Abstract
Deafness is an etiologically heterogeneous trait with many known genetic, environmental causes or a combination thereof. The identification of more than 120 independent genes for deafness has provided profound new insights into the pathophysiology of hearing. However, recent findings indicate that a large proportion of both syndromic and non-syndromic forms of deafness in the Chinese population are caused by defects in a small number of genes. Studies of the genetic epidemiology and molecular genetic features revealed that there is a clear relevance of genes causing deafness in Chinese deaf patients as well as a unique spectrum of common and rare deafness gene mutations in the Chinese population. This review is focused on the genetic aspects of non-syndromic and mitochondrial deafness, in which unique molecular genetic features of hearing impairment have been identified in the Chinese population. The current China population is approximately 1.3 billion. It is estimated that 30 000 infants are born with congenital sensorineural hearing loss each year. Better understanding of the genetic causes of deafness in the Chinese population is important for accurate genetics counseling and early diagnosis for timely intervention and treatment options.
Similar content being viewed by others
Introduction
Hearing loss (HL) is a common congenital disorder and one of the most distressing disorders affecting humanity. Deafness can result from a mutation in a single or a combination of mutations of genes; environmental causes, such as medical problems, environment exposure, trauma and medications, or interactions between genetic and environmental factors.1, 2 Worldwide, congenital deafness occurs in 1 in 1000 live births, and more than half of these cases are hereditary.2, 3 Of the estimated minimum of 50% of cases with inherited hearing loss, 70% of these are non-syndromic and 80% of these are autosomal recessive.4 Non-syndromic hearing loss (NSHL) is most often sensorineural. It can be divided into DFNA (autosomal dominant deafness, 15–20%), DFNB (autosomal recessive deafness, 80%), DFN (X-linked deafness, 1%), and mitochondrial deafness of at least 1%.4, 5 To date, 121 deafness loci have been reported, 54 DFNA loci and 67 DFNB loci. More than 40 genes for monogenetic NSHL and even more for syndromic HL have been cloned (http://webhost.ua.ac.be/hhh/).6
Despite the genetic heterogeneity of deafness, there is evidence that a large proportion of non-syndromic forms of hearing loss in Chinese populations is caused by a limited number of recurrent mutations resulting from common founders or a mutational hot spot. It is well known that there are racial and ethnic differences in the causes, expression, and prevalence of various Mendelian diseases and it is not surprising that groups with a common ancestry also share genetic risk factors for particular human disorders.7, 8 Some recessive diseases could become prevalent in a population either because the heterozygous state is associated with a selective advantage and/or because there was a change in frequency of the mutant allele caused by genetic drift or loss of the beneficial allele.9 Thus, knowledge of a person's ancestral origin can be informative for the provision of health care, including testing, diagnosis and treatment when genetic factors are involved.10 A series of studies have revealed genes responsible for Chinese deafness patients, and the differences in mutation spectrum between Chinese (probably other Asian populations) and populations with European ancestry.
According to the China Disabled Persons’ Federation website, China has approximately 21 million people with hearing loss out of the 60 million disabled. Although there has been a significant advance in the knowledge of the molecular basis of hereditary deafness in recent years, there have been few studies in China to evaluate the effects of genetic etiologic factors. The magnitude of the problem in China remains largely undefined. State-of-the-art molecular testing is now available for the most common causes of hereditary hearing loss. However, only with knowledge of prevalent population-specific mutations, can an effective strategy for detecting deafness genes and an appropriate genetic counseling be developed. The importance of identifying the genetic basis for deafness among Jews and Japanese population have been reported earlier.11, 12 Here, we focus on the implications of identifying the molecular genetic features in the largest population in the world for patient care, molecular diagnosis and better genetic counseling.
Non-syndromic recessive deafness (DFNB)
DFNB1 (GJB2/GJB6)
Connexin 26 has been found to be the most common cause of deafness in many ethnic populations around the world. GJB2 encodes a gap junction protein expressed in the cochlea and is thought to be important for recycling potassium ions that flow into sensory hair cells as part of the transduction current. Mutations of GJB2, which is responsible for DFNB1 and DFNA3, are the most frequent cause of inherited hearing loss.13, 14, 15 More than 100 different mutations in GJB2 have been identified in patients with non-syndromic deafness and a significant difference in the frequency and distribution of the mutations have been observed in different populations (http://davinci.crg.es/deafness/). They are responsible for approximately half the cases in the United States, Europe, Australia, and Israel, and have been reported in other populations as well. Most interestingly, a single mutation, 35delG, accounts for up to 70% of Northern and Southern European, as well as American Caucasian populations, with a carrier frequency ranging from 1.3 to 2.8%.16, 17 Another GJB2 mutation, the 167delT, accounts for 40% of the pathologic alleles in the Jewish deaf population18 and has a 4% carrier frequency among Ashkenazi Jews.19 The 235delC mutation is the most prevalent in Eastern Asian (Japanese, Chinese and Korean) populations, R143W has been described as a recessive cause of deafness in Ghana and the W24X, the most common GJB2 mutation in India, has also been found in high prevalence in Slovak and Spanish gypsies (www.gendeaf.org). A series of reports have described genotype–phenotype correlations for most GJB2 genotypes, enabling, to a certain extent, to predict the phenotype expression on the basis of the genotype. However, for the majority of these genotypes, a certain degree of phenotypic variability was still observed, which was most striking for 35delG homozygous patients. Hearing loss of 35delG homozygous patients ranges from mild to profound.20, 21 There was inter- and intrafamilial phenotypic variability and the hearing loss ranged from mild (least often) over moderate to severe and profound (most frequent).
More than twenty GJB2 mutations have been reported in Chinese patients (Figure 1). Mutation screening of GJB2 in Chinese deaf probands revealed that the deletion of cytosine at position 235 (235delC) mutation was the most prevalent mutation (20.3% of alleles), accounting for 81% of the pathologic alleles in multiplex cases and 67% in simplex cases.22 We further confirmed that 235delC is the most common pathologic mutation in inherited deafness attributed to GJB2 in the Chinese population.23 We screened DNA samples from 3004 patients with non-syndromic hearing impairment from 26 regions of China and found that the 235delC allele frequency varies from 0.9 to 19.5%. These results indicate that mutations in GJB2 are a major cause of inherited and sporadic congenital deafness in the Chinese population. We have recently sequenced the coding region of GJB2 gene from 2063 unrelated probands with non-syndromic deafness and identified a total of 24 mutations (Figure 1 and Table 1) (Dai et al., unpublished data). Eight of these mutations have not been previously reported. The novel mutations include six frame shifts (c.99delT, c.155_158delTCTG, c.311_324del14, c.388_397del10, c.512_515insAACG and c.605_651in46), a single amino acid deletion (c.424_ 426delTCC/del142F), and one nonsense mutation (p.W3X). The two most prevalent mutations including c.235delC and c.299_c.300delAT account for 82% of all mutant alleles identified.
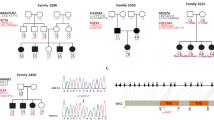
Schematic representation of the relative linear location of the mutations of GJB2 gene in the Chinese population. Novel mutations are in bold letters (Dai et al., unpublished data). The structural motifs of connexins are NT: amino-terminus; M1-M4: transmembrane domains 1–4; E1/E2: extracellular domains; CL: cytoplasmic loop; CT: carboxyl-terminus.
The status of the 235delC mutation, which seems to be a unique mutation in Asian populations, is comparable to the 35delG mutation in Caucasian populations. Patients with homozygous or compound heterozygous 235delC mutation exhibit a comparative phenotype varying from mild to profound loss of hearing. The high frequency of the 35delG GJB2 allelic variant in the white population has been shown to be the result of a founder effect, rather than a mutational hot spot.41 The 167delT mutation present in Ashkenazi Jews has also been attributed to a founder effect.19 Both 35delG and 167delT mutations are absent or exceptionally low in Asian populations, in whom, the 235delC is the most prevalent, with carrier rates ranging from 1.0 to 1.3%. This mutation accounts for up to 80% of pathogenic GJB2 alleles among Japanese,14, 42, 43 and Koreans.44 Interestingly, the 235delC has not been detected in South Asian populations from India, Pakistan, Bangladesh and Sri Lanka, in whom the prevalent GJB2 mutations are W24X and W77X.45, 46 The high frequency of the 235delC mutation in East Asian populations has subsequently been shown to be the result of a founder effect, rather than a mutational hot spot, and 235delC among all East Asian populations was derived from a common ancestral founder.47, 48
A role for GJB6, the gene adjacent to GJB2 on chromosome 13, was first suggested in 1999. The most common mutation in GJB6 is a 342-kb deletion, which causes NSHL when homozygous, or when present in trans with a recessive GJB2 mutation. The GJB6 deletion may account for 10% of all DFNB1 alleles with an extremely wide range based on ethnic origin. GJB2 and GJB6 genes are expressed in the cochlea where they can combine to form multi-unit hemichannels in the cell membrane, and function as an integral component of the potassium regulation in the inner ear. The 342-kb deletion (GJB6-D13S1830) mutation is most frequent in Spain, France, the United Kingdom, Israel and Brazil (5.9–9.7% of all DFNB1 alleles); it is less frequent in the USA, Belgium and Australia (1.3–4.5% of all DFNB1 alleles), and is very rare in Southern Italy,49 but has been found, present in Northern Italy and in Germany at frequencies similar to those of other European countries.50, 51 The deletion was not found in Austria52 Turkey53, 54 and China.55, 56, 57, 58
DFNB2 (MYO7A)
Myosin VIIA, located at 11q13.5, is an unconventional myosin that has been implicated in recessively inherited USH1B,7 recessively inherited atypical USH,59, 60 non-syndromic autosomal recessive hearing impairment (DFNB2)24 and autosomal dominant hearing impairment (DFNA11).61 Mutations in MYO7A, are associated with Usher syndrome type 1B in many populations, and more than one hundred different mutant alleles of MYO7A have been reported (www.hgmd.cf.ac.uk/ac/gene.php?gene=MYO7A). We have previously shown that three recessive alleles (pR2444P, IVS3nt-2a → g and val1199insT) of the MYO7A gene cause DFNB2 in families from China.24 In a recent study of 24 consanguineous Pakistani families segregating deafness as a recessive trait by Riazuddin et al.62 a total of 23 of these families were found to segregate USH1B whereas one family segregated DFNB2. Data of functional analyses of MYO7A mutations is consistent with the hypothesis that DFNB2-causative mutations lead to a less severe phenotype than the USH1B-associated mutations because the resulting proteins retain some degree of normal activity.
DFNB4 (SLC26A4)
Pendred syndrome is an inherited disorder that accounts for as much as 10% of hereditary deafness.63 The syndrome is characterized by congenital severe-to-profound sensorineural hearing loss (SNHL) and euthyroid goiter. Goiter is not present at birth and develops in early puberty (40%) or adulthood. Vestibular function is abnormal in the majority of affected persons. Recessive mutations of the SLC26A4 (PDS) gene on chromosome 7q can cause sensorineural deafness with goiter64 or NSHL without goiter.65 Mutations in the SLC26A4 gene are identified in about 50% of multiplex families. These mutations disrupt in vitro transmembrane anion/base exchange activity of the SLC26A4 polypeptide, pendrin.66 So far, more than 100 mutations causing Pendred syndrome and non-syndromic (EVA) sensorineural hearing loss have been reported in the PDS gene (http://www.healthcare.uiowa.edu/labs/pendredandbor//slcMutations.htm). Moreover, a significant difference in the frequency and distribution of the mutations has been observed in different populations.
To date, more than 40 mutations have been reported among the Chinese deaf patients (Figure 2). The IVS7-2A>G mutation was the most common form accounting for 22.3–57.62% of all the mutant alleles in Chinese patients with hearing loss associated with enlarged vestibular aqueduct (EVA) or both EVA and Mondini dysplasia.25, 26 Among Asian populations, a distinct spectrum of SLC26A4 mutations in patients associated with EVA with highly prevalent founder mutations has been observed, such as p.H723R in Japan and Korea and IVS7-2A>G in Taiwan and mainland China.67, 32, 68, 69, 34 Park et al.32 performed a mutation screening of the seven exons of the SLC26A4 gene in 274 East Asians and 318 South Asians deaf probands with sporadic or familial severe-to-profound prelingual deafness, including 86 Chinese affected subjects. They demonstrated mutations in SLC26A4 about 5.5% of both groups and identified three mutations (S252P, IVS7-2 A>G, N392Y) in 5 out of 86 (5.8%) Chinese probands. Tsukamoto et al.28 reported causative mutations have been identified in 90% of typical Pendred families in Japan, 78% among them had sensorineural hearing loss associated with EVA and 53% of mutations were the H723R mutation. Campbell et al.70 stated that L236P, T416P and IVS8+1G>A were the three frequent mutations in the Caucasian population.70 In the Spanish population the most frequent mutation is Q514K (17%).71
The IVS7-2A>G mutation was first reported in a Turkish family72 and subsequently detected in deaf subjects from Japan, Korea and China.32, 28, 73, 74 IVS7-2A>G has not been found in western populations, including deaf patients in the United States,70, 73, 75, 76 http://www.genetests.org/;77 We have recently reported a comprehensive study of the prevalence of IVS7-2A>G mutation in an analysis of 3271 deaf subjects from 27 regions of 24 provinces of China. The detection rate was found to range from 28% to below 1% based on ethnicity. The results showed that the genetic test for the IVS7-2A>G mutation alone would identify the molecular cause in up to 8–12% of deaf patients in few eastern and central regions of China. However, the IVS7-2A>G mutation occurs at a much lower frequency (0.000–0.019) among ethnic groups in the southwest and northwest regions of China.78 The data suggest that for subjects that are negative for the IVS7-2A>G mutation, further mutational analysis of the full SLC26A4 gene or other deafness-related genes will be necessary.
Non-syndromic dominant hearing loss (DFNA)
DFNA2 (GJB3)
The Connexin 31 gene (GJB3) mapped to chromosome 1p33–p35 encodes for a protein that is 270 amino acids long. Mutations in the GJB3 gene have been reported to be responsible for both DFNA and DFNB.45, 35, 36 More than 10 different mutations have been found in patients with deafness from Chinese, Brazilian and Spanish populations.74, 36, 79, 80 However, the pathogenicity of most of these sequence alterations still remains questionable. Mutations in GJB3 have originally been shown to underlie an autosomal dominant form of non-syndromic deafness (DFNA2) in Chinese patients; a missense mutation (E183K) and a nonsense mutation (R180X) of GJB3 were found to be associated with high-frequency hearing loss.36 We subsequently showed that mutations in GJB3 also underlie recessive non-syndromic form of deafness in this population.79 These findings indicate that GJB3 variations are common in non-syndromic hearing loss patients in the Chinese population. In contrast, the studies described so far have shown that variations in GJB3 have no or a low genetic relevance in European and Caucasian populations in general. Extensive sequence analysis of the coding region and the 5′ UTR of GJB3 in all five first DFNA2 families originating from Indonesia, the United States, Belgium and the Netherlands81, 82 revealed no mutations.83 Nonetheless, the dominant D66del mutation in GJB3 linked to neuropathy and deafness was identified in one Spanish deaf patient.74 The failure in finding HL causing mutations in GJB3 in a screening of sporadic cases with NSHL performed in California and in Austria further confirms population relevance of GJB3 mutations in the Chinese population.84, 85
In a screening of 108 GJB2 heterozygous Chinese patients with non-syndromic deafness without obvious dominant inheritance patterns, we recently showed that mutations in Cx26 and Cx31 genes can interact to cause hearing loss in digenic heterozygotes in humans. Direct physical interaction of Cx26 with Cx31 is supported by data showing that Cx26 and Cx31 have overlapping expression patterns in cochlea and by the detection of heteromeric Cx26/Cx31 connexons, using coimmunoprecipitation of mouse cochlear membrane proteins. Furthermore, in vitro cotransfection of the two-tagged connexin proteins showed that they were able to co-assemble in vitro in the same junction plaque Liu et al.86
DFNA5
DFNA5 was originally mapped to chromosome 7p15 in an extended Dutch kindred with autosomal dominant progressive hearing loss starting in the high frequencies. With increasing age the middle and low frequencies also became affected.87 To date, five mutations have been reported in the DFNA5 gene. Four of them caused skipping of exon 8, resulting in premature termination of the open reading frame. All four mutations segregated with late-onset non-syndromic hearing impairment in the families.88, 89, 90 Among the mutations identified, a CTT deletion in the polypyrimidine tract of intron 7 and an IVS8+4A → G transition in the splice-donor site of intron 8 have been described in Chinese families with non-syndromic deafness, indicating that mutations in DFNA5 may be major known causes of hereditary hearing loss in Chinese patients.89 A truncating mutation, c.640insC, in exon 5 of DFNA5 was recently reported in an Iranian family. Interestingly, the mutation is not a disease-causing mutation, and was even found present in normal-hearing subjects in this family.91 This finding may be regarded as support to the previously formulated hypothesis that only exon 8 skipping can lead to DFNA5-associated hearing loss. Recent studies suggest that DFNA5 belongs to a novel GSDMDC family. As different mutations in the DFNA5 gene have been shown to result in hearing loss, it is tempting to speculate that this gene may play some crucial role in the physiological process of hearing.
DFNA9
The DFNA9 locus on chromosome 14q12–q13 is known to be associated with vestibular dysfunction.92 The cochlin, encoded by COCH is an extracellular matrix protein that contains a region homologous to a domain in factor C of Limulus, also known as LCCL domain and two von Willebrand factor A-like domains (vWFA1 and 2).93 Interestingly, the majority of the previously described COCH mutations are missense point mutations located within the LCCL domain of cochlin.94, 95, 96, 97, 98, 99 They have been associated with progressive hearing loss and concomitant vestibular symptoms. Street et al.100 reported a p.C542F mutation in an American DFNA9 family with hearing loss as well as oculomotor and vestibular disturbances. This mutation located within the vWFA2 domain of cochlin, representing the first reported DFNA9 mutation outside of the LCCL domain. We subsequently identified a different mutation, p.C542Y, at the same location in vWFA2 domain in a large Chinese family with late-onset autosomal dominant non-syndromic progressive sensorineural hearing loss. In addition, a heterozygous missense mutation (p.M512T) at an evolutionarily conserved methionine residue was also identified in the vWFA2 domain in a small family.101 More importantly, none of the C542F, C542Y and M512T mutation carriers had vestibular complaints (dizziness, vertigo, head movement-dependent oscillopsia and instability in the dark) in their lifetime.100, 101 A genotype–phenotype correlation could thus be established when comparing the vestibular involvement between the DFNA9 families harboring mutations in the LCCL domain and in the vWFA2 domain.
DFNA11
Mutations in Myosin VIIA have been found to be responsible for the DFNA11 deafness. Initially, an in-frame 9-bp deletion in exon 22 of MYO7A was identified to cause DFNA11 in a Japanese family.61 The affected members suffered from postlingual, bilateral, non-syndromic sensorineural hearing loss, with gradual progression at all frequencies and minor vestibular problems. No evidence was found for retinitis pigmentosa.102 So far, the DFNA11 families have been identified in other populations including USA, Italy, Dutch and Germany.103 Recently, a DFNA11 family with late-onset hearing loss ranging from 20 to 47 years old was reported, involving the high-frequency first. The locus was mapped in a Chinese pedigree with an autosomal dominant non-syndromic hearing loss. A maximum two point LOD score of 5.71 at θ=0.05 on chromosome 11q, was obtained for marker D11S937. The analysis of recombinant haplotypes mapped the locus between markers (D11S165–D11S1874).104 A missense mutation at the motor region has been recently identified in this Chinese DFNA11 family (Yuan et al., unpublished data). Interestingly, all mutations involved in the DFNA11 families are located at the motor domain of the Myosin VIIA protein.
DFNA39
Xiao et al.105 described two Chinese DFNA39 families, characterized by the combination of bilateral progressive high-frequency sensorineural hearing loss, with dentinogenesis imperfecta-1 (DGI1). DGI1 is an autosomal dominant dental disease associated with abnormal dentin production and mineralization. Distinct mutations within the dentin sialophosphoprotein gene have been found responsible for the clinical manifestations of Dentinogenesis imperfecta 1 with or without DFNA39.
Mitochondrial DNA mutations
Mitochondrial (mt) pathology has been implicated in both inherited and acquired hearing loss. Sensorineural hearing loss (SNHL) is present in 42–70% of individuals with mt cytopathies and can be syndromic and non-syndromic.106, 107, 108 Mutations in mtDNA have also been found to be associated with both aminoglycoside-induced and non-syndromic deafness.108, 109 The matrilineal transmission of aminoglycoside-induced deafness in the Chinese population was first reported in isolated pedigrees in the early 1990s and mutations in the mtDNA were suggested as the likely cause.30, 110 The most commonly used ototoxic drugs in China are aminoglycoside antibiotics including streptomycin, gentamycin and kanamycin. Aminoglycoside antibiotics were widely, and in many cases over-used nationwide in China from the 1960s to 1980s, as they were considered cost-effective in controlling infections by a wide spectrum of bacteria. Despite the large number of reports on aminoglycoside-induced hearing loss in China and in other countries, aminoglycosides and a few new synthetic derivatives from traditional aminoglycosides, such as amikacin, netilmicin, etimicin, isepamicin, are still being used in clinics all over China, especially in low-income areas including mountainous regions. Different prevalences of these predisposing mutations might contribute to the high incidence of aminoglycoside ototoxicity in certain regions of China. The incidence of aminoglycoside-induced deafness has increased in recent years in China. Some studies of deaf-school populations have indicated that aminoglycoside antibiotics may account for 13–66% of profound deafness with a prevalence of 0.035%.111 Our data shows that 88% of severe-profound mtDNA deafness carrying the homoplasmic A1555G mutation had exposure to aminoglycosides.112
The mitochondrial 12S rRNA (MTRNR1) appears to be a hot spot for mtDNA mutations. The MTRNR1 mutations likely alter the secondary structure of the 12S rRNA molecule to make it more similar to its bacterial counterpart, the 16S rRNA, the target of aminoglycoside.113 This might explain the use of aminoglycosides and the cumulating effect of these MTRNR1 mutations. Sequence analyses of the mitochondrial genome in patients with aminoglycoside ototoxicity have led to the identification of several ototoxic mtDNA mutations in the 12S rRNA gene. The homoplasmic A1555G and mutations at position 961 of the 12S rRNA gene have been implicated with aminoglycoside-induced and non-syndromic hearing loss in many families of different ethnic origins.29, 114, 115, 116, 117, 30, 110, 118, 119, 37, 120 Similarly, the homoplasmic C1494T mutation in the highly conserved decoding site of the same gene has been associated with aminoglycoside-induced and non-syndromic hearing loss in a large Chinese family.121 Furthermore, the 961 C-insertion and the A827G mutation have been reported to be linked to aminoglycoside ototoxicity or non-syndromic hearing loss,30, 110, 119, 37, 120 and possibly the T1005C and A1116G mutations can contribute to ototoxic hearing loss.111 Another mutation, the T1095C in the 12S rRNA gene has been associated with hearing impairment in several genetically unrelated families including Italian122, 123 and Chinese deaf patients.111, 121
In the familial cases of aminoglycoside ototoxity, these known predisposing mutations in the 12S rRNA gene account for a significant proportion of cases.111, 114 The A1555G mutation has been found in high frequency compared to other mutations in the sporadic populations with aminoglycoside ototoxity. The prevalence of the A1555G mutation varies in hearing-impaired populations from different ethnic origins. The incidence of A1555G mutation in non-syndromic deafness is much lower compared with those with aminoglycoside ototoxity. In Asian non-syndromic hearing-impaired patients, the prevalence of the A1555G mutation seems to be higher than in Caucasians: 2.9% in Chinese,111 3% in Japanese,124 and 5.3% in Indonesian.125 There was variation in the incidence of the A1555G mutation in Caucasian deaf populations: 0.6% in Cincinnati white patients,30, 110 1.8 and 0.7% in Hungarian and Germany populations, respectively,126 1.8% in Turkish pediatric population,127 2.4% in a Danish cohort 128 and 2.5% in a UK cohort.129 Li et al.130 conducted a systematic mutational screening of the mitochondrial 12S rRNA gene in a Chinese pediatric population with sporadic aminoglycoside-induced and NSHL.131 They showed that aminoglycoside ototoxicity is responsible for 48% of cases. Mutations at position 961 in the MTRNR1 gene accounted for ∼1.7 and 4.4% cases of aminoglycoside-induced and NSHL, respectively in this Chinese clinical population. The frequency of the A1555G mutation is ∼13 and ∼2.9% in these Chinese pediatric subjects with aminoglycoside-induced and non-syndromic hearing loss, respectively. Our recent study shows that the mtDNA A1555G is responsible for deafness for a large proportion of NSHL, accounting for 15.5% of SNHL cases from the special clinics for deaf in China.112
The A1555G mutation is thought to be a susceptibility mutation, which needs the combined action of environmental factors and/or nuclear modifying genes to cause hearing impairment.130, 132, 133, 134, 135 Variants in mitochondrial tRNAGlu A14693G, tRNAThr T15908C, tRNAArg T10454C, tRNASer(UCN) G7444A and tRNACys G5821A may influence the phenotypic manifestation of deafness-associated 12S rRNA A1555G mutation.110, 118, 136, 38, 137 Interestingly, the tRNACys T5802C, tRNAThr A15924C and ND5 T12338C alterations have been shown to have a potential modifier role in increasing the penetrance and expressivity of the deafness-associated 12S rRNA A1555G mutation in Han-affected Chinese patients.138 Furthermore, Liao et al.139 reported that the ND4 G11696A mutation may act in synergy with the primary deafness-associated 12S rRNA A1555G mutation in a four-generation Chinese family, thereby increasing the penetrance and expressivity of hearing loss in this Chinese pedigree.
The mitochondrial tRNASer(UCN) (MTTS1) appears to be another hot spot for mtDNA mutations associated with NSHL. Five deafness-associated mutations, A7445G,107, 140, 141 A7445C142 7472insC,143, 144 T7510C145 and T7511C146, 147 have been identified in this gene.
Non-syndromic loci with unknown specific genes DFNA41/42/53
There are also genes for autosomal dominant and recessive as well as X or Y-linked hereditary deafness that is yet to be cloned in the Chinese population. We have previously reported the mapping of the DFNA41 and DFNA53,148, 149, 150 located on chromosome 12q24-qter and on 14q11.2–q12 respectively, in large multi-generational Chinese families. Another locus (DFNA42) for autosomal dominant non-syndromic hearing loss identified at 5q31.1–32 has also been reported in a Chinese pedigree.151 Finally, an X-linked recessive deafness syndrome with muteness as well as Y-linked inheritance of non-syndromic hearing impairment has also been described in Chinese deaf families.152, 153
Concluding remarks
We have described in this review genes and spectrum of the mutations found in the Chinese deafness population. So far, mutations in GJB2, GJB3, Myo7A, SLC26A4 and the A1555G mitochondrial mutation were found to be the major causes of hearing loss in Chinese deaf patients. It has been shown that the 235delC mutation in the GJB2 gene and IVS7-2A>G (c.919-2A>G) in the SLC26A4 gene are common founder mutations, thus explaining their frequency. Mutations in the SLC26A4 gene are second to GJB2 mutations as a currently identifiable genetic cause of SNHL in China.23, 78 Differences in spectrum of the GJB3 and Myo7A mutations have also been described in Chinese cases of SNHL. Genetic testing is now available for a number of genes involved in hearing loss. However, only with the knowledge of prevalent population-specific mutations, can an appropriate and effective strategy for detecting deafness genes be developed. This will facilitate precise clinical diagnosis, accurate genetic counseling and will provide a framework for the implementation and assessment of any future gene therapy.
References
Fraser, G. R. The Causes of Profound Deafness in Childhood (Johns Hopkins University Press: Baltimore, 1976).
Morton, N. E. Genetic epidemiology of hearing impairment. Ann. N. Y. Acad. Sci. 630, 16–31 (1991).
Marazita, M. L., Ploughman, L. M., Rawlings, B., Remington, E. & Nance, W. E. Genetic epidemiological studies of early-onset deafness in the US school-age population. Am. J. Med. Genet. 46, 486–491 (1993).
ACMG. Genetics evaluation guidelines for the etiologic diagnosis of congenital hearing loss. Genetic Evaluation of Congenital Hearing Loss Expert Panel: ACMG statement. Genet Med. 4, 162–171 (2002).
Hone, S. W. & Smith, R. J. Medical evaluation of pediatric hearing loss: laboratory, radiographic, and genetic testing. Otolaryngol. Clin. North Am. 35, 751–764 (2002).
Fischel-Ghodsian, N., Blanchard, S., Kaplan, J., Guilford, P., Gibson, F., Walsh, J. et al. Mitochondrial RNA processing and translation: link between mitochondrial mutations and hearing loss? Mol. Genet. Metab. 65, 97–104 (1998).
Weil, D., Blanchard, S., Kaplan, J., Guilford, P., Gibson, F., Walsh, J. et al. Defective myosin VIIA gene responsible for Usher syndrome type 1B. Nature. 374, 60–61 (1995).
Burchard, E. G., Ziv, E., Coyle, N., Gomez, S. L., Tang, H., Karter, A. J. et al. The importance of race and ethnic background in biomedical research and clinical practice. N. Engl. J. Med. 348, 1170–1175 (2003).
Whitlock, M. C. Genetic drift. In Encyclopedia of Evolution (ed. Pagel, M.), 400–403 (Oxford University Press, USA, 2002).
Rathore, S. S. & Krumholz, H. M. Race, ethnic group, and clinical research. BMJ 327, 763–764 (2003), doi:10.1136/bmj.327.7418.763.
Ben-Yosef, T. & Friedman, T. B. The genetic bases for syndromic and nonsyndromic deafness among Jews. Trends Mol. Med. 9, 496–502 (2003).
Usami, S., Wagatsuma, M., Fukuoka, H., Suzuki, H., Tsukada, K., Nishio, S. et al. The responsible genes in Japanese deafness patients and clinical application using Invader assay. Acta. Otolaryngol. 128, 446–454 (2008).
Cohn, E. S. & Kelley, P. M. Clinical phenotype and mutations in connexin 26 (DFNB1/GJB2), the most common cause of childhood hearing loss. Am. J. Med. Genet. 89, 130–136 (1999).
Fuse, Y., Doi, K., Hasegawa, T., Sugii, A., Hibino, H. & Kubo, T. Three novel connexin26 gene mutations in autosomal recessive nonsyndromic deafness. Neuroreport. 10, 1853–1857 (1999).
Kelley, P. M., Harris, D. J., Comer, B. C., Askew, J. W., Fowler, T., Smith, S. D et al. Novel mutations in the connexin 26gene (GJB2) that cause autosomal recessive (DFNB1) hearing loss. Am. J. Hum. Genet. 62, 792–799 (1998).
Gasparini, P., Estivill, X., Volpini, V., Totaro, A., Castellvi-Bel, S., Govea, N. et al. Linkage of DFNB1 to nonsyndromic neurosensory autosomal recessive deafness in Mediterranean families. Eur. J. Hum. Genet. 5, 83–88 (1997).
Green, G. E., Scott, D. A., McDonald, J. M., Woodworth, G. G., Sheffield, V. C. & Smith, R. J. Carrier rates in the midwestern United States for GJB2 mutation causing inherited deafness. JAMA 28, 2211–2216 (1999).
Sobe, T., Vreugde, S., Shahin, H., Berlin, M., Davis, N. & Kanaan, M. The prevalence and expression of inherited connexin 26 mutations associated with nonsyndromic hearing loss in the Israeli population. Hum. Genet. 106, 50–57 (2000).
Morell, R. J., Kim, H. J., Hood, L. J., Goforth, L., Friderici, K. & Fisher, R. Mutations in the connexin 26 gene (GJB2) among Ashkenazi Jews with nonsyndromic recessive deafness. N. Engl. J. Med. 339, 1500–1505 (1998).
Cryns, K., Orzan, E., Murgia, A., Huygen, P. L. M., Moreno, F., del Castillo, I. et al. A genotype-phenotype correlation for GJB2 (connexin 26) deafness. Med. Genet. 41, 147–154 (2004).
Hilgert, N., Huentelman, M. J., Thorburn, A. Q., Fransen, E., Dieltjens, N., Mueller-Malesinska, M. et al. Phenotypic variability of patients homozygous for the GJB2 mutation 35delG cannot be explained by the influence of one major modifier gene. Eur. J. Hum. Genet. doi:10.1038/ejhg.2008.201 (2008).
Liu, X. Z., Xia, X. J., Ke, X. M., Ouyang, X. M., Du, L. L., Liu, Y. H. et al. The prevalence of connexin 26 (GJB2) mutations in the Chinese population. Hum. Genet. 111, 394–397 (2002).
Dai, P., Yu, F., Han, B., Yuan, Y., Li, Q., Wang, G. et al. The prevalence of the 235delC GJB2 mutation in a Chinese deaf population. Genet. Med. 9, 283–289 (2007).
Liu, X. Z., Walsh, J., Tamagawa, Y., Kitamura, K., Nishizawa, M., Steel, K. P. et al. Autosomal dominant non-syndromic deafness caused by a mutation in the myosin VIIA gene. (Letter). Nat. Genet. 17, 268–269 (1997a).
Hu, H., Wu, L., Feng, Y., Pan, Q., Long, Z., Li, J. et al. Molecular analysis of hearing loss associated with enlarged vestibular aqueduct in the mainland Chinese: a unique SLC26A4 mutation spectrum. J. Hum. Genet. 52, 492–497 (2007).
Wang, Q. J., Zhao, Y. L., Rao, S. Q., Guo, Y. F., Yuan, H., Zong, L. et al. A distinct spectrum of SLC26A4 mutations in patients with enlarged vestibular aqueduct in China. Clin. Genet. 72, 245–254 (2007).
Wu, C. C., Yeh, T. H., Chen, P. J. & Hsu, C. J. Prevalent SLC26A4mutations in patients with enlarged vestibular aqueduct and/or Mondini dysplasia: a unique spectrum of mutations in Taiwan, including a frequent founder mutation. Laryngoscope. 115, 1060–1064 (2005).
Tsukamoto, K., Suzuki, H., Harada, D., Namba, A., Abe, S. & Usami, S. Distribution and frequencies of PDS (SLC26A4) mutations in Pendred syndrome and nonsyndromic hearing loss associated with enlarged vestibular aqueduct: a unique spectrum of mutations in Japanese. Eur. J. Hum. Genet. 11, 916–922 (2003).
Hutchin, T., Haworth, I., Higashi, K., Fischel-Ghodsian, N., Stoneking, M., Saha, N. et al. A molecular basis for human hypersensitivity to aminoglycoside antibiotics. Nucleic Acids Res. 21, 4174–4179 (1993).
Li, R., Greinwald, J. H., Yang, L., Choo, D. I., Wenstrup, R. J. & Guan, X. M. Molecular analysis of mitochondrial 12S rRNA and tRNASer(UCN) genes in paediatric subjects with nonsyndromic hearing loss. J. Med. Genet. 41, 615–620 (2004).
Yong, A. M., Goh, S. S., Zhao, Y., Eng, P. H., Koh, L. K. & Khoo, D. H. Two Chinese families with Pendred’s syndrome—radiological imaging of the ear and molecular analysis of the pendrin gene. J. Clin. Endocrinol. Metab. 86, 3907–3911 (2001).
Park, H. J., Shaukat, S., Liu, X. Z., Hahn, S. H., Naz, S., Ghosh, M. et al. Origins and frequencies of SLC26A4 (PDS) mutations in east and south Asians: global implications for the epidemiology of deafness. J. Med. Genet. 40, 242–248 (2003).
Fugazzola, L., Cerutti, N., Mannavola, D., Crino, A., Cassio, A., Gasparoni, P. et al. Differential diagnosis between Pendred and pseudo-Pendred syndromes: clinical, radiologic, and molecular studies. Pediatr. Res. 51, 479–484 (2002).
Yang, J. J., Tsai, C. C., Hsu, H. M., Shiao, J. Y., Su, C. C. & Li, S. Y. Hearing loss associated with enlarged vestibular aqueduct and Mondini dysplasia is caused by splice-site mutation in the PDS gene. Hear Res. 199, 22–30 (2005).
Grifa, A., Wagner, C. A., D′Ambrosio, L., Melchionda, S., Bernardi, F., Lopez-Bigas, N. et al. Mutations in GJB6 cause nonsyndromic autosomal dominant deafness at DFNA3 locus. Nat. Genet. 23, 16–18 (1999).
Xia, J. H., Liu, C. Y., Tang, B. S., Pan, Q., Huang, L., Dai, H. P. et al. Mutations in the gene encoding gap junction protein beta-3 associated with autosomal dominant hearing impairment. Nat. Genet. 20, 370–373 (1998).
Casano, R. A., Johnson, D. F., Bykhovskaya, Y., Torricelli, F., Bigozzi, M. & Fischel-Ghodsian, N. Inherited susceptibility to aminoglycoside ototoxicity: genetic heterogeneity and clinical implications. Am. J. Otolaryngol. 20, 151–156 (1999).
Yuan, H., Qian, Y., Xu, Y., Cao, J., Bai, L., Shen, W et al. Cosegregation of the G7444A mutation in the mitochondrial COI/tRNA(Ser(UCN)) genes with the 12S rRNA A1555G mutation in a Chinese family with aminoglycoside-induced and nonsyndromic hearing loss. Am. J. Med. Genet. 138, 133–140 (2005).
Dai, P., Yu, F., Kang, D. Y., Zhang, X., Liu, X., Mi, W. Z. et al. Diagnostic methods and clinic application for mtDNA A1555G and GJB2 and SLC26A4 genes in deaf patients. Zhonghua Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 40, 769–773 (2005) (in Chinese).
Zhang, X. Y., Zhang, S. L., Ke, B. S., Jiang, Z. S. & Sun, R. Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 21, 168–170 (2004) (in Chinese).
Van Laer, L., Coucke, P., Mueller, R. F., Caethoven, G., Flothmann, K., Prasad, S. D. et al. A common founder for the 35delG GJB2 gene mutation in connexin 26 hearing impairment. J. Med. Genet. 38, 515–518 (2001).
Abe, S., Usami, S., Shinkawa, H., Kelley, P. M. & Kimberling, W. J. Prevalent connexin 26 gene (GJB2) mutations in Japanese. J. Med. Genet. 37, 41–43 (2000).
Kudo, T., Ikeda, K., Kure, S., Matsubara, Y., Oshima, T., Watanabe, K. et al. Novel mutations in the connexin 26 gene (GJB2) responsible for childhood deafness in the Japanese population. J. Med. Genet. 90, 141–145 (2000).
Park, H. J., Hahn, S. H., Chun, Y. M., Park, K. & Kim, H. N. Connexin26 mutations associated with nonsyndromic hearing loss. Laryngoscope. 110, 1535–1538 (2000).
Kelsell, D. P., Dunlop, J., Stevens, H. P., Lench, N. J., Liang, N., Parry, G. et al. Cx26 mutation in hereditary nonsyndromic sensorineural deafness. Nature 387, 80–83 (1997).
Scott, D. A. & Kraft, M. L. Connexin mutation and hearing loss. Nature 391, 32 (1998).
Ohtsuka, A., Yuge, I., Kimura, S., Namba, A., Abe, S., Van Laer, L. et al. GJB2 deafness gene shows a specific spectrum of mutations in Japan, including a frequent founder mutation. Hum. Genet. 112, 329–333 (2003).
Yan, D., Park, H. J., Ouyang, X. M., Pandya, A., Doi, K., Erdenetungalag, R. et al. Evidence of a founder effect for the 235delC mutation of GJB2 (connexin 26) in east Asians. Hum. Genet. 114, 44–50 (2003).
Del Castillo, I., Moreno-Pelayo, M. A., Del Castillo, F. J., Brownstein, Z., Marlin, S., Adina, Q. et al. Prevalence and evolutionary origins of the del (GJB6-D13S1830) mutation in the DFNB1 locus in hearing impaired subjects: a multicenter study. Am. J. Hum. Genet. 73, 1452–1458 (2003).
Gualandi, F., Ravani, A., Berto, A., Burdo, S., Trevisi, P., Ferlini, A. et al. Occurrence of del (GJB6-D13S1830) mutation in Italian non-syndromic hearing loss patients carrying a single GJB2 mutated allele. Acta. Otolaryngol. Suppl. 552, 29–34 (2004).
Bolz, H., Schade, G., Ehmer, S., Kothe, C. & Hess, M. G. A Phenotypic variability of non-syndromic hearing loss in patients heterozygous for both c.35delG of GJB2 and the 342-kb deletion involving GJB6. Hear. Res. 188, 42–46 (2004).
Gunther, B., Steiner, A., Nekahm-Heis, D., Albegger, K., Zorowka, P., Utermann, G. & Janecke, A. The 342-kb deletion in GJB6 is not present in patients with non-syndromic hearing loss from Austria. Hum. Mutat. 22, 180 (2003).
Tekin, M., Duman, T., Bo∂oçlu, G., Incesulu, A., Comak, E., Ilhan, I. & Akar, N. Spectrum of GJB2 mutations in Turkey comprises both Caucasian and Oriental variants: roles of parental consanguinity and assortative mating. Hum. Mutat. 21, 552–553 (2003).
Uyguner, O., Emiroglu, M., Uzumcu, A., Hafiz, G., Ghanbari, A., Baserer, N. et al. Frequencies of gap- and tight-junction mutations in Turkish families with autosomal-recessive non-syndromic hearing loss. Clin. Genet. 64, 65–69 (2003).
Liu, Y., Ke, X., Qi, Y., Li, W. & Zhu, P. Connexin26 gene (GJB2): prevalence of mutations in the Chinese population. J. Hum. Genet. 47, 688–690 (2002).
Yuan, Y., Huang, D., Dai, P., Zhu, Q., Liu, X., Wang, G. et al. GJB6 gene mutation analysis in Chinese nonsyndromic deaf population. Lin Chuan Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 21, 3–6 (2007) (in Chinese)..
Yuan, Y., Huang, D., Dai, P., Zhu, X., Yu, F., Zhang, X. et al. Mutation analysis of GJB2, GJB3 and GJB6 gene in deaf population from special educational school of Chifeng city. Lin Chuan Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 22, 14–17, 21. (2008)(in Chinese).
Riazalhosseini, Y., Nishimura, C., Kahrizi, K., Shafeghati, Y., Daneshi, A. & Jogataie, M. T. Delta (GJB6-D13S1830) is not a common cause of nonsyndromic hearing loss in the Iranian population. Iranian Med. 8, 104–108 (2005).
Liu, X. Z., Hope, C., Walsh, J., Newton, V., Ke, X. M., Liang, C. Y. et al. Mutations in the myosin VIIA gene cause a wide phenotypic spectrum, including atypical Usher syndrome. Am. J. Hum. Genet. 63, 909–912 (1998).
Zina, Z. B., Masmoudi, S., Ayadi, H., Chaker, F., Ghorbel, A. M., Drira, M. et al. From DFNB2 to Usher syndrome: Variable expressivity of the same disease. Am. J. Med. Genet. 101, 181–183 (2001).
Liu, X. Z., Walsh, J., Mburu, P., Kendrick-Jones, J., Cope, M. J., Steel, K. P. et al. Mutations in the myosin VIIA gene cause non-syndromic recessive deafness. Nat. Genet. 16, 188–190 (1997b).
Riazuddin, S., Nazli, S., Ahmed, Z. M., Yang, Y., Zulfiqar, F., Shaikh, R. S. et al. Mutation Spectrum of MYO7A and Eveluation of a Novel Nonsyndromic Deafness DFNB2 Allele with Residual Function. Hum. Mutat. 29, 502–511 (2008).
Fraser, G. R. Association of congenital deafness with goitre (Pendred’s syndrome): a study of 207 families. Ann. Hum. Genet. 28, 201–249 (1965).
Everett, L. A., Glaser, B., Beck, J. C., Idol, J. R., Buchs, A., Heyman, M. et al. Pendred syndrome is caused by mutations in a putative sulphate transporter gene (PDS). Nat. Genet. 17, 411–422 (1997).
Li, X. C., Everett, L. A., Lalwani, A. K., Desmukh, D., Friedman, T. B., Green, E. D. et al. A mutation in PDS causes non-syndromic recessive deafness. Nat. Genet. 18, 215–217 (1998).
Scott, D. A., Wang, R., Kreman, T. M., Andrews, M., McDonald, J. M., Bishop, J. R. et al. Functional differences of the PDS gene product are associated with phenotypic variation in patients with Pendred syndrome and non-syndromic hearing loss (DFNB4). Hum. Mol. Genet. 9, 1709–1715 (2000).
Usami, S., Abe, S., Weston, M. D., Shinkawa, H., Van Camp, G. & Kimberling, W. J. Non-syndromic hearing loss associated with enlarged vestibular aqueduct is caused by PDS mutations. Hum. Genet. 104, 188–192 (1999).
Harada, D., Namba, A., Abe, S. & Usami, S. Distribution and frequencies of PDS (SLC26A4) mutations in Pendred syndrome and nonsyndromic hearing loss associated with enlarged vestibular aqueduct: a unique spectrum of mutations in Japanese. Eur. J. Hum. Genet. 11, 916–922 (2003).
Park, H. J., Lee, S. J., Jin, H. S., Lee, J. O., Go, S. H., Jang, H. S., Moon, S. K. et al. Genetic basis of hearing loss associated with enlarged vestibular aqueducts in Koreans. Clin. Genet. 67, 160–165 (2005).
Campbell, C., Cucci, R. A., Prasad, S., Green, G. E., Edeal, J. B., Galer, C. E et al. Pendred syndrome, DFNB4, and PDS/SLC26A4 identification of eight novel mutations and possible genotype-phenotype correlations. Hum. Mutat. 17, 403–411 (2001).
Pera, A., Dossena, S., Rodighiero, S., Gandía, M., Bottà, G., Meyer, G. et al. Functional assessment of allelic variants in the SLC26A4 gene involved inPendred syndrome and nonsyndromic EVA. Proc. Natl Acad. Sci. USA. 105, 18608–18613 (2008).
Coucke, P. J., Van Hauwe, P., Everett, L. A., Demirhan, O., Kabakkaya, Y., Dietrich, N. L. et al. Identification of two different mutations in the PDS gene in an inbred family with Pendred syndrome. J. Med. Genet. 36, 475–477 (1999).
Gonzalez Trevino, O., Karamanoglu Arseven, O., Ceballos, C. J., Vives, V. I., Ramirez, R. C., Gomez, V. V et al. Clinical and molecular analysis of three Mexican families with Pendred′s syndrome. Eur. J. Endocrinol. 144, 585–593 (2001).
López-Bigas, N., Olivé, M., Rabionet, R., Ben-David, O., Martínez-Matos, J. A., Bravo, O. et al. Connexin 31 (GJB3) is expressed in the peripheral and auditory nerves and causes neuropathy and hearing impairment. Hum. Mol. Genet. 10, 947–952 (2000).
Pryor, S. P., Madeo, A. C., Reynolds, J. C., Sarlis, N. J., Arnos, K. S., Nance, W. E. et al. SLC26A4/PDS genotype-phenotype correlation in hearing loss with enlargement of the vestibular aqueduct (EVA): evidence that Pendred syndrome and non-syndromic EVA are distinct clinical and genetic entities. J. Med. Genet. 42, 159–165 (2005).
López-Bigas, N., Olivé, M., Rabionet, R., Ben-David, O., Martínez-Matos, J. A., Bravo, O. et al. Connexin 31 (GJB3) is expressed in the peripheral and auditory nerves and causes neuropathy and hearing impairment. Hum. Mol. Genet. 10, 947–952 (2001).
Blons, H., Feldmann, D., Duval, V., Messaz, O., Denoyelle, F., Loundon, N. et al. Screening of SLC26A4 (PDS) gene in Pendred’s syndrome: a large spectrum of mutations in France and phenotypic heterogeneity. Clin. Genet. 66, 333–340 (2004).
Dai, P., Li, Q., Huang, D., Yuan, Y., Kang, D., Miller, D. T. et al. SLC26A4 c.919-2A>G varies among Chinese ethnic groups as a cause of hearing loss. Genet. Med. 10, 586–592 (2008).
Liu, X. Z., Xia, X. J., Xu, L. R., Pandya, A., Liang, C. Y., Blanton, S. H. et al. Mutations in connexin31 underlie recessive as well as dominant non-syndromic hearing loss. Hum. Mol. Genet. 9, 63–67 (2000).
Alexandrino, F., Oliveira, C. A., Reis, F. C., Maciel-Guerra, A. T. & Sartorato, E. L. Screening for mutations in the GJB3 gene in Brazilian patients with nonsyndromic deafness. J. Appl. Genet. 45, 249–254 (2004).
Coucke, P., Van Camp, G., Djoyodiharjo, B., Smith, S. D., Frants, R. R., Padberg, G. W. et al. Linkage of autosomal dominant hearing loss to the short arm of chromosome 1 in two families. N. Engl. J. Med. 331, 425–431 (1994).
Van Camp, G., Coucke, P. J., Kunst, H., Schatteman, I., Van Velzen, D., Marres, H. et al. Linkage analysis of progressive hearing loss in five extended families maps the DFNA2 gene to a 1.25-Mb region on chromosome 1p. Genomics. 41, 70–74 (1997).
Van Hauwe, P., Coucke, P. & Van Camp, G. The DFNA2 locus for hearing impairment: two genes regulating K+ ion recycling in the inner ear. Br. J. Audiol. 33, 285–289 (1999).
Mhatre, A. N., Weld, E. & Lalwani, A. K. Mutation analysis of Connexin 31 (GJB3) in sporadic non-syndromic hearing impairment. Clin. Genet. 63, 154–159 (2003).
Frei, K., Ramsebner, R., Hamader, G., Lucas, T., Schoefer, C., Baumgartner, W. D. et al. Lack of association between Connexin 31 (GJB3) alterations and sensorineural deafness in Austria. Hear Res. 194, 81–86 (2004).
Liu, X. Z., Yuan, Y., Yan, D., Ding, E. H., Ouyang, X. M., Fei, Y. et al. Digenic inheritance of non-syndromic deafness caused by mutations at the gap junction proteins Cx26 and Cx31. Hum. Genet. 125, 53–62 (2009).
Van, G. C., Paul, C., Balemans, W., van Velzen, D., van de Bilt, C., Van Laer, L. et al. Localization of a gene for non-syndromic hearing loss (DFNA5) to chromosome 7p15. Hum. Mol. Genet. 4, 2159–2163 (1995).
Van Laer, L., Huizing, E. H., Verstreken, M., van Zuijlen, D., Wauters, J. G., Bossuyt, P. J. et al. Nonsyndromic hearing impairment is associated with a mutation in DFNA5. Nat. Genet. 20, 194–197 (1998).
Yu, C., Meng, X., Zhang, S., Zhao, G., Hu, L. & Kong, X. A 3-nucleotide deletion in the polypyrimidine tract of intron 7 of the DFNA5 gene causes nonsyndromic hearing impairment in a Chinese family. Genomics 82, 575–579 (2003).
Bischoff, A. M., Luijendijk, M. W., Huygen, P. L., van Duijnhoven, G., De Leenheer, E. M., Oudesluijs, G. G et al. A novel mutation identified in the DFNA5 gene in a Dutch family: a clinical and genetic evaluation. Audiol. Neurootol. 9, 34–46 (2004).
Van Laer, L., Meyer, N. C., Malekpour, M., Riazalhosseini, Y., Moghannibashi, M., Kahrizi, K. et al. A novel DFNA5 mutation does not cause hearing loss in an Iranian family. J. Hum. Genet. 52, 549–552 (2007).
Fransen, E. & Van Camp, G. The COCH gene: a frequent cause of hearing impairment and vestibular dysfunction? Br. J. Audiol. 33, 297–302 (1999).
Trexler, M., Bányai, L. & Patthy, L. The LCCL module. Eur. J. Biochem. 267, 5751–5757 (2000).
Robertson, N. G., Lu, L., Heller, S., Merchant, S. N., Eavey, R. D., McKenna, M. et al. Mutations in a novel cochlear gene cause DFNA9, a human nonsyndromic deafness with vestibular dysfunction. Nat. Genet. 20, 299–303 (1998).
de Kok, Y. J. M., Bom, S. J. H., Brunt, T. M., Kemperman, M. H., van Beusekom, E., van der Velde-Visser, S. D. et al. A Pro51Ser mutation in the COCH gene is associated with late onset autosomal dominant progressive sensorineural hearing loss with vestibular defects. Hum. Mol. Genet. 8, 361–366 (1999).
Kamarinos, M., McGill, J., Lynch, M. & Dahl, H. Identification of a novel COCH mutation, I109N, highlights the similar clinical features observed in DFNA9 families. Hum. Mutat. 17, 351–355 (2001).
Nagy, I., Horvath, M., Trexler, M., Repassy, G. & Patthy, L. A novel COCH mutation, V104del, impairs folding of the LCCL domain of cochlin and causes progressive hearing loss. J. Med. Genet. 41, e9 (2004).
Usami, S., Takahashi, K., Yuge, I., Ohtsuka, A., Namba, A., Abe, S. et al. Mutations in the COCH gene are a frequent cause of autosomal dominant progessive cochleo-vestibular dysfuntion, but not of Meniere’s disease. Eur. J. Hum. Genet. 11, 744–748 (2003).
Collin, R. W., Pauw, R. J., Schoots, J., Huygen, P. L., Hoefsloot, L. H., Cremers, C. W et al. Identification of a novel COCH mutation, G87W, causing autosomal dominant hearing impairment (DFNA9). Am. J. Med. Genet. A. 140, 1791–1794 (2006).
Street, V. A., Kallman, J. C., Robertson, N. G., Kuo, S. F., Morton, C. C. & Phillips, J. O. A novel DFNA9 mutation in the vWFA2 domain of COCH alters a conserved cysteine residue and intrachain disulfide bond formation resulting in progressive hearing loss and site-specific vestibular and central oculomotor dysfunction. Am. J. Med. Genet. 139A, 86–95 (2005).
Yuan, H. J., Han, D. Y., Sun, Q., Sun, H. J., Tao, R., Yan, D. et al. Absence of vestibular symptoms in two Chinese families with novel mutations in the vWFA2 domain of COCH: evidence of genotype-phenotype correlation in DFNA. Clin. Genet. 73, 391–394 (2008).
Tamagawa, Y., Ishikawa, K., Ishikawa, K., Ishida, T., Kitamura, K., Makino, S. et al. Clinical presentation of DFNA11 (MYO7A). Adv. Otorhinolaryngol. 61, 79–84 (2002).
Luijendijk, M. W., Van Wijk, E., Bischoff, A. M., Krieger, E., Huygen, P. L., Pennings, R. J et al. Identification and molecular modelling of a mutation in the motor head domain of myosin VIIA in a family with autosomal dominant hearing impairment (DFNA11). Hum. Genet. 115, 149–156 (2004).
Yuan, H., Han, D. Y., Wang, Q. J., Zong, L. & Zhao, Y. L. Gene mapping for autosomal dominant nonsyndromic hearing loss DFNA11. Zhonghua Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 42, 422–427 (2007) (in Chinese)..
Xiao, S., Yu, C., Chou, X., Yuan, W., Wang, Y., Bu, L., Fu, G. et al. Dentinogenesis imperfecta 1 with or without progressive hearing loss is associated with distinct mutations in DSPP. Nat. Genet. 27, 201–204 (2001).
Jacobs, H. T. Mitochondrial deafness. Ann. Med. 9, 483–491 (1997).
Fischel-Ghodsian, N., Prezant, T. R. & Fournier, P. Mitochondrial mutation associated with non-syndromic deafness. Am. J. Otolaryngol. 16, 403–408 (1995).
Fischel-Ghodsian, N. Mitochondrial deafness mutations reviewed. Hum. Mut. 13, 261–270 (1999).
Van Camp, G. & Smith, R. J. Maternally inherited hearing impairment. Clin. Genet. 57, 409–414 (2000).
Li, R., Xing, G., Yan, M., Cao, X., Liu, X. Z., Bu, X. et al. Cosegregation of C-insertion at position 961 with A1555G mutation of mitochondrial 12S rRNA gene in a large Chinese family with maternally inherited hearing loss. Am. J. Med. Genet. 124A, 113–117 (2004).
Li, Z., Li, R., Chen, J., Liao, Z., Zhu, Y., Qian, Y. et al. Mutational analysis of the mitochondrial 12S rRNA gene in Chinese pediatric subjects with aminoglycoside-induced and non-syndromic hearing loss. Hum. Genet. 117, 9–15 (2005).
Liu, X. Z., Angel, iS., Ouyang, X. M., Liu, W., Ke, X. M., Liu, Y. H. et al. Audiological and genetic features of the mtDNA mutations. Acta. Otolaryngol. 128, 732–738 (2008b).
Ballana, E., Morales, E., Rabionet, R., Montserrat, B., Ventayol, M., Bravo, O. et al. Mitochondrial 12S rRNA gene mutations affect RNA secondary structure and lead to variable penetrance in hearing impairment. Biochen. Biophys. Res. Commun. 341, 950–957 (2006).
Prezant, T. R., Agapian, J. V., Bohlman, M. C., Bu, X., Oztas, S., Qiu, W. Q. et al. Mitochondrial ribosomal RNA mutation associated with both antibiotic-induced and non-syndromic deafness. Nat. Genet. 4, 289–294 (1993).
Matthijs, G. S., Claes, S., Longo-Bbenza, B. & Cassiman, J. J. Non-syndromic deafness associated with a mutation and a polymorphism in the mitochondrial 12S ribosomal RNA gene in a large Zairean pedigree. Eur. J. Hum. Genet. 4, 46–51 (1996).
Pandya, A., Xia, X., Radnaabazar, J., Batsuuri, J., Dangaansuren, B., Fischel-Ghodsian, N. et al. Mutation in the mitochondrial 12S ribosomal-RNA gene in 2 families from Mongolia with matrilineal aminoglycoside ototoxicity. J. Med. Genet. 34, 169–172 (1987).
Estivill, X., Govea, N., Barceló, E., Badenas, C., Romero, E., Moral, L. et al. Familial progressive sensorineural deafness is mainly due to the mtDNA A1555G mutation and is enhanced by treatment of aminoglycosides. Am. J. Hum. Genet. Jan. 62, 27–35 (1998).
Young, W. Y., Zhao, L., Qian, Y., Wang, Q., Li, N. & Greinwald, J. H. Jr. Extremely low penetrance of hearing loss in four Chinese families with the mitochondrial 12S rRNA A1555G mutation. Biochem. Biophys. Res. Commun. 328, 1244–1251 (2005).
Bacino, C., Prezant, T. R., Bu, X., Fournier, P. & Fischel-Ghodsian, N. Susceptibility mutations in the mitochondrial small ribosomal RNA gene in aminoglycoside induced deafness. Pharmacogenetics. 5, 165–172 (1995).
Tang, H. Y., Hutcheson, E., Neill, S., Drummond-Borg, M., Speer, M. & Alford, R. L. Genetic susceptibility to aminoglycoside ototoxicity: how many are at risk? Genet. Med. 4, 336–345 (2002).
Zhao, L., Young, W. Y., Li, R., Wang, Q., Qian, Y. & Guan, M. X. Clinical evaluation and sequence analysis of the complete mitochondrial genome of three Chinese patients with hearing impairment associated with the 12S rRNA T1095C mutation. Biochem. Biophys. Res. Commun. 325, 1503–1508 (2004).
Tessa, A., Giannotti, A., Tieri, L., Vilarinho, L., Marotta, G. & Santorelli, F. M. Maternally inherited deafness associated with a T1095C mutation in the mDNA. Eur. J. Hum. Genet. 9, 147–149 (2001).
Thyagarajan, D., Bressman, S., Bruno, C., Przedborski, S., Shanske, S., Lynch, T. et al. A novel mitochondrial 12SrRNA point mutation in parkinsonism, deafness, and neuropathy. Ann. Neurol. 48, 730–736 (2000).
Usami, S., Abe, S., Akita, J., Namba, A., Shinkawa, H., Ishii, M. et al. Prevalence of mitochondrial gene mutations among hearing impaired patients. J. Med. Genet. 37, 38–40 (2000).
Malik, S. G., Pieter, N., Sudoyo, H., Kadir, A. & Marzuki, S. Prevalence of the mitochondrial DNA A1555G mutation in sensorineural deafness patients in island Southeast Asia. J. Hum. Genet. 48, 480–483 (2003).
Kupka, S., Tóth, T., Wróbel, M., Zeißler, U., Szyfter, W., Szyfter, K. et al. Mutation A1555G in the 12S rRNA gene and its epidemiological importance in German, Hungarian, and Polish patients. Hum. Mutat. 19, 308–309 (2002).
Tekin, M., Duman, T., Bo∂´oçlu, G., Incesulu, A., Comak, E., Fitoz, S. et al. Frequency of mtDNA A1555G and A7445G mutations among children with prelingual deafness in Turkey. Eur. J. Pediatr. 162, 154–158 (2003b).
ØStergaard, E., Montserrat-Sentis, B., Grønskov, K. & Brøndum-Nielsen, K. The A1555G mtDNA mutation in Danish hearing-impaired patients: frequency and clinical signs. Clin. Genet. 62, 303–305 (2002).
Jacobs, H. T., Hutchin, T. P., Käppi, T., Gillies, G., Minkkinen, K., Walker, J. et al. Mitochondrial DNA mutations in patients with postlingual, nonsyndromic hearing impairment. Eur. J. Hum. Genet. 13, 26–33 (2005).
Guan, M. X., Fischel-Ghodsian, N. & Attardi, G. A biochemical basis for the inherited susceptibility to aminoglycoside ototoxicity. Hum. Mol. Genet. 9, 1787–1793 (2000).
Hu, D. N., Qu, W. Q., Wu, B. T., Fang, L. Z., Zhou, F., Gu, Y. P. et al. Genetic aspects of antibiotic induced deafness: mitochondrial inheritance. J. Med. Genet. 28, 79–83 (1991).
Guan, M. X., Fischel-Ghodsian, N. & Attardi, G. Nuclear background determines biochemical phenotype in the deafness-associated mitochondrial 12S rRNA mutation. Hum. Mol. Genet. 10, 573–580 (2001).
Bykhovskaya, Y., Shohat, M., Ehrenman, K., Johnson, D., Hamon, M., Cantor, R. M et al. Evidence for complex nuclear inheritance in a pedigree with nonsyndromic deafness due to a homoplasmic mitochondrial mutation. Am. J. Med. Genet. 77, 421–426 (1998).
Bykhovskaya, Y., Estivill, X., Taylor, K., Hang, T., Hamon, M., Casano, R. A. et al. Candidate locus for a nuclear modifier gene for maternally inherited deafness. Am. J. Hum. Genet. 66, 1905–1910 (2000).
Bykhovskaya, Y., Yang, H., Taylor, K., Hang, T., Tun, R. Y., Estivill, X. et al. Modifier locus for mitochondrial DNA disease: linkage and linkage disequilibrium mapping of a nuclear modifier gene for maternally inherited deafness. Genet. Med. 3, 177–180 (2001).
Young, W. Y., Zhao, L., Qian, Y., Li, R., Chen, J., Yuan, H. et al. Variants in mitochondrial tRNAGlu tRNAArg and tRNAThr may influence the phenotypic manifestation of deafness-associated 12S rRNA A1555G mutation in three Chinese families with hearing loss. Am. J. Med. Genet. Part. 140A, 2188–2197 (2006).
Zhao, L., Wang, Q., Qian, Y., Li, R., Cao, J., Hart, L. C. et al. Clinical evaluation and mitochondrial genome sequence analysis of two Chinese families with aminoglycoside-induced and nonsyndromic hearing loss. Biochem. Biophys. Res. Commun. 336, 967–973 (2005).
Chen, B., Sun, D., Yang, L., Zhang, C., Yang, A., Zhu, Y. et al. Mitochondrial ND5 T12338C, tRNA(Cys) T5802C, and tRNA(Thr) G15927A variants may have a modifying role in the phenotypic manifestation of deafness-associated 12S rRNA A1555G mutation in three Han Chinese pedigrees. Am. J. Med. Genet. 146, 1248–1258 (2008).
Liao, Z., Zhao, J., Zhu, Y., Yang, L., Yang, A., Sun, D. et al. The ND4 G11696A mutation may influence the phenotypic manifestation of the deafness-associated 12S rRNA A1555G mutation in a four-generation Chinese family. Biochem. Biophys. Res. Commun. 362, 670–676 (2007).
Reid, F. M., Vernham, G. A. & Jacobs, H. T. A novel mitochondrial point mutation in a maternal pedigree with sensorineural deafness. Hum. Mutat. 3, 243–247 (1994).
Sevior, K. B., Hatamochi, A., Stewart, I. A., Bykhovskaya, Y., Allen-Powell, D. R. & Fischel-Ghodsian, N. Mitochondrial A7445G mutation in two pedigrees with palmoplantar keratoderma and deafness. Am. J. Med. Genet. 75, 179–185 (1998).
Pandya, A., Xia, X. J., Erdenetungalag, R., Amendola, M., Landa, B., Radnaabazar, J. et al. Heterogenous point mutations in the mitochondrial tRNASer(UCN) precursor coexisting with the A1555G mutation in deaf students from Mongolia. Am. J. Hum. Genet. 65, 1803–1806 (1999).
Jaksch, M., Klopstock, T., Kurlemann, G., Dörner, M., Hofmann, S., Kleinle, S. et al. Progressive myoclonus epilepsy and mitochondrial myopathy associated with mutations in the tRNASer(UCN) gene. Ann. Neurol. 44, 635–640 (1998).
Verhoeven, K., Ensink, R. J., Tiranti, V., Huygen, P. L., Johnson, D. F., Schatteman, I. et al. Hearing impairment and neurological dysfunction associated with a mutation in the mitochondrial tRNASer(UCN) gene. Eur. J. Hum. Genet. 7, 45–51 (1999).
Hutchin, T. P. & Cortopassi, G. A. Mitochondrial defects and hearing loss. Cell Mol. Life Sci. 57, 1927–1937 (2000).
Ishikawa, K., Tamagawa, Y., Takahashi, K., Kimura, H., Kusakari, J., Hara, A. et al. Nonsyndromic hearing loss caused by mitochondrial T7511C mutation. Laryngoscope. 112, 1494–1499 (2002).
Chapiro, E., Feldmann, D., Denoyelle, F., Sternberg, D., Jardel, C., Eliot, M. M. et al. Two large French pedigrees with non syndromic sensorineural deafness and the mitochondrial DNA T7511C mutation: evidenc for a modulatory factor. Eur. J. Hum. Genet. 10, 851–856 (2002).
Blanton, S. H., Liang, C. Y., Cai, M. W., Pandya, A., Du, L. L., Landa, B. et al. A novel locus for autosomal dominant non-syndromic deafness (DFNA41) maps to chromosome 12q24-qter. J. Med. Genet. 39, 567–570 (2002).
Yan, D., Ouyang, X. M., Zhu, X., Du, L. L., Chen, Z. Y. & Liu, X. Z. Refinement of the DFNA41 locus and candidate genes analysis. J. Hum. Genet. 50, 516–522 (2005).
Yan, D., Ke, X., Blanton, S. H., Ouyang, X. M., Pandya, A., Du, L. L. et al. A novel locus for autosomal dominant non-syndromic deafness, DFNA53, maps to chromosome 14q11.2-q12. J. Med. Genet. 43, 170–174 (2006).
Xia, J., Deng, H., Feng, Y., Zhang, H., Pan, Q., Dai, H. et al. A novel locus for autosomal dominant nonsyndromic hearing loss identified at 5q31.1-32 in a Chinese pedigree. J. Hum. Genet. 47, 635–640 (2002).
Wang, Q. J., Yang, W. Y., Wu, Z. M., Li, Q. Z., Guo, W. W. & Qiu, C. Y. Genetic analysis in a Chinese deaf-mute family with X linked recessive inheritance. Yi. Chuan. 26, 579–583 (2004a) (in Chinese).
Wang, Q. J., Lu, C. Y., Li, N., Rao, S. Q., Shi, Y. B., Han, D. Y. et al. Y-linked inheritance of non-syndromic hearing impairment in a large Chinese family. J. Med. Genet. 41, e80–e80 (2004b).
Acknowledgements
This study is supported by a grant from NIH DC R01 05575 (USA) and NSFC 30528025 (China) to XZL. Accession numbers of the genes and the url presented in the review are as follows: National Center for Biotechnology Information (NCBI) Entrez Database, http://www.ncbi.nlm.nih.gov/Entrez/ (for GJB2 cDNA [NM_004004]; GJB3 cDNA [BC012918]; DFNA5 cDNA [AF073308]; COCH cDNA [NM_004086]; SLC26A4 cDNA [NM_000441]; MYO7A cDNA [NM_000260]; Homo sapiens mitochondrion [NC_001807]).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Ouyang, X., Yan, D., Yuan, H. et al. The genetic bases for non-syndromic hearing loss among Chinese. J Hum Genet 54, 131–140 (2009). https://doi.org/10.1038/jhg.2009.4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2009.4
Keywords
This article is cited by
-
Analysis of deafness susceptibility gene of neonates in northern Guangdong, China
Scientific Reports (2024)
-
A novel missense variant in ESRRB gene causing autosomal recessive non-syndromic hearing loss: in silico analysis of a case
BMC Medical Genomics (2022)
-
Preventing autosomal-dominant hearing loss in Bth mice with CRISPR/CasRx-based RNA editing
Signal Transduction and Targeted Therapy (2022)
-
A multiplex PCR amplicon sequencing assay to screen genetic hearing loss variants in newborns
BMC Medical Genomics (2021)
-
Genetic Susceptibility Study of Chinese Sudden Sensorineural Hearing Loss Patients with Vertigo
Current Medical Science (2021)