Abstract
Norway spruce of northern Europe expanded at the end of the last glacial out of one refugium in Russia. To provide a detailed insight into how the genetic structure in the northern European lineage of this species has been shaped by postglacial migration, recurrent pollen flow and marginality, we here compare variation at seven highly variable nuclear microsatellite loci in 37 populations (1715 trees) with mitochondrial DNA variation. Microsatellite diversity was high (HE=0.640) and genetic differentiation was low (FST=0.029). The microsatellite structure supported a mitochondrial DNA (mtDNA)-based hypothesis of two migration routes out of a single Russian refugium; one northwestern over Finland to northern Scandinavia, and one southwestern across the Baltic Sea into southern Scandinavia. Microsatellite diversity was maintained along the southwestern migration routes, whereas a significant decrease was observed towards the north. In contrast, the mtDNA diversity suggested higher amounts of historical gene flow towards the north than along the southwestern migration route. This suggests that the loss of nuclear diversity after postglacial colonization has been efficiently replenished by pollen-mediated gene flow in the south. Towards the north, smaller effective population size because of more limited seed and pollen production may have caused decreased nuclear diversity and increased inbreeding, reflecting the ecological marginality of the species in the north.
Similar content being viewed by others
Introduction
Phylogeographical inferences have in earlier works typically relied on organelle markers representing single gene histories. To fully address the population history of an organism, several distinct genealogies from independent genetic markers are needed (Ballard and Whitlock, 2004). By combining mitochondrial (mt) or chloroplast (cp) DNA sequences with nuclear markers, demographic processes acting on different time scales will be captured because of organelle and different modes of inheritance of nuclear markers, effective population size and mutation rate (Hewitt, 2001, Semerikov and Lascoux, 2003).
A number of recent studies contrasting genetic population structures at organelle and nuclear loci have gained an improved understanding of past and present population demographic events for many species (Gamache et al., 2003; Heuertz et al., 2004a, 2004b; Magri et al., 2006). In these studies, maternally inherited seed-dispersed organelle markers typically revealed distinct genetic groups associated with glacial refugia and colonization routes. The increased information content that resulted from the use of multiple nuclear markers in these studies, did indeed provide a better resolution than maternally inherited markers alone, and facilitated the detection of additional refugia and meeting zones. Moreover, differentiation identified with maternally inherited markers and nuclear markers has often been found to be independent (Petit et al., 2005), and populations fixed for a single organelle haplotype do not necessarily show low diversity at nuclear markers (Heuertz et al., 2004b; Magri et al., 2006).
For Norway spruce (Picea abies L. Karst), a comprehensive data set of the maternally inherited mtDNA marker, nad1, and palaeoecological data, were recently combined to infer glacial refugia, primary routes of postglacial colonization and genetic consequences of postglacial colonization (Tollefsrud et al., 2008b). Norway spruce, one of the most important ecological and economical forest tree species in Europe, occurs in two disjunct natural ranges, a northern and a southern (Schmidt-Vogt, 1977). These disjunct ranges correspond to two divergent genetic lineages that have likely been separated over several periods of glaciation (Lagercrantz and Ryman, 1990; Vendramin et al., 2000; Heuertz et al., 2006; Tollefsrud et al., 2008b). As inferred from both palaeo and mtDNA data, postglacial colonization of the southern range took place from several distinct refugia, whereas the northern range was colonized from a single, large refugium situated in Russia (Terhürne-Berson, 2005; Latalowa and van der Knaap, 2006; Tollefsrud et al., 2008b).
On the basis of mtDNA results, the northern lineage of Norway spruce consists of a single, large gene pool (Tollefsrud et al., 2008b). A shallow substructuring of this gene pool indicates that postglacial colonization followed two main migration routes: one northwestern route from Russia to Finland, following the mainland north of the Gulf of Bothnia and further to Norway and Sweden, and one southwestern route over the Baltics, crossing directly over the Baltic Sea into southern Scandinavia (Tollefsrud et al., 2008b). Today, the Russian plain populations of Norway spruce exhibit high mtDNA gene diversity and are little differentiated. During postglacial colonization, mtDNA diversity generally decreased away from the Russian refugium, but substantial diversity was nevertheless maintained over large distances, suggesting phalanx colonization in concordance with the pollen data (Giesecke and Bennett, 2004). Severe bottlenecks seem to have occurred mainly in the northernmost part of Scandinavia and in western Finland, where the populations typically hold a single mtDNA haplotype (Tollefsrud et al., 2008b).
Mitochondrial diversity in Norway spruce does not seem to be influenced by its interfertile congener, Siberian spruce (Picea obovata Lebed.). Sizing and sequence analysis of nad1 from Siberian spruce samples show a distinct division between Norway spruce and Siberian spruce east of the Ural Mountains, following the river Ob (Tollefsrud et al., 2008a). On the other hand, pollen flow from Siberian spruce may have influenced the nuclear diversity in Norway spruce as Siberian spruce cp haplotypes are found at low frequencies in Norway spruce stands (Tollefsrud et al., 2008a).
To obtain deeper insights into the past and present processes shaping the genetic structuring that has taken place in the northern lineage of Norway spruce, we here investigate nuclear microsatellite variation and contrast it with the earlier reported mtDNA variation (Tollefsrud et al., 2008b). This will give a better understanding of the relative effects of postglacial colonization versus the past and present pollen flow, as the diversity of highly variable microsatellites will recover more quickly from founder events owing to larger effective population size and spread by both seeds and pollen. We first examine whether nuclear microsatellite variation provides additional and complementary information regarding the migration routes. Second, we investigate how nuclear diversity and differentiation is distributed in relation to distance from the refugium and to peripheral populations.
Materials and methods
Population sampling
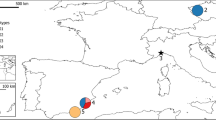
Twigs from Picea abies were collected from an average of 46 trees, each separated by at least 20 m, from each of 37 putatively autochthonous stands (Figure 1), and subjected to microsatellite analysis (in total 1715 trees; see Appendix A). Mitochondrial DNA variation was obtained from an average of 17 trees per stand (Tollefsrud et al., 2008b and Appendix A; in total 644 trees). This sample covered the North European distribution area (Schmidt-Vogt, 1977) and the West coast of Norway, which is outside the continuous range (Fægri, 1950). To ensure collection from natural stands as far as possible, we gathered information on stand history from local foresters’ offices and only included stands with mtDNA haplotypes belonging to the northern European lineage, as reported in Tollefsrud et al. (2008b).
Geographical distribution of the six genetic (BAPS) groups inferred from variation at seven nuclear microsatellite loci in 37 Picea abies populations (a). The green outline illustrates the distribution of the northern lineage of Norway spruce (after Schmidt-Vogt, 1977). The neighbour-joining tree (b) is based on Cavallia-Sforza and Edwards chord distances between the BAPS groups. Bootstrap values are given in percentages and are based on 1000 replications. Stand numbers are according to Appendix A.
Test for null alleles
To empirically test for the presence of null alleles at microsatellite loci, trees along with their seeds were collected from a seed orchard of northern European origin. Conifer seeds have haploid megagametophyte storage tissue with the same genotype as the corresponding ovule. For trees showing a homozygote profile for an allele, DNA from eight endosperms was isolated and tested for the potential presence of null alleles. Null allele heterozygotes were able to be detected because the haploid megagametophytes of a tree segregate for the presence and absence of the corresponding microsatellite band.
DNA extraction, amplification and sizing
DNA was extracted from frozen needles sampled from adult trees and from endosperm tissue for the null allele test, using the DNeasy 96 Plant Kit or the DNeasy Plant Mini Kit (Qiagen, Hilden, Germany). A total of 30 microsatellite primers from Pfeiffer et al. (1997) and Scotti et al. (2002a, 2002b) were tested for variation and peak quality. Seven primers, EATC1D02A, EATC1B01, EATC2B02, EATC1E3, EATC2G05 (Scotti et al., 2002a), EAC2C08 (Scotti et al., 2002b), SPAC1F7 (Pfeiffer et al., 1997), were selected for further screening. These loci were variable and provided a high quality amplification product with a minimum of slippage. The loci EATC1D02A, EATC1B01, EATC2B02, EAC2C08 and SPAC1F7 were amplified in 25 μl containing 1 × PCR buffer (GE Healthcare, Piscataway, NJ, USA), 2.5 mM MgCl2, 0.2 mM of each dNTP, 0.2 μM of each primer, 1 U Taq DNA polymerase T. aquaticus (GE Healthcare) and ca. 30 ng DNA template. The loci EATC1D2A, EATC1B01, EATC2B02 and EAC2C08 were amplified with the touchdown profile described in Scotti et al. (2002a). The profile used for SPAC1F7 followed Pfeiffer et al. (1997). The loci EATC1E3 and EATC2G05 were amplified together in 20 μl containing a 1 × Qiagen multiplexing kit, 0.2 μM of each primer and ca. 20 ng DNA template using the following protocol: 15 min at 95 °C, 28 cycles of 94 °C for 30 s, 58 °C for 90 s, 72 °C for 60 s and a final extension at 60 °C for 30 min. DNA from endosperm tissue was amplified twice to ensure that non-amplified products were because of null alleles and not because of PCR failure. PCR products labelled with various fluorescent dyes (FAM, HEX and TAMRA) were loaded on the capillary system sequencer, MegaBACE1000 (GE Healthcare), with the size standard MegaBACE ET400 (GE Healthcare). Peaks were analysed and fragment lengths determined with the MegaBACE FRAGMENT PROFILER software version 1.2 (GE Healthcare). All peaks and binning were manually checked.
The mt marker used in this study comprises a fragment of the second intron of the nad1 gene including two highly variable minisatellites of 32 and 34 base pairs (bp) and insertion/deletions and mutations in the flanking regions of the tandem repeats. Variation in the minisatellite-flanking sequences separates northern populations from the southern ones, whereas copy–number variation in the two minisatellites is very variable on a regional scale. Populations in the northern range mainly show variation in the 34 bp repeat (Sperisen et al., 2001; Tollefsrud et al., 2008b).
Variation in the nad1 fragment from 30 of the stands used in this study had been screened earlier from at least 13 trees per stand (Tollefsrud et al., 2008b). For the seven remaining stands, each represented by at least 14 trees, variation in nad1 was screened for this study. The nad1 fragment was amplified and cut with the restriction enzyme, EcoRV, following Tollefsrud et al. (2008b). Parts of the samples were sized on an ABI 310 genetic analyzer (Applied Biosystems, Foster City, CA, USA) as described in Tollefsrud et al. (2008b). The remaining samples were run on a MegaBACE1000 (GE Healthcare) using the size standard MegaBACE ET900 (GE Healthcare). Peaks were analysed and fragment lengths were determined using the MegaBACE FRAGMENT PROFILER software version 1.2 (GE Healthcare). Sixteen samples with known size were used as control samples on the MegaBACE1000 to calibrate differences in size resolution between the two sizing systems.
Genetic diversity and differentiation
Linkage disequilibrium for all microsatellite loci pairs in each population was tested in the programme FSTAT (Goudet, 2001), using randomization. Microsatellite diversity was measured as gene diversity (expected heterozygosity, HE) (Nei, 1987) and allelic richness (AS) using FSTAT. Allelic richness was calculated using rarefaction (Hurlbert, 1971) standardized to the minimum sample size of 25 diploid individuals. The total number of alleles and the allele range per locus were also recorded. Conformity to Hardy–Weinberg (HW) proportions was tested using the exact test (Guo and Thompson, 1992) based on Markov chain iterations as implemented in GENEPOP v4.0 (Rousset, 2008). The extent of deviation from HW proportions was evaluated using FIS estimates across loci for each stand. The influence of each locus on the stand level of FIS was examined to test whether some loci influenced the FIS level more than others.
Null allele frequencies for each microsatellite locus and each stand were estimated following the expectation maximum algorithm of Dempster et al. (1977) using FREENA (Chapuis and Estoup, 2007). This algorithm was chosen because it provided the most accurate estimate of several algorithms tested in Chapuis and Estoup (2007).
For the microsatellite data, an unbiased estimate of Wright's fixation index theta (FST) (Weir and Cockerham, 1984) was calculated as a measure of genetic differentiation in FSTAT. The significance of genetic differentiation among pairs of stands was tested by permutating the genotypes a 1000 times among samples. FST may be overestimated in the case of null alleles (Chapuis and Estoup, 2007) and underestimated when within-population heterozygosity is high (Hedrick, 2005; Jost, 2008). We controlled for the potential effect of null alleles on genetic differentiation by calculating FST values using the excluding null allele (ENA) method by Chapuis and Estoup (2007) in FREENA. To take high levels of heterozygosity into account, standardized FST (F'ST) values following Hedrick (2005) were calculated dividing the uncorrected FST values by its upper limits (F′ST=FST/FST max). To calculate FST max, we used RecodeData version 0.1 (Meirmans, 2006) to create an FSTAT file with maximally differentiated populations (that is, the data file was recoded such that all alleles were population specific). Mantel tests between pairwise FST values, pairwise ENA-corrected FST values and pairwise F'ST values were performed in ARLEQUIN 3.11 (Excoffier et al., 2005) using 1000 random permutations to evaluate whether the different FST calculations were significantly correlated.
Analyses of molecular variance (AMOVA) partitioning variation among genetic groups, among populations and within populations were carried out using ARLEQUIN 3.11. Critical significance levels of multiple tests were adjusted for by sequential Bonferroni correction (Rice, 1989).
For the mt data, within population genetic diversity (HS), total genetic diversity (HT) and genetic differentiation among populations (GST) were calculated according to Pons and Petit (1995) using the programme, CONTRIB. Allelic richness (AS) based on a rarefaction factor of 13 was calculated in FSTAT. Taking the different levels of variation obtained from the mt and nuclear markers into account, standardized GST values (G′ST) following Hedrick (2005) were calculated using RecodeData version 0.1 in conjunction with FSTAT.
Population genetic structure
The population structure determined using microsatellite data was initially analysed adopting three statistical approaches: the Bayesian methods implemented in the programme BAPS 4.14 (Corander et al., 2003, 2004) and STRUCTURE 2.2 (Pritchard et al., 2000; Falush et al., 2003, 2007), and Genetic Landscape Shape analysis (Miller, 2005). The population structure based on mt data has been presented by Tollefsrud et al. (2008b); here, we only performed Genetic Landscape Shape analysis of the mtDNA data to facilitate direct comparison between the geographical patterns of differentiation observed between the two marker types.
BAPS and STRUCTURE attempt to reveal the population genetic structure by placing individuals or predefined groups in K numbers of clusters with optimal HW and gametic phase equilibrium among clusters. K is not chosen in advance, but is varied within a reasonable range. BAPS 4.14 was first run using individual clustering, but this resulted in an unreasonably high number of clusters. Because of this, we used our predefined stands as units in the analysis. The number of clusters, K, was set from 1 to 37 with ten replicates for each K in five independent runs. We ran STRUCTURE 2.2 using both the admixture model and the dominant marker model using the option ‘recessive alleles’ to deal with null alleles. We used a burn-in period of 5 × 105 and 2 × 106 iterations with correlated allele frequencies under an admixture model. K was set from 1 to 10, and 10 replicates were run for each K-value. Similarity among the runs for the same K was calculated according to Nordborg et al. (2005), using an R script written by D Ehrich available at http://www.nhm.uio.no. We identified the number of groups as the value of K, in which the increase in likelihood began to flatten out and the results of replicate runs were identical.
The Genetic Landscape Shapes analyses were produced using the programme, Alleles In Space (AIS; Miller, 2005). The procedure was initiated by constructing a Delaunay triangulation network among all the sampling coordinates. Average interindividual genetic distances were calculated between the stands connected in the network. Considering that there was substantial variation in geographical distances between the sampling areas, we used distance-corrected genetic distances. Next, a simple interpolation procedure was used to infer genetic distances at locations on a uniformly spaced grid overlaying the entire sample landscape (for a more detailed description, see Miller et al., 2006). A three-dimensional surface plot of the set of interpolated genetic distances was produced, where X and Y coordinates corresponded to geographical locations within the Delaunay network and surface plot heights (Z) to genetic distance. We used sequences as input matrix for the nad1 data set, with coding of the insertion/deletions and the repeated sequences following Tollefsrud et al. (2008b). Gaps were treated as a fifth character in AIS.
Genetic distances (DCE, Cavallia-Sforza and Edwards) among stands or genetic clusters obtained from the Bayesian cluster analysis were calculated in the MICROSATELLITE ANALYSER (MSA 4.05) programme (Dieringer and Schlötterer, 2003). For statistical support, loci were bootstrapped a 1000 times. A neighbour-joining (NJ) tree was constructed on the basis of DCE distances using NEIGHBOR and TREEVIEW in the PHYLIP Software package (Felsenstein, 2004).
We traced the patterns of changes in diversity parameters within stands following postglacial expansion out of the Russian plain (Latalowa and van der Knaap, 2006; Tollefsrud et al., 2008b) by calculating the correlations (Pearson's r) for the genetic parameters obtained from the microsatellites (HE NUC, AS NUC, and FIS NUC), mtDNA (HS MT, HT MT, and AS MT), longitude, latitude and distance from the refugial gene pool. The refugial gene pool was represented by the stand, RUS3 (41.00° E, 57.58° N; Tollefsrud et al., 2008b). Pearson's r was calculated using R version 2.5.0 (R Development Core Team, 2007).
To test whether the genetic differentiation followed isolation by distance (IBD), the correlation between FST/(1-FST) and the natural logarithm of geographical distances was calculated with the Mantel test suggested by Rousset (1997), using ARLEQUIN 3.11, with 10 000 random permutations.
We tested whether the populations departed from mutation-drift equilibrium using the BOTTLENECK programme (Cornuet and Luikart, 1996; Luikart and Cornuet, 1998; Piry et al., 1999). The significance of potential bottlenecks was assessed using the Sign and the Wilcoxon tests for heterozygosity excess, of which the Wilcoxon test is considered the most powerful and robust (Piry et al., 1999).
Results
Genetic diversity and differentiation
Significant linkage disequilibrium between the pairs of microsatellite loci within stands was only detected in five out of 777 tests (α=0.05) after sequential Bonferroni correction (k=37). All loci were thus considered to be genetically independent when analysed further.
The nuclear genetic variation was high overall (Table 1). The mean gene diversity over loci was 0.640 and the mean number of alleles per locus was 22. The gene diversity (HE) showed high variation among loci, ranging from 0.279 at EATC1E3 to 0.903 at EAC2C08. The number of alleles per locus ranged from 9 (EATC1E3) to 40 (EAC2C08). A total of 154 different alleles were detected, of which 31 occurred at an overall frequency >0.05. Twenty-seven private alleles were found, distributed over 19 stands. The private alleles were mostly at the extremes of the allele size distribution and occurred at very low frequencies. Nuclear gene diversity and allelic richness per stand ranged from 0.492 to 0.688 and from 4.967 to 9.586, respectively (Appendix A).
The single-locus tests for HW proportions over stands showed significant departure (α=0.05) in 163 out of 259 cases, distributed over all seven loci. One hundred and thirty of them were significant after sequential Bonferroni correction (α=0.05, k=7; Appendix A). Significant multilocus deviation from Hardy-Weinberg Equilibrium (HWE) was observed in all stands after sequential Bonferroni correction (α=0.05, k=37). Assuming HWE, the estimated null allele frequencies over stands varied from 0.288 at EATC1D2A to 0.042 at EAC2C08 (Table 1). At four of the loci, the null allele frequency estimates were very low (0.076–0.042; Table 1). The expected number of null homozygotes ((Pn)2 * n; Table 1) was lower than the observed value for four loci. In the direct ‘count’ of null alleles in the endosperm tissue from homozygous mother trees, null alleles were observed for the five loci EATC1D2A, EATC1B02, EATC1E3, EATC2G05 and SPAC1F7. Unfortunately, because the number of homozygote trees for which we had endosperms was low (n ranging from 2–13 per locus) and because only a fraction of the potential alleles were detected (n ranging from 2–7), no reliable empirical estimate of the null allele frequencies could be determined. In the data set obtained, EATC1D2A had six null alleles, EATC1B02 and EATC1E3 had two each and SPAC1F7 and EATC2G05 had one each. All loci that had a null allele showed a 1:1 segregation (determined by the χ2-test, data not shown).
The overall genetic differentiation at the microsatellite loci was low (FST=0.029, s.d.=0.004), though highly significant (P<0.001). After ENA correction, the overall FST was slightly lower (FST ENA=0.026), and the standardized FST was slightly higher (F′ST=0.071). The pairwise FST values ranged from 0.000 to 0.147. The highest values were obtained for the northernmost stand (NN17) and for two stands outside the continuum on the West coast of Norway (SN36 and SN37). Significant (P<0.05) genetic differentiation was found in 518 out of 666 pairs after sequential Bonferroni correction. The ENA-corrected pairwise FST estimates were very similar and strongly correlated with the uncorrected pairwise FST values (R2=0.935, P<0.00001) in the Mantel test, suggesting that all stands were similarly affected by null alleles. The pairwise standardized Hedrick (2005) F′ST estimates were generally elevated, but strongly correlated with the uncorrected pairwise FST values (R2=0.899, P<0.00001) in the Mantel test.
In the 37 stands, we detected 10 mt haplotypes (721, 754, 755, 789, 819, 823, 857, 891, 925 and 959; Appendix A). None of these haplotypes was new compared with Tollefsrud et al. (2008b), and they all belonged to the northern lineage of Norway spruce (Tollefsrud et al., 2008b). In the present data set, mt mean gene diversity (HS) was 0.238, mean allelic richness (AS) was 2.426, total gene diversity (HT) was 0.349 and differentiation among populations (GST) was 0.317. G′ST for the mt data was about five times higher than that for the nuclear data (G′ST MT=0.400, G′ST NUC=0.081).
Genetic subdivision
The BAPS analysis of the microsatellite data identified six clusters (Figure 1). The largest group contained all stands from Russia, the Baltics and Finland, and one stand from northern Sweden. The second largest group contained most of the stands from southern Scandinavia. These two groups clustered together with high support in the NJ tree of the BAPS groups (Figure 1b). The third largest BAPS group contained four stands from central Sweden. The three remaining BAPS groups contained single stands located at the northern and western range margins of the Norway spruce. These three single stands were the most divergent ones within the NJ tree of the BAPS groups (Figure 1b). In the AMOVA analysis, only 1.44% was explained by the BAPS groups, 1.77% was explained by variation among stands within the groups and 96.84% of the variation was explained by within-stand variation (P<0.001). All of the variance components were highly significant (P<0.001).
STRUCTURE failed to reveal any biologically reasonable groups. Under both the admixture model and the dominant marker model, three groups were chosen, as the increase in the likelihood flattened out at K>3 and similarity among the runs for K=3 was close to 1. All populations were however almost equally partitioned among the three groups.
The NJ tree of individual stands (Figure 2) showed a structure that was similar to the BAPS groups, but little support was obtained. One large cluster consisted of stands from southern Scandinavia and the Baltics, and two stands from western Russia. A second large cluster consisted of Russian, Finnish and northern Scandinavian stands. An additional cluster, not observed in the BAPS analysis, consisted of the northernmost stand, NN17, together with NS18, FIN16 and one stand from the southern Russian Urals (RUS1).
Neighbour-joining tree for 37 Picea abies stands based on nuclear microsatellite variation at seven loci. Distances between populations are based on Cavallia-Sforza and Edwards chord distances. The lines drawn around the two main clusters are based on geographical origins of the stands. BAL: Baltic States and Belarus, CS: central Sweden, FIN: Finland, NN: northern Norway, NS: northern Sweden, RUS: Russia, SN: southern Norway, SS: southern Sweden. Bootstrap values >50% based on 1000 replications are given. Stand numbers are according to Appendix A and Figure 1.
Overall, the genetic landscape shape analysis of the microsatellite data was consistent with the BAPS and NJ analyses (Figure 3a). The eastern populations showed little differentiation. Along the southern edge from east to west, there was a largely smooth genetic surface indicating low barriers to gene flow. Two primary regions of elevated genetic distances were revealed in the west, one along the northern edge (central Sweden) and one in the westernmost part of the range (southwestern Norway). By contrast, the genetic landscape shape analysis of the mtDNA data (Figure 3b) revealed a high peak in northern Finland and northern Norway (Figure 3b). For the mtDNA data, as for the microsatellite data, a smooth surface of genetic distances was evident along the southern edge from east to west.
Genetic Landscape Shape interpolation analysis for 37 Picea abies stands based on (a) nuclear microsatellite variation at seven loci, and (b) mitochondrial DNA variation in the second intron of nad1. The x and y axes show the geographic locations within a Delaunay triangulation network constructed among the sampled stands. Surface plot heights reflect average between interindividual genetic distances.
Geographic trends
We found clear geographical trends in genetic diversity for both the nuclear DNA (Table 2 and Figure 4) and mtDNA (Table 2). The correlations between the nuclear parameter AS NUC and the three mtDNA parameters were positive and significant, indicating a general coherence among the nuclear and the mtDNA diversity patterns. The mtDNA parameters and the nuclear parameter AS NUC decreased significantly with distance from the oldest region as identified by pollen data (Tollefsrud et al., 2008b). Still, with respect to longitude and latitude, different trends in the mtDNA and the nuclear DNA diversity were evident. Although the mt parameters were significantly negatively correlated with increasing distance from the oldest population (RUS3) and with decreasing longitude, nuclear gene diversity (HE NUC) and allelic richness (AS NUC) did not change significantly with longitude (Table 2). With increasing latitude, both HE NUC and AS NUC decreased significantly, whereas only HT MT decreased significantly northwards. Negative correlations between latitude and AS NUC were observed at all loci, and for HE NUC at five loci (results not shown).
Population gene diversity (a; HE) and allelic richness (b; AS) in Picea abies based on nuclear microsatellites, plotted as deviation from the mean values. Higher than average values are represented by grey circles, lower than average values are represented by white circles. The green outline illustrates the distribution of the northern lineage of Norway spruce (after Schmidt-Vogt, 1977).
FIS NUC over all loci significantly increased with latitude, suggesting increased mating within stands northwards (assuming that the occurrence of null alleles affected the stands similarly, as shown in the Mantel test of pairwise FST and ENA corrected FST values). Considering that significant positive correlations were observed at only three out of seven loci (EATC1D02A, r=0.417**, EATC2G05 r=0.621*** and SPAC1F7 r=0.428**), the increase in FIS northward should be interpreted with caution.
Stands with high nuclear gene diversity typically occurred in the south, including the western part of the Russian plains, the Baltics and southern Scandinavia (Figure 4b; HE). Allelic richness largely followed the same pattern, except that the stands from central Finland also had above-average allelic richness (Figure 4a; AS). Most stands in the northernmost part of the range as well as in central Sweden had below-average gene diversity and allelic richness. In the most recently colonized areas in southern and southwestern Norway, allelic richness was below average (Figure 4a), whereas the gene diversity in several of these stands was above average (Figure 4b).
Weak IBD and no population bottlenecks for the nuclear microsatellites
The correlation between nuclear genetic differentiation and geographical distance showed that the IBD pattern was very weak (R2=0.032, P=0.043). In the BOTTLENECK analysis, all populations showed heterozygote deficiency at >50% of the loci in the Sign test. No populations had significant heterozygote excess in the Wilcoxon test performed under the infinite, the two phase or the stepwise mutation model. All populations were thus characterized by significant heterozygote deficiency, indicating expansion rather than contraction. Evidence for recent bottlenecks was thus not detected in any stands. It is possible however, that bottlenecks might not have been detected due to null alleles causing lower levels of observed heterozygosity.
Discussion
The northern European stands of Norway spruce were characterized by high diversity and low differentiation at the nuclear microsatellite loci. This manifests high levels of past and present gene flow over the northern European range of the species, corroborating earlier results (Lagercrantz and Ryman, 1990; Vendramin et al., 2000; Tollefsrud et al., 2008b). The overall congruence between the structure at the nuclear loci and the mtDNA is likely to mirror common historical events in the populations. The deeper genetic structure observed in the seed-dispersed mt genome suggests that postglacial colonization through seeds established an underlying genetic structure that is still present and detectable in the nuclear genome, despite extensive pollen flow over large distances. Postglacial colonization nevertheless affected mt and nuclear diversity differently, owing to the different mode of transmission and dispersal of these markers as well as to the difference in their effective population size.
Congruent and refined structure at nuclear microsatellites compared with mt nad1
In the oldest regions of Norway spruce in northern Europe, as inferred from the pollen record (Tollefsrud et al., 2008b), microsatellite diversity was high and differentiation was very low. This is in agreement with the results of the earlier mtDNA analysis (Tollefsrud et al., 2008b). Large refugia are expected to exhibit such a genetic structure ( Hewitt, 2001) and strongly support the hypothesis that Norway spruce survived in a single large refugium in Russia during the last glacial maximum.
The statistically significant differences in nuclear allele frequencies, as reflected in the BAPS groups and partly in the NJ trees of the stands, provide independent support for the mtDNA-based hypothesis of a northwestern and a southwestern migration route out of the Russian refugium. A geographical subdivision on account of allele frequency differences (and not geographical differences in heterozygosity levels) is also supported by the high correlation among uncorrected and standardized FST values, evident from the Mantel test. Although only a small fraction of the total genetic variation was explained by the BAPS groups, the NJ tree of the BAPS groups (Figure 1b) provides additional support for the hypothesised migration routes, especially the southwestern route. Moreover, genetic landscape shape analysis revealed a smooth surface of genetic distances from Russia to southern Scandinavia, indicating low barriers to gene flow across the Baltic Sea. The finings of macrofossils dated to the early Holocene, as well as that of a living tree dated to be 9500 years old in southern Sweden (Kullman, 2008), suggest that the migration along the southwestern route across the Baltic Sea was initiated early. Dispersal could either have taken place through have taken place frequent long-distance trans-oceanic dispersal or possibly across land during the Ancylus regression of the Baltic Sea (9000–8000 years before present) when there was a mainland connection between southern Sweden and the Baltic coast (Björck, 1995).
From the microsatellite data, an additional migration from the eastern Urals to northern Fennoscandia may be inferred from a cluster in which a stand from the southern Urals grouped together with three northern Fennoscandian stands. This pattern may be because of pollen-mediated introgression from the Siberian spruce, as pollen flow from the Siberian spruce to the Norway spruce is suggested by paternally inherited cp haplotypes (Tollefsrud et al., 2008a).
Our microsatellite data for Norway spruce showed no evidence of bottlenecks. This is largely consistent with the pollen record, except for the northernmost populations where the pollen record has suggested significant bottlenecking (Giesecke and Bennett, 2004). Drift or founder events may still explain some of the genetic substructure that we find within northern European Norway spruce. For example, the divergence of the central Swedish stands as revealed in the NJ and BAPS analyses may be because of drift after early establishments. Macrofossil evidence from the Scandes Mountains of central Sweden suggests that Norway spruce already arrived in the early Holocene (Kullman, 1995; Kullman, 2002). As these early established populations probably expanded before a later colonization wave reached them (Giesecke and Bennett, 2004), their nuclear allele frequencies may have changed enough to maintain divergence from other stands. Founder events may also explain the high divergences of the peripheral populations, NN17 in northernmost Norway and that of SN36 and SN37 on the west coast of Norway, where spruce was established less than 1500 years ago (Fægri, 1950).
In contrast to the BAPS analysis, we did not find any meaningful nuclear genetic structure in the STRUCTURE analysis of Norway spruce. Both STRUCTURE and BAPS have been shown to perform poorly for data sets with FST values lower than 0.020 (Latch et al., 2006). In our study, in which the overall FST value was 0.029, lower and often insignificant genetic differentiation was found between many pairs of stands. In model-based clustering methods, such as STRUCTURE and BAPS, which define groups based on HW and linkage equilibrium, the presence of inbreeding or null alleles may further hamper the inference of the population structure (Falush et al., 2007). In the BAPS analysis, we constrained the individuals in each stand to belong to the same BAPS group, facilitating the detection of a structure.
Geographical trends of nuclear and mt diversity
Nuclear allelic richness decreased significantly with increasing distance from the Russian refugium, as expected from the general prediction of Hewitt (1999). Nuclear gene diversity did not, however, decrease with increasing distance from the refugium. This discrepancy can be explained by the fact that allelic richness is stronger affected by genetic drift than gene diversity as allelic richness is sensitive to the loss of rare alleles, whereas gene diversity depends more on the frequencies of the most common alleles (Nei et al., 1975). The clear loss of mtDNA diversity is, on the other hand, associated to the dispersal by seeds only and the lower effective population size (that is, higher genetic drift) of the haploid mt genome.
The high levels of nuclear diversity that we observed in southern Scandinavia can be explained by repeated dispersal events across the Baltic Sea. Furthermore, temporary increase in pollen flow during colonization from older, genetically more diverse populations to younger populations may also have counteracted the loss of nuclear diversity (Gamache et al., 2003; Heuertz et al., 2004b). In the presence of high levels of pollen flow, gene diversity is also expected to decrease less relative to allelic richness (Comps et al., 2001), in concordance with the pattern that we observed in southern Scandinavia. Pollen flow from non-autochthonous Norway spruce stands or even pollen flow from stands in the central European distribution range, may also have acted to increase the nuclear diversity level in southern parts of Scandinavia. Planting of non-autochthonous provenances is, however, common throughout the range (see Laikre et al., 2006 for Sweden) and could be expected to influence the diversity similarly over the range.
We found clear and significant decrease in both allelic richness and gene diversity towards the north as well as an increase in genetic differentiation based on the microsatellites. Such a pattern is often observed at species’ range margins where smaller effective population sizes and stronger geographical isolation make the populations more susceptible to the loss of genetic variation (Ouborg et al., 2006; Eckert et al., 2008). In the north, Norway spruce flowers infrequently (Schmidt-Vogt, 1977), and unfavourable conditions for seed ripening as well as an increased selection pressure are likely to reduce the effective population size. In Norway spruce, it has also been shown that pollen production decreases with latitude (Sarvas, 1957; Luomajoki, 1993). Reduced pollen availability towards the north may lead to increased inbreeding towards the north. The relative differences in FIS over all loci may thus reflect the differences in inbreeding. It should be stressed that the high correlation obtained in the Mantel test among pairwise FST and the ENA-corrected pairwise FST values indicates that all populations have been similarly affected by potential null alleles. Increased inbreeding towards the north was also evident in the outcrossing, wind-pollinated Picea sitchensis, in which population sizes and densities decreased towards the northern range margins (Mimura and Aitken, 2007).
In Norway spruce, mt diversity did not decrease towards the north, and particularly high mtDNA diversity was observed in the northern Finnish stands. Moreover, the increased differentiation towards Finland (as reflected in the genetic landscape shape analysis, Figure 3b) can be interpreted as a result of frequent long-distance dispersal events that supplement the migrating fronts with new variants maintaining diversity (Tollefsrud et al., 2008b). We, therefore, suggest that the loss of nuclear diversity towards the north rather reflects the northern climatic constraints of Norway spruce than the loss of diversity during postglacial colonization. An opposite trend was found in the north-American species Picea marina, in which postglacial colonization northwards induced a loss in mtDNA diversity whereas pollen flow maintained the nuclear microsatellite diversity towards the north. On the other hand, this species may be better adapted to cold climatic constraints as it is frequently found on tundra frost sites (Farjón, 1990).
A very weak pattern of IBD was found among stands in the northern range of Norway spruce. IBD will be at its maximum at an equilibrium between genetic drift and gene flow and will depend on the time since the populations were established and the distance from the ancestral population (Slatkin, 1993). Therefore, when the radiation time is short, only populations in close proximity to the ancestral population will show signs of IBD (Crispo and Hendry, 2005). The time to reach equilibrium between genetic drift and gene flow will also depend on the effective population size of the involved populations and the generation time of the organism (that is, longer time to equilibrium when the populations are large and when the generation time is long). As Norway spruce fulfilled its present northern range only 1500–2000 years ago (Fægri, 1950; Giesecke and Bennett, 2004; Latalowa and van der Knaap, 2006) and its average generation time is 40 years, spruce have thus experienced only 25–50 generations in its most recent regions, which, according to theory, can explain the lack of isolation by distance.
Mating within stands
The heterozygote deficiency that we detected in the Norway spruce stands seems to be in accordance with studies from other parts of its distribution range, both at isozyme loci (Lagercrantz and Ryman, 1990) and at different sets of microsatellite loci (Maghuly et al., 2006; Scotti et al., 2006; Meloni et al., 2007). Deviation from HW proportions was found within stands, frequently at loci largely unaffected by null alleles (Table 1, Appendix A). Inbreeding, in addition to null alleles, should thus be considered to explain the heterozygote deficiency. Norway spruce is a largely outcrossing species (Burczyk et al., 2004), and inbreeding in Norway spruce is thus most likely because of mating among relatives within stands. Although the proportion of self-pollination was estimated by Koski (1973) to average 10–20%, a strong selection was observed against seedlings resulting from selfing (Koski, 1973). Another factor explaining the deviation from HWE may be the very few effective outcross males contributing to pollination in the Norway spruce (Burczyk et al., 2004). Fertility components that are known to vary strongly from year to year for trees (Finkeldey and Ziehe, 2004) may possibly explain such an uneven contribution from outcrossing trees.
Conclusions
We found that the earlier inference of two migration routes out of the Russian refugium (Tollefsrud et al., 2008b) is consistent with the structuring at independent microsatellite loci. We found contrasting diversity patterns between mtDNA and nuclear DNA, suggesting that even if historical gene flow was high, present-day climatic constraints may reduce nuclear diversity in the north, primarily because of reduced pollen production. Considering a future scenario with a warmer and more humid climate in the north, Norway spruce may increase its propagation potential in the northernmost part of its range and recover from its present-day constraints.
References
Achereé V, Faivre-Rampant P, Jeandroz S, Besnard G, Markussen T, Aragones A et al. (2004). A full saturated linkage map of Picea abies including AFLP, SSR, ESTP, 5S rDNA and morphological markers. Theor Appl Genet 108: 1602–1613.
Ballard JWO, Whitlock MC (2004). The incomplete natural history of mitochondria. Mol Ecol 13: 729–744.
Björck S (1995). A review of the history of the Baltic Sea, 13.0–8.9 ka BP. Quat Int 27: 19–44.
Burczyk J, Lewandowski A, Chalupka W (2004). Local pollen dispersal and distant gene flow in Norway spruce (Picea abies L.) Karst.). For Ecol Manage 197: 39–48.
Chapuis MP, Estoup A (2007). Microsatellite null alleles and estimation of population differentiation. Mol Biol Evol 24: 621–631.
Comps B, Gomory D, Letouzey J, Thiebaut B, Petit RJ (2001). Diverging trends between heterozygosity and allelic richness during postglacial colonization in the European beech. Genetics 157: 389–397.
Corander J, Waldmann P, Marttinen P, Sillanpaa MJ (2004). BAPS 2: enhanced possibilities for the analysis of genetic population structure. Bioinformatics 20: 2363–2369.
Corander J, Waldmann P, Sillanpaa MJ (2003). Bayesian analysis of genetic differentiation between populations. Genetics 163: 367–374.
Cornuet JM, Luikart G (1996). Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144: 2001–2014.
Crispo E, Hendry AP (2005). Does time since colonization influence isolation by distance? A meta-analysis. Conserv Genet 6: 665–682.
Dempster AP, Laird NM, Rubin DB (1977). Maximum likelihood from incomplete data via Em-algorithm. J R Stat Soc Series B-Methodol 39: 1–38.
Dieringer D, Schlötterer C (2003). MICROSATELLITE ANALYSER (MSA): a platform independent analysis tool for large microsatellite data sets. Mol Ecol Notes 3: 167–169.
Eckert CG, Samis KE, Lougheed SC (2008). Genetic variation across species geographical ranges: the central-marginal hypothesis and beyond. Mol Ecol 17: 1170–1188.
Excoffier L, Laval G, Schneider S (2005). Arlequin ver. 3.0: An integrated software package for population genetics data analysis. Evol Bioinf Online 1: 47–50.
Falush D, Stephens M, Pritchard JK (2003). Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 164: 1567–1587.
Falush D, Stephens M, Pritchard JK (2007). Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7: 574–578.
Farjón A (1990). Pinaceae: drawings and descriptions of the genera Abies, Cedrus, Pseudolarix, Keteleeria, Nothotsuga, Tsuga, Cathaya, Pseudotsuga, Larix and Picea. Koeltz, Königstein.
Felsenstein J (2004). PHYLIP (Phylogeny Inference Package). Department of Genome Sciences, University of Washington: Seattle, WA, USA.
Finkeldey R, Ziehe M (2004). Genetic implications of silvicultural regimes. For Ecol Manage 197: 231–244.
Fægri K (1950). Omkring granens innvandring i Vest-Norge (The immigration of Norway spruce to West-Norway). Tidsskrift for skogbruk 58: 292–301.
Gamache I, Jaramillo-Correa JP, Payette S, Bousquet J (2003). Diverging patterns of mitochondrial and nuclear DNA diversity in subarctic black spruce: imprint of a founder effect associated with postglacial colonization. Mol Ecol 12: 891–901.
Giesecke T, Bennett KD (2004). The Holocene spread of Picea abies (L.) Karst. in Fennoscandia and adjacent areas. J Biogeogr 31: 1523–1548.
Goudet J (2001). FSTAT(version 2.9.3.): A program to estimate and test gene diversities and fixation indices. www.unil.ch/izea/softwares/fstat.html.
Guo SW, Thompson EA (1992). Performing the exact test of Hardy-Weinberg proportion for multiple alleles. Biometrics 48: 361–372.
Hedrick PW (2005). A standardized genetic differentiation measure. Evolution 59: 1633–1638.
Heuertz M, De Paoli E, Kallman T, Larsson H, Jurman I, Morgante M et al. (2006). Multilocus patterns of nucleotide diversity, linkage disequilibrium and demographic history of Norway spruce [Picea abies (L.) Karst]. Genetics 174: 2095–2105.
Heuertz M, Fineschi S, Anzidei M, Pastorelli R, Salvini D, Paule L et al. (2004a). Chloroplast DNA variation and postglacial recolonization of common ash (Fraxinus excelsior) in Europe. Mol Ecol 13: 3437–3452.
Heuertz M, Hausman JF, Hardy OJ, Vendramin GG, Frascaria-Lacoste N, Vekemans X (2004b). Nuclear microsatellites reveal contrasting patterns of genetic structure between western and southeastern European populations of the common ash (Fraxinus excelsior L.). Evolution 58: 976–988.
Hewitt GM (1999). Post-glacial re-colonization of European biota. Biol J Linn Soc 68: 87–112.
Hewitt GM (2001). Speciation, hybrid zones and phylogeography—or seeing genes in space and time. Mol Ecol 10: 537–549.
Hurlbert SH (1971). The nonconcept of species diversity: a critique and alternative parameters. Ecology 52: 577–586.
Jost L (2008). GST and its relatives do not measure differentiation. Mol Ecol 17: 4015–4026.
Koski V (1973). On self-pollination, genetic load, and subsequent inbreeding in some conifers. Commun Inst For Fenn 78: 1–40.
Kullman L (1995). New and firm evidence for midholocene appearance of Picea abies in the Scandes Mountains, Sweden. J Ecol 83: 439–447.
Kullman L (2002). Boreal tree taxa in the central Scandes during the Late-Glacial: implications for Late-Quaternary forest history. J Biogeogr 29: 1117–1124.
Kullman L (2008). Press release. http://www.info.umu.se/Nyheter/Pressmeddelande.aspx?id=3061.
Lagercrantz U, Ryman N (1990). Genetic structure of Norway spruce (Picea abies): Concordance of morphological and allozymic variation. Evolution 44: 38–53.
Laikre L, Palme A, Josefsson M, Utter F, Ryman N (2006). Release of alien populations in Sweden. Ambio 35: 255–261.
Latalowa M, van der Knaap WO (2006). Late Quaternary expansion of Norway spruce Picea abies (L.) Karst. in Europe according to pollen data. Qua Sci Rev 25: 2780–2805.
Latch EK, Dharmarajan G, Glaubitz JC, Rhodes OE (2006). Relative performance of Bayesian clustering software for inferring population substructure and individual assignment at low levels of population differentiation. Conserv Genet 7: 295–302.
Luikart G, Cornuet JM (1998). Empirical evaluation of a test for identifying recently bottlenecked populations from allele frequency data. Conserv Biol 12: 228–237.
Luomajoki A (1993). Climatic adaptation of Norway spruce (Picea abies (L.) Karsten) in Finland based on male flowering phenology. Acta For Fenn 242: 1–26.
Maghuly F, Pinsker W, Praznik W, Fluch S (2006). Genetic diversity in managed subpopulations of Norway spruce Picea abies (L.) Karst. For Ecol Manage 222: 266–271.
Magri D, Vendramin GG, Comps B, Dupanloup I, Geburek T, Gomory D et al. (2006). A new scenario for the Quaternary history of European beech populations: palaeobotanical evidence and genetic consequences. New Phytol 171: 199–221.
Meirmans PG (2006). Using the AMOVA framework to estimate a standardized genetic differentiation measure. Evolution 60: 2399–2402.
Meloni M, Perini D, Binelli G (2007). The distribution of genetic variation in Norway spruce (Picea abies Karst.) populations in the western Alps. J Biogeogr 34: 929–938.
Miller MP (2005). Alleles In Space (AIS): Computer software for the joint analysis of interindividual spatial and genetic information. J Hered 96: 722–724.
Miller MP, Bellinger MR, Forsman ED, Haig SM (2006). Effects of historical climate change, habitat connectivity, and vicariance on genetic structure and diversity across the range of the red tree vole (Phenacomys longicaudus) in the Pacific Northwestern United States. Mol Ecol 15: 145–159.
Mimura M, Aitken SN (2007). Increased selfing and decreased effective pollen donor number in peripheral relative to central populations in Picea sitchensis (Pinaceae). Am J Bot 94: 991–998.
Nei M (1987). Molecular Evolutionary Genetics. Columbia University Press: New York, USA.
Nei M, Maruyama T, Chakraborty R (1975). Bottleneck effect and genetic-variability in populations. Evolution 29: 1–10.
Nordborg M, Hu TT, Ishino Y, Jhaveri J, Toomajian C, Zheng HG et al. (2005). The pattern of polymorphism in Arabidopsis thaliana. Plos Biol 3: 1289–1299.
Ouborg NJ, Vergeer P, Mix C (2006). The rough edges of the conservation genetics paradigm for plants. J Ecol 94: 1233–1248.
Petit RJ, Duminil J, Fineschi S, Hampe A, Salvini D, Vendramin GG (2005). Comparative organization of chloroplast, mitochondrial and nuclear diversity in plant populations. Mol Ecol 14: 689–701.
Pfeiffer A, Olivieri AM, Morgante M (1997). Identification and characterization of microsatellites in Norway spruce (Picea abies K). Genome 40: 411–419.
Piry S, Luikart G, Cornuet JM (1999). BOTTLENECK: A computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90: 502–503.
Pons O, Petit RJ (1995). Estimation, variance and optimal sampling of gene diversity. I. Haploid locus. Theor Appl Genet 90: 462–470.
Pritchard JK, Stephens M, Donnelly P (2000). Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
R Development Core Team (2007). R: A Language and Environment for Statistical Computing. R foundation for Statistical Computing, Vienna, Austria.
Rice WR (1989). Analysing tables of statistical tests. Evolution 43: 223–225.
Rousset F (1997). Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145: 1219–1228.
Rousset F (2008). GENEPOP ′007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Res 8: 103–106.
Sarvas R (1957). Studies on the seed setting of Norway spruce. In: Mork E, Bonnevie-Svendsen C (eds). Studies on the seed setting of Norway Spruce. J.Chr. Gundersen boktrykkeri og bokbinderi: Oslo. pp 533–556.
Schmidt-Vogt H (1977). Die Fichte. Verlag Paul Parey: Hamburg and Berlin.
Scotti I, Magni F, Paglia GP, Morgante M (2002a). Trinucleotide microsatellites in Norway spruce (Picea abies): their features and the development of molecular markers. Theor Appl Genet 106: 40–50.
Scotti I, Paglia G, Magni F, Morgante M (2006). Population genetics of Norway spruce (Picea abies Karst.) at regional scale: sensitivity of different microsatellite motif classes in detecting differentiation. Ann For Sci 63: 485–491.
Scotti I, Paglia GP, Magni F, Morgante M (2002b). Efficient development of dinucleotide microsatellite markers in Norway spruce (Picea abies Karst.) through dot-blot selection. Theor Appl Genet 104: 1035–1041.
Semerikov VL, Lascoux M (2003). Nuclear and cytoplasmic variation within and between Eurasian Larix (Pinaceae) species. Am J Bot 90: 1113–1123.
Slatkin M (1993). Isolation by distance in equilibrium and nonequilibrium populations. Evolution 47: 264–279.
Sperisen C, Büchler U, Gugerli F, Matyas G, Geburek T, Vendramin GG (2001). Tandem repeats in plant mitochondrial genomes: application to the analysis of population differentiation in the conifer Norway spruce. Mol Ecol 10: 257–263.
Terhürne-Berson R (2005). Changing distribution patterns of selected conifers in the Quaternary of Europe caused by climatic variations. PhD Thesis, Rheinischen Friedrich-Wilhelms-Universität Bonn: Germany.
Tollefsrud MM, Brochmann C, Sperisen C (2008a). Paternal introgression from Siberian spruce (Picea obovata) to Norway spruce (P. abies): tracing pollen and seed flow with chloroplast and mitochondrial DNA. In: Tollefsrud MM's PhD Thesis: Phylogeography, diversity and hybridization in Norway spruce. University of Oslo, Norway.
Tollefsrud MM, Kissling R, Gugerli F, Johnsen Ø, Skrøppa T, Rachid C et al. (2008b). Genetic consequences of glacial survival and postglacial colonization in Norway spruce: combined analysis of mitochondrial DNA and fossil pollen. Mol Ecol 17: 4134–4150.
Vendramin GG, Anzidei M, Madaghiele A, Sperisen C, Bucci G (2000). Chloroplast microsatellite analysis reveals the presence of population subdivision in Norway spruce (Picea abies K.). Genome 43: 68–78.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Acknowledgements
We acknowledge our colleagues who have collected spruce samples: Darius Danusevicius, Bo Ekenäs, Bengt Eriksson, Hans Gustavsson, Gjert Heiberg, Igor Yakovlev, Risto Jalkanen, Christian Kierulf, Tiiu Koff, Merete Larsmon, Raine Larsson, Bo Leijon, Tore Løfgren, Teijo Nikkanen, Tommy Nordmark, Hans Nyeggen, Claes Olsson, Andrey Prokazin, Boris Raevsky, Jan Sotkajervi, Anders Uusitalo, Anders Vikström, Sigbjørn Øen, Geir Østereng, Åge Østgaard. Skogsstyrelsen in Sweden by Jonas Bergquist kindly organized sampling from many locations in Sweden and their collectors are acknowledged. Mari Mette Tollefsrud did the microsatellite analysis in the GenExpress laboratory, University of Florence, Italy, and we greatly acknowledge Federico Sebastiani and Anna Buonamici for their help. At the Norwegian Forest and Landscape Institute, Inger Heldal, Helene Haug Pagander, Veronica Yakovleva, Anne Nilsen and Merethe Fredhall are acknowledged for their help with DNA isolation. Two anonymous reviewers are acknowledged for their constructive comments on the manuscript. This research was supported by the Research Council of Norway Grant no. 143276/140 and Grant no. 155041140 and by the Norwegian Forest and Landscape Institute.
Author information
Authors and Affiliations
Corresponding author
Appendix A
Appendix A
Mitochondrial haplotype (nad1) data and nuclear microsatellite data for seven loci in Picea abies. Mitochondrial data are taken from Tollefsrud et al. (2008b), except for seven stands, for which data were obtained for this study (marked with *). Population abbreviations: RUS: Russia, BAL: Baltic States and Belarus, FIN: Finland, NN: Northern Norway, CS: Central Sweden, SS: Southern Sweden, SN: Southern Norway. For the mitochondrial data: NMT: sample size per stand, HS MT: mean genetic diversity, HT MT: total genetic diversity, AS MT: allelic richness. For the nuclear microsatellite data: NNUC: sample size per stand, HE NUC: gene diversity, AS NUC: allelic richness, FIS NUC: inbreeding coefficient estimated over all loci. * in the FIS NUC column indicates significant deviation from Hardy Weinberg equilibrium (HWE) after sequential Bonferroni correction. The loci which deviated from HW in the single locus exact test per stand are represented by A: EATC1D2A, B: EATC1B02, C: EATC2B02, D: EATC1E3, E: EATC2G05, F: SPAC1F7, G: EATC2C08; bold letters represent significant departure from HWE after sequential Bonferroni correction, grey letters represent significant departure from HWE after strict Bonferroni correction. The genetic (BAPS) group of the stand is given. Table A1
Rights and permissions
About this article
Cite this article
Tollefsrud, M., Sønstebø, J., Brochmann, C. et al. Combined analysis of nuclear and mitochondrial markers provide new insight into the genetic structure of North European Picea abies. Heredity 102, 549–562 (2009). https://doi.org/10.1038/hdy.2009.16
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2009.16
Keywords
This article is cited by
-
Continental-wide population genetics and post-Pleistocene range expansion in field maple (Acer campestre L.), a subdominant temperate broadleaved tree species
Tree Genetics & Genomes (2023)
-
Norway spruce postglacial recolonization of Fennoscandia
Nature Communications (2022)
-
A novel and diverse set of SNP markers for rangewide genetic studies in Picea abies
Conservation Genetics Resources (2022)
-
Genetic diversity of Norway spruce ecotypes assessed by GBS-derived SNPs
Scientific Reports (2021)
-
Towards the dynamic conservation of Serbian spruce (Picea omorika) western populations
Annals of Forest Science (2020)