Abstract
We report duplications of maternally derived chromosome 11p15 involving CDKN1C encoding a negative regulator for cell proliferation in three Japanese patients (cases 1 and 2 from family A and case 3 from family B) with Silver–Russell syndrome (SRS) phenotype lacking hemihypotrophy. Chromosome analysis showed 46,XX,der(16)t(11;16)(p15.3;q24.3)mat in case 1, 46,XY,der(16)t(11;16)(p15.3;q24.3)mat in case 2 and a de novo 46,XX,der(17)t(11;17)(p15.4;q25.3) in case 3. Genomewide oligonucleotide-based array comparative genomic hybridization, microsatellite analysis, pyrosequencing-based methylation analysis and direct sequence analysis revealed the presence of maternally derived extra copies of the distal chromosome 11p involving the wild-type CDKN1C (a ~7.98 Mb region in cases 1 and 2 and a ~4.43 Mb region in case 3). The results, in conjunction with the previous findings in patients with similar duplications encompassing CDKN1C and in those with intragenic mutations of CDKN1C, imply that duplications of CDKN1C, as well as relatively mild gain-of-function mutations of CDKN1C lead to SRS subtype that usually lack hemihypotrophy.
Similar content being viewed by others
Introduction
Silver–Russell syndrome (SRS) is a congenital developmental disorder characterized by pre- and postnatal growth failure, relative macrocephaly, hemihypotrophy and fifth-finger clinodactyly.1 Recent studies have shown that epimutation (hypomethylation) of the paternally derived H19-differentially methylated region (DMR) at the imprinting control region 1 (ICR1) on chromosome 11p15.5 and maternal uniparental disomy 7 account for ~45 and ~5% of SRS patients, respectively.1 Thus, underlying (epi)genetic factors still remain to be clarified in a substantial fraction of SRS patients, although several rare (epi)genetic aberrations have been identified in a small fraction of SRS patients.1
CDKN1C (cyclin-dependent kinase inhibitor 1C) is a maternally expressed gene that resides at the ICR2 just proximal to the ICR1.2 CDKN1C encodes a negative regulator for cell proliferation and, consistent with this, loss-of-function mutations of CDKN1C cause Beckwith–Wiedemann syndrome associated with overgrowth.2, 3 Furthermore, recent studies have shown that gain-of-function mutations of CDKN1C result in IMAGe syndrome (IMAGeS) characterized by intrauterine growth restriction, metaphyseal dysplasia, adrenal hypoplasia congenita and male genital abnormalities,2 whereas less severe gain-of-function mutations of CDKN1C have been identified in a large family with maternally inherited SRS.4 Thus, it has been suggested that relatively severe and mild CDKN1C gain-of-function effects lead to IMAGeS and SRS, respectively.4, 5 Notably, IMAGeS patients satisfy the diagnostic criteria for SRS proposed by Nechine et al.5, 6 and IMAGeS and SRS patients with CDKN1C mutations invariably lack hemihypotrophy characteristic of SRS.4, 5, 6
Here, we report three patients with SRS and duplications of maternally derived chromosome 11p15.5 involving CDKN1C. The results, in conjunction with previous findings, imply that duplications of CDKN1C, as well as relatively mild gain-of-function mutations of CDKN1C lead to SRS subtype that usually lack hemihypotrophy.
Case reports
Patients
We studied three Japanese patients (cases 1–3) from two families (Figure 1). Cases 1–3 satisfied the SRS diagnostic criteria proposed by Netchine et al.,6 although they lacked hemihypotrophy (Table 1, see its footnote for Netchine SRS criteria). Oligohydramnios characteristic of SRS7 was also noticed during the pregnancies of cases 2 and 3. They exhibited no IMAGeS-like phenotypes such as radiologically discernible skeletal dysplasia, an episode suggestive of adrenal dysfunction or undermasculinized genitalia in male case 2.
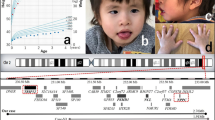
The pedigrees of families A and B and a photograph of case 3. In family A, cases 1 and 2 have an unbalanced translocation involving the distal part of chromosome 11p, the mothers of cases 1 and 2, as well as the brother of case 1 have a balanced translocation involving the distal part of chromosome 11p and the remaining subjects have a normal karyotype. In family B, case 3 has an unbalanced translocation involving the distal part of chromosome 11p and the parents have a normal karyotype. Case 3 exhibits SRS-compatible phenotypes such as prominent forehead, triangular face with relative macrocephaly and micrognathia, ear anomalies and short and curved fifth fingers, but is free from hemihypotrophy.
Cytogenetic and molecular studies
This study was approved by the Institute Review Board Committee at Hamamatsu University School of Medicine and was performed using peripheral leukocyte samples and primers shown in Supplementary Table S1 after obtaining written informed consent. The methods for molecular studies were as reported previously.7 We also obtained written informed consent to publish the facial photograph of case 3 from the parents.
Chromosome analysis showed 46,XX,der(16)t(11;16)(p15.3;q24.3)mat in case 1, 46,XY,der(16)t(11;16)(p15.3;q24.3)mat in case 2 and a de novo 46,XX, der(17)t(11;17)(p15.4;q25.3) in case 3 (Figure 1). Then, genomewide oligonucleotide-based array comparative genomic hybridization was carried out using a catalog human array (2 × 400 K format, ID G4448A) (Agilent Technologies, Santa Clara, CA, USA), revealing the presence of three copies of the distal parts of chromosome 11p involving the ICR1 and the ICR2 in cases 1–3 (a ~7.98 Mb region in cases 1 and 2 and a ~4.43 Mb region in case 3) (Figure 2a). No discernible deletion was identified on the distal chromosome 16q in cases 1 and 2, indicating the position of the chromosome 16q breakpoint at the very telomeric portion, whereas a ~200 kb deletion was detected in the telomeric portion of chromosome 17q in case 3. There was no other copy number alteration that was not registered in the Database of Genomic Variants (http://dgv.tcag.ca/dgv/app/home). Microsatellite analysis was carried out for four loci on the duplicated chromosome 11p, showing the presence of two alleles of maternal origin and a single allele of paternal origin in cases 1–3 (Figure 2b and Supplementary Table S2). Subsequently, pyrosequencing-based methylation analysis was performed for four CpG dinucleotides (CG1–CG4) within the H19-DMR and six CpG dinucleotides (CG5–CG10) within the KvDMR1 using bisulfite-treated leukocyte genomic DNA samples and methylation index (MI, the ratio of methylated clones) was obtained for each of CG1–CG10 using PyroMark Q24 (Qiagen, Valencia, CA, USA) (Figure 2c). In cases 1–3, the MIs for CG1–CG4 were mildly decreased or around the lower limit of the normal range and those for CG5–CG10 were mildly increased or around the upper limit of the normal range. Direct sequence analysis showed no discernible mutation on the CDKN1C coding region.
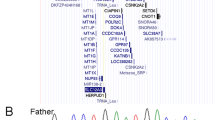
Representative molecular findings. (a) Array comparative genomic hybridization analysis. The black, red and green dots represent signals indicative of the normal, increased (log2 signal ratio>+0.5) and decreased (log2 signal ratio<−0.8) copy numbers, respectively. The log2 signal ratios of +0.5 and −1.0 indicate the presence of three copies and a single copy of the corresponding regions, respectively. The red and the green rectangles represent increased and decreased copy number regions, respectively. The yellow rectangles denote the regions encompassing the ICR1 and the ICR2. (b) Microsatellite analysis for D11S4088 proximal to the KvDMR1. Unequal amplification of the heterozygous peaks in each subject is consistent with short products being more easily amplified than long products and comparison of area under curves of the 212 bp and the 234 bp alleles between case 3 and the mother indicates the presence of two 212 bp alleles and a single 234 bp allele in case 3. This implies that the maternal 212 and 234 bp alleles and the paternal 212 bp allele have been transmitted to case 3. (c) Pyrosequencing-based methylation analysis of the H19-DMR at the ICR1 and the KvDMR1 at the ICR2, using bisulfite-treated genomic DNA. The cytosine residues at the CpG dinucleotides within the H19-DMR is methylated after paternal transmission (filled circles) and unmethylated after maternal transmission (open circles), whereas those within the KvDMR1 is unmethylated after paternal transmission (open circles) and methylated after maternal transmission (filled circles). Paternally and maternally expressed genes are shown in blue and red, respectively. For the H19-DMR, a segment encompassing 21 CpG dinucleotides was PCR amplified with PyF1 and PyR1 primers and a sequence primer (SP1) was hybridized to a single-stranded PCR product. Subsequently, the MIs were obtained for four CpG dinucleotides (CG1–CG4) (indicated with a yellow rectangle). The blue rectangle indicates the CTCF binding site 6. The CpG dinucleotide between CG1 and CG2 was not examined, because it constitutes a C/T SNP (indicated with gray circles). The KvDMR1 was similarly examined using PyF2 and PyR2 primers and SP2 and the MIs were obtained for CG5–CG10. The MIs are summarized in the bottom table. F, father; and M, mother; P, patient.
Discussion
Cases 1–3 had SRS without hemihypotrophy (body asymmetry) in the presence of maternally derived extra copies of the distal chromosome 11p involving the ICR1 and the ICR2. This implies that the SRS phenotype lacking hemihypotrophy in cases 1–3 is primarily caused by two copies of maternally expressed genes on the two ICRs. In this regard, of duplicated maternally expressed genes, CDKN1C functions as a negative growth regulator8 and CDKN1C gain-of-function mutations have been identified in SRS and IMAGeS,2, 4, 5 whereas neither H19 nor KCNQ1 appears to have a positive role in growth regulation. Indeed, H19 is regarded as a possible tumor suppressor gene9 and KCNQ1 encoding a voltage-gated potassium channel is involved in cardiac arrhythmias.10 Thus, it is likely that SRS phenotype lacking hemihypotrophy in cases 1–3 is primarily caused by the presence of two functional copies of the wild-type CDKN1C. It should be pointed out, however, that although the der(16)t(11;16)(p15.3;q24.3) chromosome in cases 1 and 2 had no discernible chromosome 16q deletion, the der(17)t(11;17)(p15.4;q25.3) chromosome in case 3 was missing the ~200 kb telomeric 17q region that harbors several genes. In addition, there are multiple nonimprinted genes on the duplicated chromosome 11p15 regions. Thus, altered dosage of such genes may have exerted a certain effect on growth patterns of cases 1–3.
An extra copy of maternally derived chromosome 11p15 involving CDKN1C has been identified in 16 patients (Table 1) (for detailed clinical features of each case, see Supplementary Table S3).11, 12, 13, 14, 15, 16, 17, 18, 19 Notably, although they frequently show SRS-like phenotype, hemihypotrophy (body asymmetry) has been found only in a single case12 and none of them exhibit IMAGeS-like skeletal, adrenal or genital manifestation. This provides further support for the notion that two copies of maternally derived CDKN1C, as well as mild gain-of-function mutations of CDKN1C usually lead to SRS subtype lacking hemihypotrophy.
References
Eggermann, T. Russell-Silver syndrome. Am. J. Med. Genet. C Semin. Med. Genet. 154C, 355–364 (2010).
Arboleda, V.A., Lee, H., Parnaik, R., Fleming, A., Banerjee, A. & Ferraz-de-Souza, B. et al. Mutations in the PCNA-binding domain of CDKN1C cause IMAGe syndrome. Nat. Genet. 44, 788–792 (2012).
Soejima, H. & Higashimoto, K. Epigenetic and genetic alterations of the imprinting disorder Beckwith–Wiedemann syndrome and related disorders. J. Hum. Genet. 58, 402–409 (2013).
Brioude, F., Oliver-Petit, I., Blaise, A., Praz, F., Rossignol, S. & Le Jule, M. et al. CDKN1C mutation affecting the PCNA-binding domain as a cause of familial Russell Silver syndrome. J. Med. Genet. 50, 823–830 (2013).
Kato, F., Hamajima, T., Hasegawa, T., Amano, N., Horikawa, R. & Nishimura, G. et al. IMAGe syndrome: clinical and genetic implications based on investigations in three Japanese patients. Clin. Endocrinol. 80, 706–713 (2014).
Netchine, I., Rossignol, S., Dufourg, M. N., Azzi, S., Rousseau, A. & Perin, L. et al. 11p15 imprinting center region 1 loss of methylation is a common and specific cause of typical Russell-Silver syndrome: clinical scoring system and epigenetic-phenotypic correlations. J. Clin. Endocrinol. Metab. 92, 3148–3154 (2007).
Fuke, T., Mizuno, S., Nagai, T., Hasegawa, T., Horikawa, R. & Miyoshi, Y. et al. Molecular and clinical studies in 138 Japanese patients with Silver–Russell syndrome. PLoS ONE 8, e60105 (2013).
Lee, M. H., Reynisdottir, I. & Massague, J. Cloning of p57(KIP2), a cyclin-dependent kinase inhibitor with unique domain structure and tissue distribution. Genes Dev. 9, 639–649 (1995).
Hao, Y., Crenshaw, T., Moulton, T., Newcomb, E. & Tycko, B. Tumour-suppressor activity of H19 RNA. Nature 365, 764–767 (1993).
Wang, Q., Curran, M. E., Splawski, I., Burn, T. C., Millholland, J. M. & VanRaay, T. J. et al. Positional cloning of a novel potassium channel gene: KVLQT1 mutations cause cardiac arrhythmias. Nat. Genet. 12, 17–23 (1996).
Fisher, A. M., Thomas, N. S., Cockwell, A., Stecko, O., Kerr, B. & Temple, I. K. et al. Duplications of chromosome 11p15 of maternal origin result in a phenotype that includes growth retardation. Hum. Genet. 111, 290–296 (2002).
Eggermann, T., Meyer, E., Obermann, C., Heil, I., Schüler, H. & Ranke, M. B. et al. Is maternal duplication of 11p15 associated with Silver–Russell syndrome? J. Med. Genet. 42, e26 (2005).
Schönherr, N., Meyer, E., Roos, A., Schmidt, A., Wollmann, H. A. & Eggermann, T. et al. The centromeric 11p15 imprinting centre is also involved in Silver–Russell syndrome. J. Med. Genet. 44, 59–63 (2007).
South, S. T., Whitby, H., Maxwell, T., Aston, E., Brothman, A. R. & Carey, J. C. Co–occurrence of 4p16.3 deletions with both paternal and maternal duplications of 11p15: modification of the Wolf–Hirschhorn syndrome phenotype by genetic alterations predicted to result in either a Beckwith–Wiedemann or Russell–Silver phenotype. Am. J. Med. Genet. A 146A, 2691–2697 (2008).
Bliek, J., Snijder, S., Maas, S. M., Polstra, A., van der Lip, K. & Alders, M. et al. Phenotypic discordance upon paternal or maternal transmission of duplications of the 11p15 imprinted regions. Eur. J. Med. Genet. 52, 404–408 (2009).
Eggermann, T., Spengler, S., Bachmann, N., Baudis, M., Mau-Holzmann, U. A. & Singer, S. et al. Chromosome 11p15 duplication in Silver-Russell syndrome due to a maternally inherited translocation t(11;15). Am. J. Med. Genet. A 152A, 1484–1487 (2010).
Cardarelli, L., Sparago, A., De Crescenzo, A., Nalesso, E., Zavan, B. & Cubellis, M. V. et al. Silver-Russell syndrome and Beckwith–Wiedemann syndrome phenotypes associated with 11p duplication in a single family. Pediatr. Dev. Pathol. 13, 326–330 (2010).
Bonaldi, A., Mazzeu, J. F., Costa, S. S., Honjo, R. S., Bertola, D. R. & Albano, L. M. et al. Microduplication of the ICR2 domain at chromosome 11p15 and familial Silver-Russell syndrome. Am. J. Med. Genet. A 155A, 2479–2483 (2011).
Chiesa, N., De Crescenzo, A., Mishra, K., Perone, L., Carella, M. & Palumbo, O. et al. The KCNQ1OT1 imprinting control region and non-coding RNA: new properties derived from the study of Beckwith-Wiedemann syndrome and Silver–Russell syndrome cases. Hum. Mol. Genet. 21, 10–25 (2012).
Acknowledgements
This work was supported by Grants-in-Aid for Scientific Research (A) (25253023) and Research (B) (23390083) from the Japan Society for the Promotion of Science, by Grants for Research on Intractable Diseases (H22-161) from the Ministry of Health, Labor and Welfare (MHLW) and by Grant for National Center for Child Health and Development (25-10).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Nakashima, S., Kato, F., Kosho, T. et al. Silver–Russell syndrome without body asymmetry in three patients with duplications of maternally derived chromosome 11p15 involving CDKN1C. J Hum Genet 60, 91–95 (2015). https://doi.org/10.1038/jhg.2014.100
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2014.100
This article is cited by
-
Frequency and clinical characteristics of distinct etiologies in patients with Silver-Russell syndrome diagnosed based on the Netchine-Harbison clinical scoring system
Journal of Human Genetics (2022)
-
Prospective study of epigenetic alterations responsible for isolated hemihyperplasia/hemihypoplasia and their association with leg length discrepancy
Orphanet Journal of Rare Diseases (2021)
-
Methylation of Cdkn1c may be involved in the regulation of tooth development through cell cycle inhibition
Journal of Molecular Histology (2018)
-
Diagnosis and management of Silver–Russell syndrome: first international consensus statement
Nature Reviews Endocrinology (2017)