the proportion of that haplotype found in a particular population. The numbers between haplotypes indicate the nucleotide position where a mutation has occurred (−725 is the nucleotide position of rs702424 upstream from the transcription start site of TAS2R16). The dashed lines demarcate haplotypes previously associated with ‘high’ and ‘low’ salicin bitter taste sensitivity.8, 10 In addition, we have indicated haplotypes defined by variation both at rs702424 in the TAS2R16 promoter and at site 516 in the TAS2R16 coding exon (for example, ‘G-T’ haplotypes correspond to the G-allele at rs702424 and the T-allele at site 516).
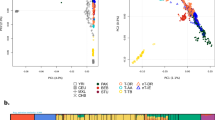
To further explore signatures of selection based on long-range LD patterns, we applied the Extended Haplotype Homozygosity (EHH)14 statistic in African samples for which we had Illumina 1M Duo genotype and TAS2R16 sequence data (N=165). Specifically, we integrated previously sequenced TAS2R16 coding exon data with ∼60 000 SNPs from the Illumina 1M Duo genotyped across the entire length of chromosome 7, and measured the decay of EHH from the core SNP (rs702424). However, we did not observe extensive EHH on chromosomes carrying the derived G-allele at rs702424 compared with chromosomes carrying the ancestral A-allele (Figure 2; Supplementary Figure 1), contrary to the pattern we would expect for a recent selective sweep of variation.
EHH plots for selected African populations. We applied the EHH statistic to identify signatures of recent positive selection based on long-range LD on chromosomes containing the rs702424 core SNP for individuals for whom we also had Illumina Human 1M-Duo genotype and TAS2R16 sequence data (N=165 individuals). The decay of haplotype homozygosity on chromosomes was measured by calculating EHH for the core rs702424 SNP and the surrounding SNPs in the order of increasing distances on either side of the core (in Mb). EHH=0 means all extended haplotypes are different, whereas EHH=1 indicates that all extended haplotypes are the same. Here, the red line represents the decay of homozygosity on chromosomes carrying the derived G-allele at rs702424, whereas the blue line signifies the decay of homozygosity on chromosomes with the ancestral A-allele at the core site. KE-AA and KE-NS are abbreviations for Kenyan Afroasiatic speakers and Kenyan Nilo-Saharan speakers, respectively. Omotic speakers and the Fulani are from Ethiopia and Cameroon, respectively. The number of chromosomes (2N) analyzed for each population is given in parentheses.
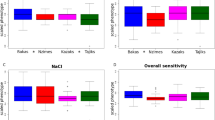
Levels of salicin (a known ligand of the TAS2R16 receptor) sensitivity were measured using a modification of the classic threshold recognition test15 in a subset of 135 individuals from East Africa. We examined the functional effect of common variation at rs702424 by comparing mean salicin score (transformed to a normal distribution) for three genotypic classes: (1) individuals homozygous for the derived G-allele at rs702424 and with at least one T-allele at site 516; (2) individuals heterozygous for variation at rs702424 and with at least one T-allele at site 516; and (3) individuals homozygous for the ancestral A-allele at rs702424 and with at least one T-allele at site 516 using a GLM univariate analysis of variance adjusting for age and sex. We observed a lower mean salicin score for individuals homozygous or heterozygous for the G-allele at rs702424 compared with individuals homozygous for the A-allele at the same site. However, the difference in mean score between these genotypic classes was not statistically significant (P>0.05).
Although previous analyses detected signatures of positive selection at site 516 in the coding exon of TAS2R16 in Africa,8, 10 we did not find evidence of selection at rs702424 in the TAS2R16 promoter based on EHH and FST statistics in a similar set of African populations in the present study. However, a study inferred recent selection for the derived mutations at rs702424 and site 516 primarily in Eurasian populations.9 Given the widespread global distribution of the derived G-allele at rs702424,9 we suggest that this mutation arose on haplotypes with the derived T-allele at site 516 before modern humans migrated from Africa ∼40 000–80 000 years ago. If, indeed, selection occurred only on haplotypes carrying the derived alleles at both rs702424 and site 516 in non-Africans as previously proposed,9 selection at site rs702424 must have occurred after the expansion of modern humans out of Africa. Alternatively, the long-range LD structure present in non-Africans may have resulted from the population bottleneck that accompanied the geographic expansion from Africa.9, 16 Nonetheless, it is clear that the G-allele at rs702424 has not been a target of positive selection in African populations unlike the derived T-allele at site 516. In addition, we did not observe a significant effect of alleles at rs702424 on salicin bitter taste recognition, supporting the recent finding that nonsynonymous variation at site 516 mainly influences salicin phenotypic variance. Although variability at rs702424 did not exhibit signals of selection in Africa and was not correlated with salicin bitter taste, continued research on TAS2R16 and other TAS2R loci (including regulatory variation), together with phenotype data, will be important for understanding both the evolution and function of these understudied receptor genes.
References
Bufe, B., Hofmann, T., Krautwurst, D., Raguse, J. D. & Meyerhof, W. The human TAS2R16 receptor mediates bitter taste in response to beta-glucopyranosides. Nat. Genet. 32, 397–401 (2002).
Greene, T. A., Alarcon, S., Thomas, A., Berdougo, E., Doranz, B. J., Breslin, P. A. et al. Probenecid inhibits the human bitter taste receptor TAS2R16 and suppresses bitter perception of salicin. PLoS ONE 6, e20123 (2011).
Hayes, J. E., Wallace, M. R., Knopik, V. S., Herbstman, D. M., Bartoshuk, L. M. & Duffy, V. B. Allelic variation in TAS2R bitter receptor genes associates with variation in sensations from and ingestive behaviors toward common bitter beverages in adults. Chem. Senses 36, 311–319 (2011).
Lee, R. J., Xiong, G., Kofonow, J. M., Chen, B., Lysenko, A., Jiang, P. et al. T2R38 taste receptor polymorphisms underlie susceptibility to upper respiratory infection. J. Clin. Invest. 122, 4145–4159 (2012).
Prince, A. The bitter taste of infection. J. Clin. Invest. 122, 3847–3849 (2012).
Shah, A. S., Ben-Shahar, Y., Moninger, T. O., Kline, J. N. & Welsh, M. J. Motile cilia of human airway epithelia are chemosensory. Science 325, 1131–1134 (2009).
Campbell, M. C., Ranciaro, A., Froment, A., Hirbo, J., Omar, S., Bodo, J. M. et al. Evolution of functionally diverse alleles associated with PTC bitter taste sensitivity in Africa. Mol. Biol. Evol. 29, 1141–1153 (2012).
Campbell, M. C., Ranciaro, A., Zinshteyn, D., Rawlings-Goss, R., Hirbo, J., Thompson, S. et al. Origin and differential selection of allelic variation at TAS2R16 associated with salicin bitter taste sensitivity in Africa. Mol. Biol. Evol. 31, 288–302 (2014).
Li, H., Pakstis, A. J., Kidd, J. R. & Kidd, K. K. Selection on the human bitter taste gene, TAS2R16, in Eurasian populations. Hum. Biol. 83, 363–377 (2011).
Soranzo, N., Bufe, B., Sabeti, P. C., Wilson, J. F., Weale, M. E., Marguerie, R. et al. Positive selection on a high-sensitivity allele of the human bitter-taste receptor TAS2R16. Curr. Biol. 15, 1257–1265 (2005).
Wooding, S., Kim, U. K., Bamshad, M. J., Larsen, J., Jorde, L. B. & Drayna, D. Natural selection and molecular evolution in PTC, a bitter-taste receptor gene. Am. J. Hum. Genet. 74, 637–646 (2004).
Tishkoff, S. A., Reed, F. A., Friedlaender, F. R., Ehret, C., Ranciaro, A., Froment, A. et al. The genetic structure and history of Africans and African Americans. Science 324, 1035–1044 (2009).
Fstat 2.9.3.2. [updated 2005; cited 2011 Jun 22]. Available at http://www2.unil.ch/popgen/softwares/fstat.htm (2002).).
Sabeti, P. C., Walsh, E., Schaffner, S. F., Varilly, P., Fry, B., Hutcheson, H. B. et al. The case for selection at CCR5-Delta32. PLoS Biol. 3, e378 (2005).
Harris, H. & Kalmus, H. The distribution of taste thresholds for phenylthiourea of 384 sib pairs. Ann. Eugen. 16, 226–230 (1951).
Schmegner, C., Hoegel, J., Vogel, W. & Assum, G. Genetic variability in a genomic region with long-range linkage disequilibrium reveals traces of a bottleneck in the history of the European population. Hum. Genet. 118, 276–286 (2005).
Acknowledgements
We thank Fathya Abdo, Eva Aluvalla, Godfrey Lema, Birhanu Mekaunintie, Alemayehu Moges, Hussein Musa, Lilian Nyindodo and Solomon Taye who helped with the field work, and the various institutions in West Central, Central and East Africa for their generous support. We thank Dongbo Hu and Wen-Ya Ko for useful discussions. We also offer our deepest appreciation to the many Africans who generously donated their DNA and time so that we can learn more about African genetic diversity. This work was supported by the US National Science Foundation (grant numbers BCS-0552486, BCS-0827436), US National Institutes of Health (grant numbers R01GM076637, 5DP1ES022577 05) to SAT and US National Institutes of Health (grant number RO1 DC02995) to PASB.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Campbell, M., Ranciaro, A., Zinshteyn, D. et al. Limited evidence for adaptive evolution and functional effect of allelic variation at rs702424 in the promoter of the TAS2R16 bitter taste receptor gene in Africa. J Hum Genet 59, 349–352 (2014). https://doi.org/10.1038/jhg.2014.29
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2014.29
This article is cited by
-
Taste buds: cells, signals and synapses
Nature Reviews Neuroscience (2017)
-
The Bitter Taste Receptor TAS2R16 Achieves High Specificity and Accommodates Diverse Glycoside Ligands by using a Two-faced Binding Pocket
Scientific Reports (2017)