Abstract
Methanotrophs in the rhizosphere play an important role in global climate change since they attenuate methane emission from rice field ecosystems into the atmosphere. Most of the CH4 is emitted via transport through the plant gas vascular system. We used this transport for stable isotope probing (SIP) of the methanotrophs in the rhizosphere under field conditions and pulse-labelled rice plants in a Chinese rice field with CH4 (99% 13C) for 7 days. The rate of 13CH4 loss rate during 13C application was comparable to the CH4 oxidation rate measured by the difluoromethane inhibition technique. The methanotrophic communities on the roots and in the rhizospheric soil were analyzed by terminal-restriction fragment length polymorphism (T-RFLP), cloning and sequencing of the particulate methane monooxygenase (pmoA) gene. Populations of type I methanotrophs were larger than those of type II. Both methane oxidation rates and composition of methanotrophic communities suggested that there was little difference between urea-fertilized and unfertilized fields. SIP of phospholipid fatty acids (PLFA-SIP) and rRNA (RNA-SIP) were used to analyze the metabolically active methanotrophic community in rhizospheric soil. PLFA of type I compared with type II methanotrophs was labelled more strongly with 13C, reaching a maximum of 6.8 atom-%. T-RFLP analysis and cloning/sequencing of 16S rRNA genes showed that methanotrophs, especially of type I, were slightly enriched in the ‘heavy’ fractions. Our results indicate that CH4 oxidation in the rice rhizosphere under in situ conditions is mainly due to type I methanotrophs.
Similar content being viewed by others
Introduction
Flooded rice fields, an important source of atmospheric methane, are widely distributed in China (Cai, 1997; Li et al., 2002; Cai et al., 2003). The total emission of CH4 from Chinese rice fields was estimated to be in the range of 8.05±3.69 Tg CH4 per year, depending on the type of rice paddy field (Cai, 1997). In these ecosystems, the main CH4 emission to the atmosphere occurs through the aerenchyma system of the rice plants (Nouchi et al., 1990; Schütz et al., 1991). In turn, oxygen from the atmosphere is transported to the rice roots enabling aerobic and microaerophilic conditions in the rhizosphere (Liesack et al., 2000). Aerobic methanotrophs in such rice rhizosphere environments play an important role in CH4 emission, since they attenuate the CH4 flux from the soil into the atmosphere covering a large range between 0 and 94% attenuation (Schütz et al., 1989; Groot et al., 2003). The activity of aerobic methanotrophs strongly depends on the environmental conditions, for example the in situ concentrations of not only CH4 and O2, but also of other compounds, such as ammonium, sulfide or ferrous iron. The latter compounds can also be microbially oxidized in the rhizosphere resulting in competition with methanotrophs for oxygen. Ammonia oxidizers may affect CH4 oxidation due to the similarity between the key enzymes catalyzing the first step of CH4 oxidation and ammonium oxidation (Mancinelli, 1995). In previous studies, inhibition (Bosse et al., 1993; Nold et al., 1999), stimulation (Bodelier et al., 2000; Eller and Frenzel, 2001) or no effect (Lilkanen and Martikainen, 2003) on aerobic methanotrophs were observed in anoxic soils and sediments when fertilized with ammonium or urea (see review by Bodelier and Laanbroek, 2004). Another important factor for CH4 emission is the composition of the methanotrophic communities, which seem to be different in bulk soil and rhizosphere and change dynamically over the season and upon fertilization with ammonium or urea (Bodelier et al., 2000; Eller and Frenzel, 2001; Macalady et al., 2002; Eller et al., 2005; Shrestha et al., 2008).
A further complication is the fact that not all members of the total methanotrophic community are equally active in attenuation of the CH4 flux. Pulse labelling of rice plants with 13C followed by stable isotope probing (SIP) of nucleic acid and membrane lipids has been demonstrated as a powerful approach to identify the metabolically active microbial communities in the rhizosphere of rice (Lu and Conrad, 2005; Lu et al., 2006, 2007). SIP allows direct observations of substrate assimilation in minimally disturbed communities, and thus represents an exciting tool for linking microbial identity and function. As methanotrophs use CH4 as carbon source, SIP of phospholipid fatty acids (PLFAs) (Boschker et al., 1998; Boschker and Middelburg, 2002) or DNA/RNA (McDonald et al., 2005) using 13C-labelled CH4 is a promising approach. Recently, rice microcosms were used to identify the methanotrophs actively assimilating CH4-carbon that was introduced into the rice rhizosphere by means of permeable tubings (Shrestha et al., 2008). However, a labelling approach under field conditions has not been conducted, and therefore the ability to compare laboratory-based results is still lacking.
Therefore, we applied SIP in a Chinese rice field by pulse labelling the rice plants with 13C-labelled CH4 through the aerenchyma systems of the rice plants into the rhizosphere. This enables an in situ labelling of methanotrophs located directly at the rice rhizosphere without disturbing the soil ecosystem. Furthermore, we tested if typical Chinese farm fertilization procedures had an effect on methanotrophic activity and on the global methane emission budget from this environment.
Materials and methods
Study site and treatments
The experimental field was located at the China National Rice Research Institute (CNRRI; Hangzhou, Zhejiang province, China; 30°25′N, 120°18′E). The fields cover an area of 494 ha, representing a typical rice growing area with annual single- and double-rice cropping systems. The climate is warm/cool humid subtropical with an annual rainfall of 1400 mm and a mean solar radiation of 12.8 MJ m−2 day−1.
The field had been treated according to the common agricultural practice of the region. Fertilizers were applied directly to the rhizosphere of rice plants at the rates of 0.09 g P and 0.41 g K per plant and additional 0.6 g urea at 48 days after transplantation. For our study, urea treatment (PKN) was done in the same field, but separated with four rows of rice plants from those plots that were only fertilized with PK. The water level was kept constant at 5–10 cm depth. The field plots were shaded with transparent black fiber tissue to avoid excessively high solar radiation during our experiment to reduce the evaporation inside the gas chambers. The soil is a silty clay soil with a pH of 6.7, organic C of 24.2 g per kg soil, total N of 2.3 g per kg soil and cation exchange capacity of 144 mmol per kg soil.
Methane flux measurement
Methane flux was measured on 4 consecutive days starting 3 days after fertilization. The measurements were always done between 12:00 and 16:00 hours local time. The cylindrical flux chambers (diameter=5 cm and height=40 cm) were made of plexiglas. For flux measurements, rice plants were covered for a period of 4 h and gas samples were withdrawn every 30 min. Methane emission rates were determined from the slope of the linearly increasing CH4 mixing ratio using a gas chromatograph with flame ionization detector.
Loss rates of 13CH4 were determined after injection of 1 ml of 13C-labelled CH4 (99 atom-% 13CH4; Sigma, Taufkirchen, Germany) into the flux chamber. The amount of 13CH4 at time t (nt) was calculated by:

mt=total amount of CH4 at time t; at=13C atom-% of CH4 at time t; ap=natural 13C atom-% of CH4. The 13C atom-% was analyzed using a gas chromatography–combustion–isotope ratio mass spectrometry as described previously (Conrad et al., 2002).
The rate of CH4 oxidation was determined by measuring the CH4 flux in the presence and absence of difluoromethane (20 ml CH2F2), an inhibitor of CH4 oxidation (Miller et al., 1998), which was injected into the flux chamber. The measurements were done as described by Krüger et al. (2001). It was made sure that the concentration of CH2F2 did not decrease over the 4-h incubation time. Methane oxidation rates were calculated by

The rates of the flux measurements are expressed in mmol CH4 per m2 per h giving the mean±s.e. of n=4 data obtained on 4 consecutive days.
13C-labelling and sampling procedures
Labelling with 13CH4 was started 36 h after fertilization. Plots were covered with closed chambers as described above. For both the N-fertilized (PKN) and the unfertilized (PK) plots the following experiments were performed: (1) rice plants were cut so that stubbles were 15 cm above the water surface and 15 ml of 13CH4 (99 atom-% 13CH4; Sigma) was added to the chamber twice a day (13CH4 cut). (2) Rice plants were left intact and treated as above (13CH4). (3) Rice plants were left intact and treated with 15 ml of unlabelled CH4 (12CH4). (4) Rice plants were left intact and no CH4 was injected into the chamber (air). At the end of 7-day labelling, the chambers were removed and the rice plants were pulled out. Four replicates were taken in each treatment. The bulk soil fell off the roots during plant tiller separation and with gentle shaking. The soil still adhering to the roots was defined as rhizospheric soil. After cutting off the shoots, the roots with the adhering soil were separated into three batches and immediately frozen in liquid nitrogen and stored at −80 °C. Transport to laboratories in Beijing (PLFA extraction) and Marburg (DNA/RNA extraction) was done within 20 h keeping the samples in a frozen state.
Phospholipid fatty acid extraction and 13C profiling
After thawing one batch of the soil–roots samples for each replicate treatment, the soil was carefully washed from roots with deionized sterile water. The soil slurries were collected separately and centrifuged at 950 g. Total PLFA was extracted from 4 g fresh soil samples using the Bligh–Dyer method (Bligh and Dyer, 1969). The total amount of PLFA was determined by gas chromatography–mass spectrometry and the stable isotope ratio of individual fatty acid methyl esters by gas chromatography–combustion–isotope ratio mass spectrometry as described by Lu et al. (2007).
The relative amounts of individual fatty acid methyl esters were calculated on the basis of the amount of PLFA 16:0. This PLFA was chosen as it is ubiquitous in prokaryotes. The fractional abundance (F) of 13C in the unlabelled (Fu) and labelled (Fl) PLFAs was used to calculate the concentration of 13C incorporated into PLFAs ([PLFA-13C]) from the total PLFA concentration ([PLFA]):

The fractional abundance expresses the amount of 13C as a proportion of the total amount of carbon in the PLFA:

DNA extraction and pmoA gene T-RFLP analysis
After thawing another batch of the soil-roots samples for each replicate treatment, the soil was carefully washed from roots with deionized sterile water. The soil slurries were collected separately and centrifuged at 950 g and used for nucleic acid extraction. DNA was extracted from rhizospheric soil according to Lueders et al. (2004) with minor modifications during the precipitation step. DNA was precipitated from the aqueous phase with 1 volume isopropanol and centrifugation at maximum speed (21 000 g) at 4 °C for 1 h. The precipitate was washed once with 500 μl of pre-cooled 70% ethanol, dried and re-suspended in 50 μl of PCR water (Sigma-Aldrich). Additional purification was done using a DNA clean-up system (Promega, Mannheim, Germany). Amplification of the particulate methane monooxygenase (pmoA) gene was performed using the primer sets 189f/682r and 189f-FAM/mb661r using nested PCR (Horz et al., 2005). Purified amplicons were digested by MspI (HpaII) (Fermentas, St Leon-Rot, Germany) at 37 °C for 3 h and inactivated at 65 °C. Aliquots of desalted digest (1–2 μl) were mixed with 12 μl of Hi-Di formamide (Applied Biosystems, Weiterstadt, Germany) and 0.25 μl of ROX-labelled MapMarker 1000 ladder (BioVentures, Murfreesboro, TN, USA), denaturated for 3 min at 95 °C, cooled on ice immediately and size separated on an ABI 310 genetic analyzer (Applied Biosystems). Electrophoresis was performed with POP-4 polymer in 47 cm by 50 μm capillaries (Applied Biosystems) with 15 kV injection voltage, 15 kV run voltage, 60 °C run temperature and 45 min analysis time. The peak heights were automatically quantified by the GeneScan software. The relative abundance of individual terminal-restriction fragments (T-RFs) were calculated based on the peak height of the individual T-RFs in relation to the total peak height of all T-RFs detected in the respective terminal-restriction fragment length polymorphism (T-RFLP) community pattern.
RNA extraction and gradient centrifugation
Total RNA was extracted from the rhizospheric soil samples 13CH4 cut+PKN and 12CH4 cut+PKN using a previously described protocol (Noll et al., 2005). RNA for reverse transcription (RT)-PCR was obtained by removal of co-extracted DNA using RNase-Free DNase (Promega) following the manufacturer's protocol. Additional RNasin (Promega) was added to inhibit RNases. The RNA pellets were re-suspended in 50 μl nuclease-free water (Promega). The absence of DNA in the resulting small-subunit ribosomal RNA (16S rRNA) preparation was checked by PCR of bacterial 16S rRNA genes and standard agarose gel electrophoresis on a 1% agarose followed by ethidium bromide staining. RiboGreen (500 ng; Invitrogen, Karlsruhe, Germany) was used to quantify RNA content in the extracts. The RNA was density resolved by equilibrium density gradient centrifugation in CsTFA (cesium trifluoroacetate) (Amersham Biosciences, Freiburg, Germany) under the conditions reported previously (Lueders et al., 2004). After centrifugation, the density of the collected gradient fractions was determined refractometrically, and RNA was precipitated and re-suspended in 30 μl nuclease-free water for subsequent community analyses. Control gradients were conducted with RNA from unlabelled soil (12CH4 cut−PKN).
RT-PCR of bacterial 16S rRNA gene and T-RFLP analysis
Terminal-restriction fragment length polymorphism analysis of density-resolved bacterial communities from gradient fractions was performed with primers Ba27f-FAM/Ba907r by RT-PCR using a one-step RT-PCR system (Access Quick1 RT-PCR-System; Promega). The reaction mixture (50 ml) contained 25 ml of 2 × reaction buffer, 0.5 mM of each primer (MWG Biotech, Ebersberg, Germany), 1.0 μl of template RNA and 1 μl (5 U) of AMV reverse transcriptase supplied by the manufacturer. The 2 × reaction buffer included 3 mM MgSO4 (final concentration 1.5 mM), Tfl DNA polymerase (unknown amount) and the deoxynucleoside triphosphates (concentration was not provided by the manufacturer). The RT reaction was carried out at 48 °C for 45 min and followed by a denaturing step at 94 °C for 2 min to inactivate the AMV reverse transcriptase and to activate the Tfl DNA polymerase. The thermal profile of the PCR included 25–30 cycles of primer annealing at 52 °C for 45 s, primer extension at 68 °C for 1 min and denaturing at 94 °C for 45 s. The final elongation step was 5 min. Amplicons were digested by MspI and separated as described above for the pmoA gene. To verify the assignments of T-RFs to our detected methanotrophs, we also tested individual methanotrophic clones indicated in sequence data by T-RFLP analysis.
Cloning, sequencing and phylogenetic analysis
Four clone libraries were created, one of pmoA genes from sample 13CH4 cut+PKN (54 clones) and three of bacterial 16S rRNA genes from density-resolved ‘heavy’ (HM) (101 clones) and ‘light’ (LM) RNA fractions (74 clones) and control (CM) (78 clones) as indicated in Figure 5. Amplicons for sequence analyses were generated with the same primers and PCR conditions as described above and cloned into Escherichia coli using the pGEM-T Vector System (Promega). The randomly selected clones were sequenced by the ADIS DNA core facility (Max Planck Institute for Plant Breeding Research, Cologne, Germany) on an ABI Prism 3700 sequencer (Applied Biosystems) using BigDye terminator cycle sequencing chemistry. Raw sequence data were assembled and checked with DNAStar (Madison, WI, USA). All sequences were compared with those in the public databases by using NCBI megaBlast and the highest matched sequences were obtained from GenBank database (http://www.ncbi.nih.gov). Chimeras were checked for with the Bellerophon server (http://foo.maths.uq.edu.au/~huber/doc/doc.html) (Huber et al., 2004) for all sequences and chimera sequences were excluded for further analysis. Phylogenetic analysis were conducted using the ARB software package (Ludwig et al., 2004). A neighbor-joining tree was calculated for bacterial 16S rRNA gene sequences including a 50% base frequency filter. Neighbour-joining, maximum likelihood and tree-puzzle programs were used for calculating pmoA-based phylogenetic trees in which all showed the same topology.
Sequence data have been submitted to EMBL under accession numbers AM909841 to AM910093 for 16S rRNA gene and AM910094 to AM910147 for pmoA gene.
Results
Methane oxidation activity
We fertilized two plots of the same rice field with either phosphorous and potassium (PK treatment) or with phosphorous, potassium and nitrogen (PKN treatment) using the fertilization procedure typical for Chinese agriculture. Nine days after fertilization, ammonia concentrations in bulk soil were two times higher in the PKN treatment (27.6±0.7 mg N per kg dry soil) compared with the PK treatment (11.5±0.4 mg N per kg dry soil), while nitrate concentrations were low in both treatments (0.6±0.1 and 1.1±0.1 mg N per kg dry soil, respectively). Three to seven days after fertilization, CH4 emission flux from the rice field plots was measured using the closed chamber technique. The flux was measured in the presence and absence of difluoromethane (CH2F2) as specific inhibitor of methanotrophs (Miller et al., 1998), the difference being the rate of CH4 oxidation (Table 1). Both CH4 oxidation activity and the loss of 13CH4 in the chambers to which 13C-labelled CH4 was added indicate that about 2–18% of the produced CH4 was oxidized (Table 1). The rates of CH4 oxidation or 13CH4 loss were on the same order of magnitude irrespectively of whether the fields were fertilized with urea or not.
Isotopic enrichment of PLFAs
Rhizospheric soil samples were retrieved from rice field plots after different treatments, that is, fertilization with (PKN) and without (PK) urea, cutting plants above the water surface (cut), and addition of 13CH4, 12CH4 or no CH4 (air) to the headspace of the flux chambers. Cutting plants above the water surface does not impede the CH4 flux into the atmosphere (Seiler et al., 1984; Conrad and Schütz, 1988). The relative abundance of total PLFA was very similar in all the soil samples irrespectively of the treatment. The PLFA included Gram-positive bacterial biomarkers (15:0i, 15:0a, 17:0i, cy17:0d7,8 and 10Me-16:0) and Gram-negative biomarkers (16:1ω7, 18:1ω7 and cy19:0d8,9) (Figure 1).
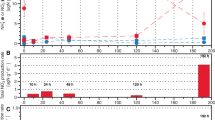
(a) Concentrations of individual phospholipid fatty acids (PLFAs) at the end of the field experiment using samples from all the different treatments including N fertilization and cutting of plants; mean±s.e., n=32. The small error bars show that concentrations of PLFAs were similar in the different treatments. (b and c) Concentrations of 13C-labelled PLFA in the soil at the end of the field experiment with (PKN) and without (PK) addition of N fertilizer on plots with (b) intact plants and (c) plants cut above the water surface, both labelled with 13CH4; mean±s.e.; n=4.
Pulsed addition of 13CH4 into the gas headspace led to strong labelling of methanotrophic PLFAs, reaching up to 6.8 13C atom-% in 16:1ω7 PLFA of the 13CH4 cut+PKN treatment. Mainly, the saturated and monounsaturated PLFAs with 16 and 18-carbon atoms exhibited a significant isotopic enrichment (Figure 1). Notably, the monounsaturated 16:1ω7 and 16:1ω6 were highly 13C-labelled in the PLFA profiles, which have been often affiliated to type I methanotrophs (Nold et al., 1999; Mohanty et al., 2007). Saturated C14 fatty acid, which occurs in type I Methylomonas species (Hanson and Hanson, 1996; Evershed et al., 2006), was also labelled to a less extent. The PLFAs 18:1ω7 and 18:1ω9, occurring in type II methanotrophs (Nold et al., 1999; Mohanty et al., 2007), were relatively strongly 13C-labelled, but with less intensity than the PLFAs with 16-carbon fatty acids. There was no significant difference between the plots with and without ammonium fertilization (analysis of variance, P>0.05) (Figure 1). Likewise, the treatments with and without cutting of the plants had similar isotope incorporation PLFA profiles (Figure 1). In PLFAs extracted from the unlabelled methane control soil, there was no enrichment of 13C in the PLFAs.
Characterization of methanotroph communities
To characterize the functional gene diversity of the methanotrophic community, we applied pmoA gene-based T-RFLP fingerprint analysis of the different treatments after 7 days of incubation (Figure 2). Neither urea fertilization, nor addition of unlabelled or labelled CH4, nor cutting of the plants had a significant (analysis of variance, P>0.05) effect on the community structure of the T-RFLP patterns.
Relative abundance of terminal-restriction fragments (T-RFs) of particulate methane monooxygenase (pmoA) amplicons obtained from the various treatments at the end of the field experiment. 244-bp T-RF is representative of type II methanotrophs; the others (79, 349, 437 and 505 bp) are representative of type I methanotrophs. PK and PKN indicates the fertilizers applied; air, 12CH4 and 13CH4, the gas treatment; cut indicates that plants were cut above the water surface; mean±s.e., n=2.
To assess the pmoA gene-based diversity, we cloned and sequenced in total 54 clones from pooled PCR products from all replicates of 13CH4 cut+PKN. This treatment was chosen for cloning, as it was also used for SIP of rRNA (RNA-SIP) (see below). Out of 54 clones, 17 represented type II methanotrophs, which clustered within the Methylocystis group (Figure 3). These type II methanotrophs represented the 244-bp T-RF. All the other T-RFs represented type I methanotrophs. Type I pmoA made up approximately 80% of the total methanotrophic community. Some (15 clones) of the pmoA clones with a T-RF=79 bp, represented pmoA from a group of type I methanotrophs that was distinct from pmoA from extant methanotrophs. Other clones clustered with pmoA from extant Methylococcus/Methylocaldum, Methylomonas and Methylomicrobium/Methylobacter species (Figure 3). The T-RF of 456 bp length occurred only with a low relative abundance (<1%) in the T-RFLP analyses and was affiliated to members of the genus Methylomicrobium (Figure 3).
Phylogenetic relationship (neighbor-joining) of amino-acid sequences derived from particulate methane monooxygenase (pmoA) sequences cloned from rice soil methanotrophs (RSMs) retrieved from soil samples of the cut+PKN treatment. The terminal-restriction fragment (T-RF) sizes of clone sequences digested in silico with MspI are shown in bold. Values in parentheses give the number of clones in the indicated group. The scale bar represents 10% sequence divergence; outgroup=amoA. GeneBank accession numbers of the reference sequences are indicated.
Structure of bacterial community in rhizospheric soil
Ribosomal RNA was extracted from the rhizospheric soil of samples 13CH4 cut−PKN and 12CH4 cut−PKN, respectively. The RNA extracts were centrifuged in a density gradient and density fractions were amplified via RT-PCR with bacterial primers for subsequent T-RFLP analyses (Figure 4). Three different fractions were used to construct clone libraries of bacterial 16S rRNA, one from the control incubated with 12CH4 (CM), one from the ‘heavy’ RNA (HM) and one from the ‘light’ RNA (LM) fraction as indicated in Figure 4. In total, 101 clones were sequenced from the HM library; 65% were affiliated to Proteobacteria and the remaining 35% belonged to diverse phylogenetic taxa including Actinobacteria, Firmicutes, Acidobacteria, Gemmatimonadales, Thermomicrobia, Cyanobacteria, Nitrospirae, Verrucomicrobia, Bacteroidetes and Planctomycetes (Figure 5, Table 2) (note the recent discovery of thermoacidophilic methanotrophs belonging to the Verrucomicrobia; Dunfield et al., 2007; Pol et al., 2007; Islam et al., 2008). Of the 74 clones sequenced from the LM library and the 78 clones sequenced from the CM library, 62 and 45%, respectively, were affiliated to Proteobacteria. Deltaproteobacteria-like clone sequences dominated the retrieved sequences affiliated to Proteobacteria (34, 28 and 33% in HM, LM and CM clone library, respectively). Methanotrophic bacteria within the Alphaproteobacteria and Gammaproteobacteria were relatively more frequent in the HM library than in the LM and CM clone libraries (Table 2). In silico prediction of T-RF sizes from sequence data (Table 2) allowed the assignment of the major T-RFs observed in the bacterial fingerprints (Figure 5).
Bacterial terminal-restriction fragment length polymorphism (T-RFLP) fingerprints of 16S rRNA retrieved from density-resolved gradient fractions of (a) 13C-labelled rhizospheric soil and (b) unlabelled control soil. CsTFA (cesium trifluoroacetate) buoyant densities (g ml−1) of fractions are given. The specific fragment lengths (bp) of methanotrophic T-RFs (as mentioned in the text) are indicated by arrows. HM, LM and CM indicate the fractions from which clone libraries were generated.
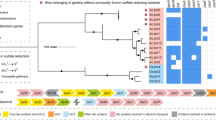
Phylogenetic relationship (neighbor-joining) of cloned bacterial 16S rRNA gene sequences retrieved from amplicons of the density-resolved ‘heavy’ (HM) and ‘light’ (LM) RNA fractions and from the control (CM) RNA. The new clone sequences and their terminal-restriction fragment (T-RF) sizes digested in silico with MspI are shown in bold. The scale bar represents 10% sequence divergence. GenBank accession numbers of the reference sequences are indicated.
Metabolically active methanotrophic community
The comparison between 16S rRNA T-RFLP fingerprinting and clone libraries indicated a relative increase of the 150-bp, 455-bp and 487-bp T-RFs in the ‘heavy’ compared with the other RNA fractions (Figure 4). The T-RFLP patterns are in agreement with the higher abundance of methanotrophs in the HM versus the LM and CM clone libraries (Table 2). Thus, it seems that methanotrophs were enriched in the ‘heavy’ fraction. The T-RFs of 455 and 487 bp represented type I methanotrophs. Only Methylosarcina and Methylomicrobium had a T-RF=487 bp when considering all the bacterial 16S rRNA sequences retrieved (Table 2). Some of type I methanotrophs had 444-bp and 479-bp T-RFs as indicted by the HM clone library (Table 2), but these T-RFs were not seen in the T-RFLP patterns (Figure 4). The T-RFs of 150 bp length were mainly found in clones belonging to Methylocystis species and exhibited a significant increase in the ‘heavy’ fractions (Figure 4). No other type II methanotrophs were found in the ‘heavy’ fractions. Hence, type I methanotrophs, including Methylobacteria, Methylosarcina, Methylomicrobium and Methylomonas, seemed to be more diverse than type II methanotrophs in the ‘heavy’ fraction (Figure 4).
Discussion
By applying 13CH4 to the phyllosphere of rice, we were able to identify for the first time active methanotrophs under field conditions. Both PLFA-SIP (SIP of phospholipid fatty acids) and RNA-SIP indicated that mainly type I methanotrophs and to a less extent type II methanotrophs assimilated CH4 in the rhizosphere.
In previous studies, CH4 was supplied to the soil–plant system by the water from the bottom of rice microcosms (Groot et al., 2003) or through permeable tubes buried into the soil (Shrestha et al., 2008). Such technical approach is suitable for microcosm experiments, but can hardly be used for field studies. Since rice plants allow the transport of O2 from the atmosphere into the soil and of CH4 from the soil into atmosphere via their gas vascular system and aerenchyma tissue in roots and shoots by a diffusional process (Nouchi et al., 1990; Schütz et al., 1991), we inferred that transport of 13C-labelled CH4 from the atmosphere into the rhizosphere is also possible and that rhizospheric methanotrophs can be labelled in this way. The transport of 13C-labelled CH4 from the atmosphere into the rhizosphere was indeed operating as shown by the incorporation of 13C into microbial PLFAs. Cutting plants above the water surface does not impede the CH4 flux into the atmosphere (Seiler et al., 1984; Conrad and Schütz, 1988). It apparently did also not impede the transport of 13CH4 into the rhizosphere as microbial PLFAs were comparably labelled as with intact plants.
After labelling for 7 days, the 13C content of some PLFAs (for example, 16:1ω7 and 16:1ω6) reached a value of almost 7 atom-%. Labelling experiments with 13CH4 diffusing from permeable tubes buried into the soil achieved only about 3 atom-% incorporation (Shrestha et al., 2008), indicating that the supply of CH4 through the plant vascular system allows a more efficient labelling of the rhizosphere microbiota. Manefield et al. (2002) determined in a pure culture that the labelling of RNA should be >10 atom-% to allow separation by density centrifugation. However, recent studies of plants pulse labelled with 13CO2 showed that root-inhabiting microbes incorporated within only 5 h sufficient amounts of 13C to allow separation of heavy RNA by density gradient centrifugation (Vandenkoornhuyse et al., 2007). This was possible although the δ13C content of the bulk root biomass was rather low (equivalent to 1.4 atom-%). Although the labelled PLFAs detected in our experiments are biomarkers for a limited group of bacteria, they nevertheless represent not only methanotrophs but also other microorganisms, so that the methanotrophs might be 13C-labelled by >7 atom-%. Therefore, we attempted RNA-SIP, which indeed was successful. The obtained results were consistent with the results of PLFA-SIP and indicated that type I methanotrophs were slightly more active in assimilation of 13CH4 than type II methanotrophs. Hence, the application of 13CH4 to the rice canopy seems to be a suitable technique for identification of rhizospheric methanotrophs active in CH4 oxidation.
The fatty acid 16:1ω7 received most of the 13C-label in our experiment (Figure 3). This fatty acid is common not only in type I methanotrophs but also in ammonia oxidizers, even in one type II methanotroph (Dedysh et al., 2007). Although ammonia oxidizers can oxidize CH4 and contain the PLFAs 16:0 and 16:1ω7 (Roslev and Iversen, 1999), they seem to play no role for CH4 oxidation in rice field soil (Bodelier and Frenzel, 1999). Furthermore, ammonia oxidizers may not be able to assimilate CH4 as a carbon source (Bedard and Knowles, 1989). Since the 13C-labelling of PLFA was similar in samples from unfertilized and urea-fertilized plots, we assume that ammonia oxidizers indeed were not involved in CH4 oxidation. A somewhat lower 13C-labelling was found for 18:1ω7 PLFA, which is present in all known type II methanotrophs (Bowmann et al., 1993). Labelling of 18:1ω9 PLFA, which also belongs to type II methanotroph, was still weaker. Hence, methanotophs characterized by C16-PLFAs incorporated more labelled CH4 than those characterized by C18-PLFAs, suggesting that type I methanotrophs were more active in the rhizosphere of rice than type II methanotrophs.
To assess methanotroph diversity within environmental samples, functional genes, such as pmoA, have also been used extensively as targets for PCR amplification (McDonald and Murrell, 1997) in rice field ecosystems (Henckel et al., 1999; Hoffmann et al., 2002; Shrestha et al., 2008). This approach has been successful with methanotrophs as their 16S rRNA phylogeny is reflected in the phylogenetic relationships of pmoA (Holmes et al., 1995), with the exception of Methylocella spp, which do not contain pmoA (Dedysh et al., 2000). Thus, the analysis of pmoA in our study suggested a large diversity, including Methylomonas, Methylococcus/Methylocaldum, Methylomicrobium, Methylobacter and Methylocystis species. An uncultured group of type I methanotrophs with the same T-RF size as Methylococcus/Methylocaldum was detected in the clone libraries of pmoA. This uncultured group has formerly been detected on rice roots from Italian rice fields (Horz et al., 2001) and littoral lake sediment (Bussmann et al., 2004; Pester et al., 2004). Collectively, the pmoA analyses confirmed that type I methanotrophs formed the dominant methanotrophic populations in the rice rhizosphere. However, little difference was observed between plots that were fertilized or not fertilized with urea. This was consistent with methane oxidation measurement and PLFA analysis. Possibly, urea fertilization affects on methanotrophic populations only for a short period after the treatment (Dan et al., 2001; Krüger and Frenzel, 2003). Urea is the most important nitrogen fertilizer in rice agriculture (Dedatta and Buresh, 1989).
Among the type II methanotrophs, only Methylocystis spp was detected by analysis of both pmoA and 16S rRNA genes. RNA-SIP indicated that the T-RF=150 bp increased in the ‘heavy’ fractions (Figure 4), which predominantly represented type II methanotrophs. Among the type I methanotrophs mainly the T-RFs of 455 and 487 bp increased with the density fraction and were only found in the clone library from the ‘heavy’ but not from the ‘light’ or ‘control’ rRNA clone libraries. The relative abundance of these different T-RFs in the different fractions indicate that both type I and type II methanotrophs incorporated 13C from the labelled CH4, but type I increased relatively more than type II methanotrophs (Table 3).
In conclusion, we found that SIP of methanotrophic bacteria is possible for rice fields under in situ conditions identifying individual groups of methanotrophs that are active in the rhizosphere and contribute to attenuate the emission of CH4 into the atmosphere. It seems that type I compared with type II methanotrophs have a larger population (T-RFLP of pmoA gene) in the rhizosphere and are also more active in assimilating 13CH4 into PLFA and RNA. A more precise determination, in particular by RNA-SIP, should be possible, if the production of unlabelled CH4 by the methanogenic community in the soil would be relatively low so that the pool of 13CH4 in the rhizosphere is not much diluted. This labelling approach may be adapted to other ecosystem studies to understand ecosystem functioning without laboratory-based limitations.
References
Bedard C, Knowles R . (1989). Physiology, biochemistry, and specific inhibitors of CH4, NH4+, and CO oxidation by methanotrophs and nitrifiers. Microbiol Rev 53: 68–84.
Bligh EG, Dyer WJ . (1969). A rapid method for total lipid extraction and purification. Can J Biochem Physiol 37: 911–917.
Bodelier PLE, Frenzel P . (1999). Contribution of methanotrophic and nitrifying bacteria to CH4 and NH4+ oxidation in the rhizosphere of rice plants as determined by new methods of discrimination. Appl Environ Microbiol 65: 1826–1833.
Bodelier PLE, Laanbroek HJ . (2004). Nitrogen as a regulatory factor of methane oxidation in soils and sediments (Review). FEMS Microbiol Ecol 47: 265–277.
Bodelier PLE, Roslev P, Henckel T, Frenzel P . (2000). Stimulation by ammonium-based fertilizers of methane oxidation in soil around rice roots. Nature 403: 421–424.
Boschker HTS, Middelburg JJ . (2002). Stable isotopes and biomarkers in microbial ecology (Review). FEMS Microbiol Ecol 40: 85–95.
Boschker HTS, Nold SC, Wellsbury P, Bos D, DeGraaf W, Pel R et al. (1998). Direct linking of microbial populations to specific biogeochemical processes by 13C-labelling of biomarkers. Nature 392: 801–805.
Bosse U, Frenzel P, Conrad R . (1993). Inhibition of methane oxidation by ammonium in the surface layer of a littoral sediment. FEMS Microbiol Ecol 13: 123–134.
Bowmann JP, Sly LI, Nichols PD, Hayward AC . (1993). Revised taxonomy of the methanotrophs: description of Methylobacter gen. nov., emendation of Methylococcus, validation of Methylosinus and Methylocystis species, and a proposal that the family Methylococcaceae includes only the group I methanotrophs. Int J Syst Bacteriol 43: 735–753.
Bussmann I, Pester M, Brune A, Schink B . (2004). Preferential cultivation of type II methanotrophic bacteria from littoral sediments (Lake Constance). FEMS Microbiol Ecol 47: 179–189.
Cai ZC . (1997). A category for estimate of CH4 emission from rice paddy fields in china. Nutrient Cycling in Agroecosystems 49: 171–179.
Cai ZC, Tsuruta H, Gao M, Xu H, Wei CF . (2003). Options for mitigating methane emission from a permanently flooded rice field. Glob Chang Biol 9: 37–45.
Conrad R, Klose M, Claus P . (2002). Pathway of CH4 formation in anoxic rice field soil and rice roots determined by 13C-stable isotope fractionation. Chemosphere 47: 797–806.
Conrad R, Schütz H . (1988). Methods of studying methanogenic bacteria and methanogenic activities in aquatic environments. In: Austin B (ed). Methods in Aquatic Bacteriology. Wiley: Chichester. pp 301–343.
Dan JG, Krüger M, Frenzel P, Conrad R . (2001). Effect of a late season urea fertilization on methane emission from a rice field in Italy. Agric Ecosyst Environ 83: 191–199.
Dedatta SK, Buresh RJ . (1989). Integrated nitrogen management in irrigated rice. Adv Soil Sci 10: 143–169.
Dedysh SN, Belova SE, Bodelier PLE, Smirnova KV, Khmelenina VN, Chidthaisong A et al. (2007). Methylocystis heyeri sp. nov., a novel type II methanotrophic bacterium possessing ‘signature’ fatty acids of type I methanotrophs. Int J Syst Evol Microbiol 57: 472–479.
Dedysh SN, Liesack W, Khmelenina VN, Suzina NE, Trotsenko YA, Semrau JD et al. (2000). Methylocella palustris gen. nov., sp nov., a new methane-oxidizing acidophilic bacterium from peat bags, representing a novel subtype of serine-pathway methanotrophs. Int J Syst Evol Microbiol 50: 955–969.
Dunfield PF, Yuryev A, Senin P, Smirnova AV, Stott MB, Hou SB et al. (2007). Methane oxidation by an extremely acidophilic bacterium of the phylum Verrucomicrobia. Nature 450: 879–882.
Eller G, Frenzel P . (2001). Changes in activity and community structure of methane-oxidizing bacteria over the growth period of rice. Appl Environ Microbiol 67: 2395–2403.
Eller G, Krüger M, Frenzel P . (2005). Comparing field and microcosm experiments: a case study on methano- and methylo-trophic bacteria in paddy soil. FEMS Microbiol Ecol 51: 279–291.
Evershed RP, Crossman ZM, Bull ID, Mottram H, Dungait JAJ, Maxfield PJ et al. (2006). C-Labelling of lipids to investigate microbial communities in the environment (Review). Curr Opinion Biotechnol 17: 72–82.
Groot TT, VanBodegom PM, Harren FJM, Meijer HAJ . (2003). Quantification of methane oxidation in the rice rhizosphere using 13C-labelled methane. Biogeochemistry 64: 355–372.
Hanson RS, Hanson TE . (1996). Methanotrophic bacteria. Microbiol Rev 60: 439–471.
Henckel T, Friedrich M, Conrad R . (1999). Molecular analyses of the methane-oxidizing microbial community in rice field soil by targeting the genes of the 16S rRNA, particulate methane monooxygenase, and methanol dehydrogenase. Appl Environ Microbiol 65: 1980–1990.
Hoffmann T, Horz HP, Kemnitz D, Conrad R . (2002). Diversity of the particulate methane monooxygenase gene in methanotrophic samples from different rice field soils in China and the Philippines. Syst Appl Microbiol 25: 267–274.
Holmes AJ, Costello A, Lidstrom ME, Murrell JC . (1995). Evidence that particulate methane monooxygenase and ammonia monooxygenase may be evolutionarily related. FEMS Microbiol Lett 132: 203–208.
Horz HP, Rich V, Avrahami S, Bohannan BJM . (2005). Methane-oxidizing bacteria in a California upland grassland soil: diversity and response to simulated global change. Appl Environ Microbiol 71: 2642–2652.
Horz HP, Yimga MT, Liesack W . (2001). Detection of methanotroph diversity on roots of submerged rice plants by molecular retrieval of pmoA, mmoX, mxaF, and 16S rRNA and ribosomal DNA, including pmoA-based terminal restriction fragment length polymorphism profiling. Appl Environ Microbiol 67: 4177–4185.
Huber T, Faulkner G, Hugenholtz P . (2004). Bellerophon: a program to detect chimeric sequences in multiple sequence alignments. Bioinformatics 20: 2317–2319.
Islam T, Jensen S, Reigstad LJ, Larsen O, Birkeland NK . (2008). Methane oxidation at 55°C and pH 2 by a thermoacidophilic bacterium belonging to the Verrucomicrobia phylum. Proc Natl Acad Sci USA 105: 300–304.
Krüger M, Frenzel P . (2003). Effects of N-fertilisation on CH4 oxidation and production, and consequences for CH4 emissions from microcosms and rice fields. Glob Chang Biol 9: 773–784.
Krüger M, Frenzel P, Conrad R . (2001). Microbial processes influencing methane emission from rice fields. Glob Chang Biol 7: 49–63.
Li J, Wang MX, Huang Y, Wang YS . (2002). New estimates of methane emissions from Chinese rice paddies. Nutrient Cycling in Agroecosystems 64: 33–42.
Liesack W, Schnell S, Revsbech NP . (2000). Microbiology of flooded rice paddies (Review). FEMS Microbiol Rev 24: 625–645.
Lilkanen A, Martikainen PJ . (2003). Effect of ammonium and oxygen on methane and nitrous oxide fluxes across sediment-water interface in a eutrophic lake. Chemosphere 52: 1287–1293.
Lu YH, Abraham WR, Conrad R . (2007). Spatial variation of active microbiota in the rice rhizosphere revealed by in situ stable isotope probing of phospholipid fatty acids. Environ Microbiol 9: 474–481.
Lu YH, Conrad R . (2005). In situ stable isotope probing of methanogenic archaea in the rice rhizosphere. Science 309: 1088–1090.
Lu YH, Rosencrantz D, Liesack W, Conrad R . (2006). Structure and activity of bacterial community inhabiting rice roots and the rhizosphere. Environ Microbiol 8: 1351–1360.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al. (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Lueders T, Manefield M, Friedrich MW . (2004). Enhanced sensitivity of DNA- and rRNA-based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ Microbiol 6: 73–78.
Macalady JL, McMillan AMS, Dickens AF, Tyler SC, Scow KM . (2002). Population dynamics of type I and II methanotrophic bacteria in rice soils. Environ Microbiol 4: 148–157.
Mancinelli RL . (1995). The regulation of methane oxidation in soil. Ann Rev Microbiol 49: 581–605.
Manefield M, Whiteley AS, Ostle N, Ineson P, Bailey MJ . (2002). Technical considerations for RNA-based stable isotope probing: an approach to associating microbial diversity with microbial community function. Rapid Commun Mass Spectrom 16: 2179–2183.
McDonald IR, Murrell JC . (1997). The particulate methane monooxygenase gene pmoA and its use as a functional gene probe for methanotrophs. FEMS Microbiol Lett 156: 205–210.
McDonald IR, Radajewski S, Murrell JC . (2005). Stable isotope probing of nucleic acids in methanotrophs and methylotrophs: a review. Org Geochem 36: 779–787.
Miller LG, Sasson C, Oremland RS . (1998). Difluoromethane, a new and improved inhibitor of methanotrophy. Appl Environ Microbiol 64: 4357–4362.
Mohanty SR, Bodelier PLE, Conrad R . (2007). Effect of temperature on composition of the methanotrophic community in rice field and forest soil. FEMS Microbiol Ecol 62: 24–31.
Nold SC, Boschker HTS, Pel R, Laanbroek HJ . (1999). Ammonium addition inhibits 13C-methane incorporation into methanotroph membrane lipids in a freshwater sediment. FEMS Microbiol Ecol 29: 81–89.
Noll M, Matthies D, Frenzel P, Derakshani M, Liesack W . (2005). Succession of bacterial community structure and diversity in a paddy soil oxygen gradient. Environ Microbiol 7: 382–395.
Nouchi I, Mariko S, Aoki K . (1990). Mechanisms of methane transport from the rhizosphere to the atmosphere through rice plants. Plant Physiol 94: 59–66.
Pester M, Friedrich MW, Schink B, Brune A . (2004). pmoA-Based analysis of methanotrophs in a littoral lake sediment reveals a diverse and stable community in a dynamic environment. Appl Environ Microbiol 70: 3138–3142.
Pol A, Heijmans K, Harhangi HR, Tedesco D, Jetten MSM, Op den Camp HJM . (2007). Methanotrophy below pH 1 by a new Verrucomicrobia species. Nature 450: 874–878.
Roslev P, Iversen N . (1999). Radioactive fingerprinting of microorganisms that oxidize atmospheric methane in different soils. Appl Environ Microbiol 65: 4064–4070.
Schütz H, Schröder P, Rennenberg H . (1991). Role of plants in regulating the methane flux to the atmosphere. In: Sharkey TD, Holland EA, Mooney HA (eds). Trace Gas Emissions by Plants. Academic Press: San Diego. pp 29–63.
Schütz H, Seiler W, Conrad R . (1989). Processes involved in formation and emission of methane in rice paddies. Biogeochemistry 7: 33–53.
Seiler W, Holzapfel-Pschorn A, Conrad R, Scharffe D . (1984). Methane emission from rice paddies. J Atmos Chem 1: 241–268.
Shrestha M, Abraham WR, Shrestha PM, Noll M, Conrad R . (2008). Activity and composition of methanotrophic bacterial communities in planted rice soil studied by flux measurements, analyses of pmoA gene and stable isotope probing of phospholipid fatty acids. Environ Microbiol 10: 400–412.
Vandenkoornhuyse P, Mahe S, Ineson P, Staddon P, Ostle N, Cliquet JB et al. (2007). Active root-inhabiting microbes identified by rapid incorporation of plant-derived carbon into RNA. Proc Natl Acad Sci USA 104: 16970–16975.
Acknowledgements
We thank Esther Surges for GC and IRMS measurements, Professor Peter Frenzel (MPI) for helpful discussion and Professor Zongxiu Sun (China National Rice Research Institute) for access to the rice fields. This work was done in the framework of a Max Planck Partner Group.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Qiu, Q., Noll, M., Abraham, WR. et al. Applying stable isotope probing of phospholipid fatty acids and rRNA in a Chinese rice field to study activity and composition of the methanotrophic bacterial communities in situ. ISME J 2, 602–614 (2008). https://doi.org/10.1038/ismej.2008.34
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2008.34
Keywords
This article is cited by
-
Type I methanotrophs dominated methane oxidation and assimilation in rice paddy fields by the consequence of niche differentiation
Biology and Fertility of Soils (2024)
-
Unique high Arctic methane metabolizing community revealed through in situ 13CH4-DNA-SIP enrichment in concert with genome binning
Scientific Reports (2022)
-
Discrepancy in exchangeable and soluble ammonium-induced effects on aerobic methane oxidation: a microcosm study of a paddy soil
Biology and Fertility of Soils (2021)
-
Atmospheric Methane Oxidizers Are Dominated by Upland Soil Cluster Alpha in 20 Forest Soils of China
Microbial Ecology (2020)
-
Changes in methane oxidation ability and methanotrophic community composition across different climatic zones
Journal of Soils and Sediments (2019)