Abstract
Background
Mutations in KRAS, BRAF, PIK3CA and PTEN expression have been in focus to predict the effect of epidermal growth factor receptor-blocking therapy in colorectal cancer (CRC). Here, information on these four aberrations was collected and combined to a Quadruple index and used to evaluate the prognostic role of these factors in CRC.
Patients
We analysed the mutation status in KRAS, BRAF and PIK3CA and PTEN expression in two separate CRC cohorts, Northern Sweden Health Disease Study (NSHDS; n=197) and Colorectal Cancer in Umeå Study (CRUMS; n=414). A Quadruple index was created, where Quadruple index positivity specifies cases with any aberration in KRAS, BRAF, PIK3CA or PTEN expression.
Results
Quadruple index positive tumours had a worse prognosis, significant in the NSHDS but not in the CRUMS cohort (NSHDS; P=0.003 and CRUMS; P=0.230) in univariate analyses but significance was lost in multivariate analyses. When analysing each gene separately, only BRAF was of prognostic significance in the NSHDS cohort (multivariate HR 2.00, 95% CI: 1.16–3.43) and KRAS was of prognostic significance in the CRUMS cohort (multivariate HR 1.48, 95% CI: 1.02–2.16). Aberrations in PIK3CA and PTEN did not add significant prognostic information.
Conclusions
Our results suggest that establishment of molecular subgroups based on KRAS and BRAF mutation status is important and should be considered in future prognostic studies in CRC.
Similar content being viewed by others
Main
Colorectal cancer (CRC) is one of the most common causes of cancer-related deaths in the western world (Jemal et al, 2008). Distant metastases represent the greatest threat to patient survival and about 40% of the patients will die from a metastatic disease. Surgical resection is today the basis for curative therapy, but a detailed understanding of the biological processes that regulate the establishment and progression of a malignant tumour may lead to improvements in non-surgical antitumour therapy. Two developmental pathways of sporadic CRC have been identified: chromosomal instability (or microsatellite stable, MSS) and microsatellite instability (MSI). Microsatellite stable tumours are considered to arise by copy number gains of oncogenes and loss of tumour suppressors, due to numerous chromosomal translocations (Grady, 2004). In contrast, MSI tumours show loss of expression of mismatch repair genes. They are less often associated with lymph node metastasis and distant spread, and MSI patients have a better prognosis than stage-matched MSS patients (Gryfe et al, 2000; Kohonen-Corish et al, 2005; Popat et al, 2005; Wright et al, 2005). Additionally, MSI tumours have been associated with CpG island methylator phenotype (CIMP) (Ahuja et al, 1997), where the groups CIMP-high, CIMP-low or CIMP-negative are based on promoter methylation frequency. We and others have reported a poorer prognosis for CRC patients with CIMP-high or CIMP-low tumours, compared with CIMP-negative tumours, especially in combination with MSS (Van Rijnsoever et al, 2003; Ward et al, 2003; Samowitz et al, 2005; Ogino et al, 2007; Shen et al, 2007; Barault et al, 2008; Dahlin et al, 2010).
Signalling through receptor tyrosine kinases in response to cytokines, growth factors and hormones is important for maintaining the metabolism, proliferation, survival and motility of a cell (Haglund et al, 2007). Many of these signals involve the oncogenic proteins KRAS, BRAF, PIK3CA and the tumour suppressor PTEN which are all downstream effectors of the epidermal growth factor receptor (EGFR) (Siena et al, 2009). Treatment targeting EGFR has been found to be efficient only if no mutations are found in KRAS or BRAF (Lievre et al, 2006). Still all patients with wild-type KRAS and BRAF do not respond to treatment (Amado et al, 2008; Bardelli and Siena, 2010; Tol et al, 2010). PIK3CA and PTEN have been suggested to harbour aberrations in 30–40% of all sporadic CRC cases (Samuels and Ericson, 2006; Frattini et al, 2007), which might explain part of this resistance. A recent study suggested that mutations in PI3K catalytic subunit (PIK3CA) may carry prognostic information in tumour stage I–III (Ogino et al, 2009), and that PIK3CA/PTEN deregulation, in addition to KRAS and BRAF mutations, may be a biomarker of resistance (Perrone et al, 2009; Sartore-Bianchi et al, 2009). Consequently, Sartore-Bianchi et al (2009) introduced the Quadruple index as a factor taking aberrations in these four factors into simultaneous consideration. Even though many studies are focusing on the molecules downstream EGFR to estimate benefit from EGFR blocking therapy, it is still not known how the mutations affect patient prognosis and tumour aggressiveness per se.
Therefore, we have in the present study analysed the mutational status of KRAS, BRAF, PIK3CA and PTEN expression separately, and combined as Quadruple index, and correlated the results to patient survival. Additionally, we related mutation status to established molecular tumour characteristics such as MSI screening status and CIMP status.
Material and methods
Patient selection
Colorectal cancer cases from two separate patient groups were included in the present study. Archival paraffin-embedded CRC tissue samples from a total 414 patients were included from the Colorectal Cancer in Umeå Study (CRUMS), all collected during primary tumour surgery over the period 1995–2003 at Umeå University Hospital, Sweden. All routinely stained sections were reviewed by one observer, who performed all histopathological classifications including stage and tumour type (mucinous or non-mucinous). Tissue blocks from the primary tumour were chosen for DNA extraction. When necessary the proportion of tumour cells was maximised by macrodissection and necrotic areas were avoided. Clinical data were obtained by reviewing the patient records and survival data were collected from the Swedish population registry during autumn 2012 with a median follow-up time of 113 months for patients still alive at the end of follow-up.
From the Northern Sweden Health Disease Study (NSHDS), archival paraffin-embedded CRC tissue from a total of 197 patients was included. The NSHDS cohort consists of three separate cohorts: the Västerbotten Intervention Project (VIP), the Northern Sweden WHO Monitoring of Trends and Cardiovascular Disease Study (MONICA) and the local Mammography Screening Project (MSP) (Hallmans et al, 2003). The CRC cases in the NSHDS cohort, protocols and selection principles used in the present study have previously been described in detail (Van Guelpen et al, 2006). Brief summary of subjects included in the NSHDS cohort: consists of both men and women in the age of 40, 50 and 60 years in VIP; both men and women ages 25–74 years in MONICA; and only women ages ∼50–70 years in MSP. Within these cohorts, a total of 226 CRC cases were identified and selected for a previous nested case-referent study (Van Guelpen et al, 2006). After exclusion of insufficient or unavailable tumour tissue samples, 197 patients were available for mutation analysis in the NSHDS cohort.
NSHDS patients were followed up until January 2008 with a median follow-up time of 102 months for patients still alive at the end of follow-up. Cancer-specific survival was collected from the Swedish population registry and patient records. Patients originally included in both cohorts were excluded from the CRUMS cohort and only reported once.
The handling of tissue samples and patient data in this study has been approved by the local ethics committee of Umeå University, Umeå, Sweden.
Mutational analysis of KRAS and PIK3CA exon 20
PCR conditions for KRAS: 50 ng DNA, 0.5 μg primer, 10 mM dNTP, 1 mM MgCl2 and 0.4U JumpStart Taq (Sigma, Stockholm, Sweden) in a total volume of 20 μl. PCR were run at 95 °C 10 min, 95 °C 15 s, 65–55 °C (−1 °C/cycle) 72 °C 30 s (touchdown for 10 cycles); 95 °C 15 s, 55 °C 15 s, 72 °C 30 s for 35 cycles and 72 °C 10 min. Primers used:
forward: 5′-tgtaaaacgacggccagtgagtttgtattaaaaggtactgg-3′.
reverse: 5′-caggaaacagctatgacctctgtatcaaagaatggtcct-3′.
PCR conditions for PIK3CA exon 20: 50 ng DNA, 0.5 μg primer, 10 mM dNTP, 3 mM MgCl2 and 0.4U JumpStart Taq (Sigma, Stockholm, Sweden) in a total volume of 20 μl. PCR were run at 95 °C 10 min, 95 °C 21 s, 59 °C 21 s, 72 °C 30 s for 40 cycles and 72 °C 10 min. Primers used:
forward: 5′-tgtaaaacgacggccagtctcaatgatgcttggctctg-3′.
reverse: 5′-caggaaacagctatgaccatgctgttcatggattgtgc-3′.
All primers were M13-tagged (forward: 5′-tgtaaaacgacggccagt-3′; reverse: 5′-caggaaacagctatg-3′) to receive a more specific PCR product during the sequencing reaction. Sequencing was performed using Big Dye v. 3.1 according to the manufacture protocol, analysed in a 3730 xl DNA Analyser (Applied Biosystems, Stockholm, Sweden). The results were evaluated in SeqScape v2 1.1 (Applied Biosystem).
BRAF V600E mutational analysis
Detection of BRAF V600E mutation was done with the Taqman allelic discrimination assay (reagents from Applied Biosystems), which has been described in detail elsewhere (Benlloch et al, 2006).
Immunohistochemical analysis of PTEN expression
Specimens were fixed in 4% formaldehyde and embedded in paraffin, according to routine procedures at the Department of Clinical Pathology, Umeå University Hospital, Sweden. Four micrometre sections were deparaffinized and rehydrated. Antigen retrieval treatment was executed using Borg solution (Biocare Medical, Concord, CA, USA) in a pressure cooker (2100 retriever, Biocare Medical). Primary monoclonal mouse PTEN antibody (Dako, Stockholm, Sweden, clone 6H 2.1, diluted 1 : 50) was used in a semiautomatic staining machine (intelliPATH FLX, Biocare Medical).
The samples were evaluated for cytoplasmic staining, and were graded 0as no staining, 1as weak staining, and 2as moderate-strong staining. Loss of PTEN expression (graded as 0) was considered as abnormal while grade 1 and 2 was considered normal. Nerve tissue and blood vessels were used as positive internal controls in each sample. Cases without internal positive control staining were considered uninformative.
A Quadruple index was created according to Sartore-Bianchi et al (2009), where negative specify cases where all selected genes (KRAS, BRAF and PIK3CA) were wild-type and normal expression of PTEN was seen. Quadruple index positivity indicates cases where at least one of the KRAS, BRAF or PIK3CA genes was mutated and/or loss of PTEN expression was found.
Microsatellite instability screening status and CIMP status
Immunohistochemical analyses of mismatch repair proteins were performed as previously described (Dahlin et al, 2010). Briefly, expression of four mismatch repair proteins, MLH1, MSH2, MSH6 and PMS2 were analysed in formalin-fixed and paraffin-embedded human CRC tissue. Tissue samples lacking nuclear staining in tumour cells for at least one of these proteins were considered to have a positive MSI screening status, referred to as MSI. Negative MSI screening status based on immunohistochemical staining is referred to as MSS.
Methylation analysis to determine tumour CIMP status was performed by the MethyLight method, with primer and probe sequences as previously described (Weisenberger et al, 2006; Dahlin et al, 2010). The per cent of methylated refence (PMR) value was caluculated for the eight genes included in the CIMP panel (CDKN2A, MLH1, CACNA1G, NEUROG1, RUNX3, SOCS1, IGF2 and CRABP1) (Dahlin et al, 2010), and a gene was considered positive for methylation when the PMR>10 (Weisenberger et al, 2006).
Tumours with no promoter hypermethylation were classified as CIMP-negative, 1–5 genes methylated as CIMP-low, and 6–8 genes as CIMP-high (Dahlin et al, 2010).
Statistical analysis
Clinico-pathological characteristics were compared using Kruskal–Wallis tests for continuous variables and χ2-tests, or Fisher’s exact tests when observed or expected frequencies were less than five for categorical variables. For cancer-specific survival analyses, Kaplan–Meier plots were used, and differences between groups were tested by log-rank tests. Cancer-specific events were defined as death with known disseminated or recurrent disease, and cases were censored at the end of follow-up or at time of death by other causes.
Patients in CRUMS who were deceased with postoperative complications within 1 month after surgery (n=16) were excluded from the survival analyses. Deaths due to postoperative complications were not recorded in NSHDS, but only four patients died within 1 month of surgery. To take into consideration other clinico-pathological factors, multivariate Cox proportional hazard models were used. For multivariate analyses, we analysed Quadruple index, KRAS and BRAF and not PIK3CA and PTEN, as the latter two were not significantly associated with prognosis in univariate analyses. The adjusting variables were selected if they affected the risk estimates for KRAS and BRAF >10% in bivariate analyses. The final multivariate model included sex, age at diagnosis, stage and tumour site. Other factors tested, but not meeting the criteria for inclusion in the multivariate analyses were aberrant p53 protein expression, mucinous histologic tumour type, preoperative radiotherapy and adjuvant chemotherapy. Microsatellite instability screening status and CIMP status were also tested but excluded due to small subgroups and thereby loss of statistical power. All statistical tests were conducted using PASW Statistics 18 (SPSS Inc., Chicago, IL, USA).
Results
Quadruple index in relation to clinico-pathological variables
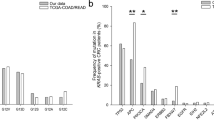
We analysed each mutation (KRAS, BRAF and PIK3CA) and PTEN expression as well as the Quadruple Index, in tumours from 197 patients in the NSHDS and 414 patients in the CRUMS cohort with respect to different clinico-pathological characteristics (Tables 1A and 1B). Seven different activating mutations in codon 12 and 13 were analysed in KRAS, and the mutation frequency was 17.9% in the NSHDS and 19.5% in the CRUMS cohort. BRAF was observed in 17.9 and 13.2% in each study population respectively (Tables 1A and 1B). When combining results from the four studied factors, only two patients had both BRAF and KRAS mutated in the NSHDS cohort (Figure 1A), while BRAF and KRAS mutations were mutually exclusive (Figure 1B) in the CRUMS cohort. Four different mutations were analysed in PIK3CA, exon 20, where the mutation frequency was 2.2% in both cohorts. Loss of PTEN expression was found in 12.5% in the NSHDS and 14.1% in the CRUMS cohort (Tables 1A and 1B). In the NSHDS cohort mutated KRAS and BRAF tumours were associated with right colon location, most distinct for BRAF (NSHDS; P<0.001). In the CRUMS cohort, BRAF mutant tumours were significantly correlated to older age (CRUMS; P=0.017) and right colon location (CRUMS; P<0.001), while KRAS mutations were significantly associated with higher tumour stage (CRUMS; P=0.030). BRAF mutations were most prevalent in mucinous tumours (Tables 1A and 1B).
The interrelationship between cases with mutations in KRAS, BRAF, PIK3CA and loss of PTEN expression in the NSHDS and the CRUMS cohorts. Total number of aberrations in NSHDS (A); KRAS (N=30), BRAF (N=31), PIK3CA (N=3), PTEN (N=18); CRUMS (B); KRAS (N=77), BRAF (N=50), PIK3CA (N=8), PTEN (N=57). Patients with missing value in any of the marker were excluded from the Figure.
The frequencies of Quadruple index positivity were 48.3% in the NSHDS and 44.0% in the CRUMS cohort. Quadruple index positivity was correlated significantly to right colon location in both patient groups (NSHDS and CRUMS; both P<0.001). Quadruple index positivity, BRAF mutations and loss of PTEN expression were significantly associated with higher tumour stage in the NSHDS, but not in the CRUMS cohort (Tables 1A and 1B).
Quadruple index in relation to MSI screening status and CIMP status
Tables 2A and 2B shows Quadruple index and each mutation (KRAS, BRAF and PIK3CA) and PTEN expression in relation to both MSI screening status and CIMP status in the NSHDS and the CRUMS cohort. Quadruple index positivity correlated significantly to CIMP-high status (NSHDS; P=0.002 and CRUMS; P<0.001) in both the NSHDS and the CRUMS cohort, and to MSI (CRUMS; P<0.001) in the CRUMS cohort. KRAS mutations were more often seen in patients with MSS (NSHDS; P=0.031 and CRUMS; P=0.002) and CIMP-low tumours (NSHDS; P=0.046 and CRUMS; P=0.001). BRAF mutations were significantly associated with MSI (NSHDS; P<0.001 and CRUMS; P<0.001) and CIMP-high (NSHDS; P<0.001 and CRUMS; P<0.001). Mutations in the PIK3CA gene significantly correlated to MSI (CRUMS; P=0.013) and CIMP-high (CRUMS; P=0.006) in the CRUMS cohort, but showed no statistical significance in the NSHDS cohort. Loss of PTEN expression did not show significant correlation to MSI screening status or CIMP status in any of the cohorts.
Survival analysis
Cancer-specific survival analyses revealed that Quadruple index positive cases had a significantly worse prognosis compared with negative cases in the NSHDS cohort (Figure 2A; univariate HR 1.98, 95% CI: 1.25–3.13). However, the Quadruple index positive cases had only a slightly poorer, but not statistically significant, prognosis in the CRUMS cohort (Figure 2B; univariate HR 1.22, 95% CI: 0.88–1.69).
When analysing each gene separately only BRAF mutations turned out to be of prognostic value in the NSHDS cohort (Figure 2E), a result that retained statistical significant also in a multivariate Cox proportional hazard model (Table 3A).
In the CRUMS cohort, on the other hand, only KRAS mutations were of prognostic value (Figure 2D), and this was seen also in multivariate analyses (Table 3B). Neither PIK3CA mutations, nor loss of PTEN expression were of prognostic significance in any of the two cohorts when analysed separately (Figure 2G–J).
Survival analyses stratified for MSI screening status and CIMP status
Patients with Quadruple index positive tumours with MSS (NSHDS; P=0.002), or CIMP-low (NSHDS; P=0.022) or CIMP-high tumours (CRUMS; P=0.042) had a worse prognosis than Quadruple index negative cases. Cancer-specific survival analyses stratified for KRAS and BRAF is shown in Figure 3. Patients with tumours harbouring BRAF mutations together with MSS (NSHDS; P=<0.001) (Figure 3G) or CIMP-low (NSHDS; P<0.001) (Figure 3O) showed an impaired survival in the NSHDS cohort. In the CRUMS cohort, tumours with KRAS mutations accompanied with MSS (Figure 3F) (CRUMS; P=0.042) or CIMP-negative (CRUMS; P=0.010) or BRAF mutations in CIMP-high tumours (CRUMS; P=0.001) (Figure 3T) showed a poorer patient prognosis. Owing to the loss of statistical power in these small subgroups, a multivariate model was not performed.
Discussion
In this study archival CRC tissue from two different cohorts from Northern Sweden, NSHDS and CRUMS, were analysed regarding mutations in the genes KRAS, BRAF, PIK3CA and loss of PTEN expression. All four aberrations investigated in this study are part of the same signalling pathway, downstream the EGFR, and to get an increased understanding for how these factors are interconnected in CRC, a Quadruple index as suggested by Sartore-Bianchi et al (2009) was created, where Quadruple index positive tumours had at least one mutation in any of the genes KRAS, BRAF, PIK3CA and/or loss of PTEN protein expression.
We found a shorter cancer-specific survival in patients with Quadruple index positive tumours in the NSHDS cohort, but the Quadruple index was not statistically significant in the CRUMS cohort. Analysing each gene separately revealed that only mutations in the BRAF gene had a significant prognostic value in the NSHDS cohort, especially in combination with MSS or CIMP-low. Only KRAS mutations, on the other hand, indicated a significantly poorer patient prognosis in the CRUMS cohort, especially together with MSS or CIMP-negative tumours. Aberrations in PIK3CA and PTEN did not add significant prognostic information. Therefore, our results do not support the use of the full Quadruple index but instead emphasise the prognostic information in KRAS and BRAF mutation status.
Taken together, these results indicate that the establishment of molecular subgroups of CRC based on KRAS and BRAF mutation status can supply important information, not only in prediction of the EGFR-treatment response but also in prediction of patient prognosis. Importantly, KRAS and BRAF mutations are nearly mutually exclusive in CRC (Jakubauskas and Griskevicius, 2010; Li et al, 2011; Krol et al, 2012).
The finding of contrary significances for KRAS and BRAF mutations in the two cohorts is not easily explained. However, it should be noted that the composition and the underlying design of the two cohorts differs significantly. For example, NSHDS consists of more women than men as a direct result of including the Mammary Screening Project as one of the three subcohorts, and BRAF mutations have more often been reported in women (Ogino et al, 2012). Furthermore, the age distribution also differs between the two cohorts and might have impact on the results. Not only the KRAS and BRAF mutations, but also molecular characteristics such as MSI screening status and CIMP status, are well known to correlate with the age and sex distribution (Nosho et al, 2009; Kalady et al, 2012). The contradictory results, however, emphasise a need for further larger studies on this topic.
One of the main strengths of this study was the two large, non-overlapping, patient groups, which were both from the same northern Swedish population but had different recruitment protocols, age range and sex distributions. The patients in the present study were generally diagnosed previous to the broad introduction of many novel therapies, including successful resection of liver metastases, into clinical practice. Treatment was thus fairly homogeneous within each tumour site and stage. Residual confounding effect due to differences in treatment is therefore unlikely. It is not possible, however, to analyse the predictive value of mutations with respect to EGFR-blocking therapy in our patient cohorts due to the lack of such treatment during the cohort recruitment. Instead, the two cohorts include all tumour stages and are suitable for studies on tumour aggressiveness and prognosis.
The present study is, to the best of our knowledge, the largest study today on this subject. Despite the use of two patient cohorts, a limitation is, however, still the relatively low number of patients, especially when analysing somewhat rare subgroups (e.g., PIK3CA mutations, MSI cases or CIMP-high cases). The fact that we could not detect any correlation between loss of PTEN expression or PIK3CA mutations and patient prognosis makes us speculate that the need for analysing all four genes, as in the Quadruple index, might be unnecessary when prognosticating cancer-specific survival. There are, however, contradictory reports indicating that both PIK3CA mutations and loss of PTEN protein expression do affect patient prognosis (Sawai et al, 2008; Li et al, 2009; Jang et al, 2010; Liao et al, 2012).
The mutation frequencies of each analysed gene found in this study were in general similar to previous reports (Rako et al, 2012; Soeda et al, 2012), except for the KRAS gene. We report a frequency of about 20%, while several other reports have reported frequencies of 30–40% (Kim et al, 2012). The low mutation frequency of KRAS in our studied populations can have several explanations. Our patient cohorts have a rather high proportion of rectal cancers, and rectal cancers have a lower KRAS mutation frequency than colon cancers. Technical differences between studies are another likely explanation, and here we have not analysed KRAS mutations in exon 61. Furthermore, most studies reporting the frequency of KRAS mutations have studied only metastatic CRCs, and KRAS-mutated CRC might be more aggressive than their wild-type counterparts.
Previous reports on PIK3CA mutation frequencies in CRC have varied considerably. In this study we report a frequency of about 2%. However, we have only analysed mutations in exon 20 in PIK3CA, not exon 9, based on recently published data showing that only mutations in exon 20 have a prognostic value (De Roock et al, 2010; Farina Sarasqueta et al, 2011), probably as this exon translates the kinase domain of PIK3CA. Additionally Muller et al (2007), recently found a PIK3CA pseudogene spanning exons 9–13 located on chromosome 22, which might be the reason for such a high reported frequency of PIK3CA exon 9 mutations.
In conclusion, by the use of two patient cohorts we show that mutations in the KRAS and BRAF genes are of prognostic importance in colorectal cancer. However, adding information on mutation status of PIK3CA and loss of PTEN does not add significant prognostic information. These results suggest that establishment of molecular subgroups based on KRAS and BRAF mutation status is important and should be considered in future prognostic studies in CRC.
Change history
28 May 2013
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Ahuja N, Mohan AL, Li Q, Stolker JM, Herman JG, Hamilton SR, Baylin SB, Issa JP (1997) Association between CpG island methylation and microsatellite instability in colorectal cancer. Cancer Res 57: 3370–3374
Amado RG, Wolf M, Peeters M, Van Cutsem E, Siena S, Freeman DJ, Juan T, Sikorski R, Suggs S, Radinsky R, Patterson SD, Chang DD (2008) Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J Clin Oncol 26: 1626–1634
Barault L, Charon-Barra C, Jooste V, de la Vega MF, Martin L, Roignot P, Rat P, Bouvier AM, Laurent-Puig P, Faivre J, Chapusot C, Piard F (2008) Hypermethylator phenotype in sporadic colon cancer: study on a population-based series of 582 cases. Cancer Res 68: 8541–8546
Bardelli A, Siena S (2010) Molecular mechanisms of resistance to cetuximab and panitumumab in colorectal cancer. J Clin Oncol 28: 1254–1261
Benlloch S, Paya A, Alenda C, Bessa X, Andreu M, Jover R, Castells A, Llor X, Aranda FI, Massuti B (2006) Detection of BRAF V600E mutation in colorectal cancer: comparison of automatic sequencing and real-time chemistry methodology. J Mol Diagn 8: 540–543
Dahlin AM, Palmqvist R, Henriksson ML, Jacobsson M, Eklof V, Rutegard J, Oberg A, Van Guelpen BR (2010) The role of the CpG island methylator phenotype in colorectal cancer prognosis depends on microsatellite instability screening status. Clin Cancer Res 16: 1845–1855
De Roock W, Claes B, Bernasconi D, De Schutter J, Biesmans B, Fountzilas G, Kalogeras KT, Kotoula V, Papamichael D, Laurent-Puig P, Penault-Llorca F, Rougier P, Vincenzi B, Santini D, Tonini G, Cappuzzo F, Frattini M, Molinari F, Saletti P, De Dosso S, Martini M, Bardelli A, Siena S, Sartore-Bianchi A, Tabernero J, Macarulla T, Di Fiore F, Gangloff AO, Ciardiello F, Pfeiffer P, Qvortrup C, Hansen TP, Van Cutsem E, Piessevaux H, Lambrechts D, Delorenzi M, Tejpar S (2010) Effects of KRAS, BRAF, NRAS, and PIK3CA mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: a retrospective consortium analysis. Lancet Oncol 11: 753–762
Farina Sarasqueta A, Zeestraten EC, van Wezel T, van Lijnschoten G, van Eijk R, Dekker JW, Kuppen PJ, Goossens-Beumer IJ, Lemmens VE, van de Velde CJ, Rutten HJ, Morreau H, van den Brule AJ (2011) PIK3CA kinase domain mutation identifies a subgroup of stage III colon cancer patients with poor prognosis. Cell Oncol 34: 523–531
Frattini M, Saletti P, Romagnani E, Martin V, Molinari F, Ghisletta M, Camponovo A, Etienne LL, Cavalli F, Mazzucchelli L (2007) PTEN loss of expression predicts cetuximab efficacy in metastatic colorectal cancer patients. Br J Cancer 97: 1139–1145
Grady WM (2004) Genomic instability and colon cancer. Cancer Metastasis Rev 23: 11–27
Gryfe R, Kim H, Hsieh ET, Aronson MD, Holowaty EJ, Bull SB, Redston M, Gallinger S (2000) Tumour microsatellite instability and clinical outcome in young patients with colorectal cancer. N Engl J Med 342: 69–77
Haglund K, Rusten TE, Stenmark H (2007) Aberrant receptor signaling and trafficking as mechanisms in oncogenesis. Crit Rev Oncog 13: 39–74
Hallmans G, Agren A, Johansson G, Stegmayr B, Jansson JH, Lindahl B, Rolandsson O, Södeberg S, Nilsson M, Johansson I, Weinehall L (2003) Cardiovascular disease and diabetes in the Northern Sweden Health and Disease Study Cohort—evaluation of risk factors and their interactions. Scand J Public Health Suppl 61: 18–24
Jakubauskas A, Griskevicius L (2010) KRas and BRaf mutational status analysis from formalin-fixed, paraffin-embedded tissues using multiplex polymerase chain reaction-based assay. Arch Pathol Lab Med 134: 620–624
Jang KS, Song YS, Jang SH, Min KW, Na W, Jang SM, Jun YJ, Lee KH, Choi D, Paik SS (2010) Clinicopathological significance of nuclear PTEN expression in colorectal adenocarcinoma. Histopathology 56: 229–239
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Murray T, Thun MJ (2008) Cancer statistics, 2008. CA Cancer J Clin 58: 71–96
Kalady MF, Dejulius KL, Sanchez JA, Jarrar A, Liu X, Manilich E, Skacel M, Church JM (2012) BRAF mutations in colorectal cancer are associated with distinct clinical characteristics and worse prognosis. Dis Colon Rectum 55: 128–133
Kim SY, Shim EK, Yeo HY, Baek JY, Hong YS, Kim DY, Kim TW, Kim JH, Im SA, Jung KH, Chang HJ (2012) KRAS Mutation Status and Clinical Outcome of Preoperative Chemoradiation With Cetuximab in Locally Advanced Rectal Cancer: A Pooled Analysis of 2 Phase II Trials. Int J Radiat Oncol Biol Phys 85: 201–207
Kohonen-Corish MR, Daniel JJ, Chan C, Lin BP, Kwun SY, Dent OF, Dhillon VS, Trent RJ, Chapuis PH, Bokey EL (2005) Low microsatellite instability is associated with poor prognosis in stage C colon cancer. J Clin Oncol 23: 2318–2324
Krol LC, t Hart NA, Methorst N, Knol AJ, Prinsen C, Boers JE (2012) Concordance in KRAS and BRAF mutations in endoscopic biopsy samples and resection specimens of colorectal adenocarcinoma. Eur J Cancer 48: 1108–1115
Li HT, Lu YY, An YX, Wang X, Zhao QC (2011) KRAS, BRAF and PIK3CA mutations in human colorectal cancer: relationship with metastatic colorectal cancer. Oncol Rep 25: 1691–1697
Li XH, Zheng HC, Takahashi H, Masuda S, Yang XH, Takano Y (2009) PTEN expression and mutation in colorectal carcinomas. Oncol Rep 22: 757–764
Liao X, Morikawa T, Lochhead P, Imamura Y, Kuchiba A, Yamauchi M, Nosho K, Qian ZR, Nishihara R, Meyerhardt JA, Fuchs CS, Ogino S (2012) Prognostic role of PIK3CA mutation in colorectal cancer: cohort study and literature review. Clin Cancer Res 18: 2257–2268
Lievre A, Bachet JB, Le Corre D, Boige V, Landi B, Emile JF, Cote JF, Tomasic G, Penna C, Ducreux M, Rougier P, Penault-Llorca F, Laurent-Puig P (2006) KRAS mutation status is predictive of response to cetuximab therapy in colorectal cancer. Cancer Res 66: 3992–3995
Muller CI, Miller CW, Hofmann WK, Gross ME, Walsh CS, Kawamata N, Luong QT, Koeffler HP (2007) Rare mutations of the PIK3CA gene in malignancies of the hematopoietic system as well as endometrium, ovary, prostate and osteosarcomas, and discovery of a PIK3CA pseudogene. Leuk Res 31: 27–32
Nosho K, Kure S, Irahara N, Shima K, Baba Y, Spiegelman D, Meyerhardt JA, Giovannucci EL, Fuchs CS, Ogino S (2009) A prospective cohort study shows unique epigenetic, genetic, and prognostic features of synchronous colorectal cancers. Gastroenterology 137: 1609–1620, e1-3
Ogino S, Kawasaki T, Kirkner GJ, Kraft P, Loda M, Fuchs CS (2007) Evaluation of markers for CpG island methylator phenotype (CIMP) in colorectal cancer by a large population-based sample. J Mol Diagn 9: 305–314
Ogino S, Nosho K, Kirkner GJ, Shima K, Irahara N, Kure S, Chan AT, Engelman JA, Kraft P, Cantley LC, Giovannucci EL, Fuchs CS (2009) PIK3CA mutation is associated with poor prognosis among patients with curatively resected colon cancer. J Clin Oncol 27: 1477–1484
Ogino S, Shima K, Meyerhardt JA, McCleary NJ, Ng K, Hollis D, Saltz LB, Mayer RJ, Schaefer P, Whittom R, Hantel A, Benson AB 3rd, Spiegelman D, Goldberg RM, Bertagnolli MM, Fuchs CS (2012) Predictive and prognostic roles of BRAF mutation in stage III colon cancer: results from intergroup trial CALGB 89803. Clin Cancer Res 18: 890–900
Perrone F, Lampis A, Orsenigo M, Di Bartolomeo M, Gevorgyan A, Losa M, Frattini M, Riva C, Andreola S, Bajetta E, Bertario L, Leo E, Pierotti MA, Pilotti S (2009) PI3KCA/PTEN deregulation contributes to impaired responses to cetuximab in metastatic colorectal cancer patients. Ann Oncol 20: 84–90
Popat S, Hubner R, Houlston RS (2005) Systematic review of microsatellite instability and colorectal cancer prognosis. J Clin Oncol 23: 609–618
Rako I, Jakic-Razumovic J, Katalinic D, Sertic J, Plestina S (2012) Mutation pattern of KRAS and BRAF oncogenes in colorectal cancer patients. Neoplasma 59: 376–383
Samowitz WS, Albertsen H, Herrick J, Levin TR, Sweeney C, Murtaugh MA, Wolff RK, Slattery ML (2005) Evaluation of a large, population-based sample supports a CpG island methylator phenotype in colon cancer. Gastroenterology 129: 837–845
Samuels Y, Ericson K (2006) Oncogenic PI3K and its role in cancer. Curr Opin Oncol 18: 77–82
Sartore-Bianchi A, Di Nicolantonio F, Nichelatti M, Molinari F, De Dosso S, Saletti P, Martini M, Cipani T, Marrapese G, Mazzucchelli L, Lamba S, Veronese S, Frattini M, Bardelli A, Siena S (2009) Multi-determinants analysis of molecular alterations for predicting clinical benefit to EGFR-targeted monoclonal antibodies in colorectal cancer. PLoS One 4: e7287
Sawai H, Yasuda A, Ochi N, Ma J, Matsuo Y, Wakasugi T, Takahashi H, Funahashi H, Sato M, Takeyama H (2008) Loss of PTEN expression is associated with colorectal cancer liver metastasis and poor patient survival. BMC Gastroenterol 8: 56
Shen L, Catalano PJ, Benson AB 3rd, O'Dwyer P, Hamilton SR, Issa JP (2007) Association between DNA methylation and shortened survival in patients with advanced colorectal cancer treated with 5-fluorouracil based chemotherapy. Clin Cancer Res 13: 6093–6098
Siena S, Sartore-Bianchi A, Di Nicolantonio F, Balfour J, Bardelli A (2009) Biomarkers predicting clinical outcome of epidermal growth factor receptor-targeted therapy in metastatic colorectal cancer. J Natl Cancer Inst 101: 1308–1324
Soeda H, Shimodaira H, Watanabe M, Suzuki T, Gamoh M, Mori T, Komine K, Iwama N, Kato S, Ishioka C (2012) Clinical usefulness of KRAS, BRAF, and PIK3CA mutations as predictive markers of cetuximab efficacy in irinotecan- and oxaliplatin-refractory Japanese patients with metastatic colorectal cancer. Int J Clin Oncol e-pub ahead of print, 26 May 2012
Tol J, Dijkstra JR, Klomp M, Teerenstra S, Dommerholt M, Vink-Borger ME, van Cleef PH, van Krieken JH, Punt CJ, Nagtegaal ID (2010) Markers for EGFR pathway activation as predictor of outcome in metastatic colorectal cancer patients treated with or without cetuximab. Eur J Cancer 46: 1997–2009
Van Guelpen B, Hultdin J, Johansson I, Hallmans G, Stenling R, Riboli E, Winkvist A, Palmqvist R (2006) Low folate levels may protect against colorectal cancer. Gut 55: 1461–1466
Van Rijnsoever M, Elsaleh H, Joseph D, McCaul K, Iacopetta B (2003) CpG island methylator phenotype is an independent predictor of survival benefit from 5-fluorouracil in stage III colorectal cancer. Clin Cancer Res 9: 2898–2903
Ward RL, Cheong K, Ku SL, Meagher A, O’Connor T, Hawkins NJ (2003) Adverse prognostic effect of methylation in colorectal cancer is reversed by microsatellite instability. J Clin Oncol 21: 3729–3736
Weisenberger DJ, Siegmund KD, Campan M, Young J, Long TI, Faasse MA, Kang GH, Widschwendter M, Weener D, Buchanan D, Koh H, Simms L, Barker M, Leggett B, Levine J, Kim M, French AJ, Thibodeau SN, Jass J, Haile R, Laird PW (2006) CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet 38: 787–793
Wright CM, Dent OF, Newland RC, Barker M, Chapuis PH, Bokey EL, Young JP, Leggett BA, Jass JR, Macdonald GA (2005) Low level microsatellite instability may be associated with reduced cancer specific survival in sporadic stage C colorectal carcinoma. Gut 54: 103–108
Acknowledgements
The authors thank Mrs Kerstin Näslund for excellent technical help concerning the immunohistochemistry. We also thank all the participants in both CRUMS and NSHDS. This work was supported by Swedish Cancer Society, the Swedish Research Council, Cutting-Edge Research Grant from the County Council of Västerbotten, Sweden, and the Cancer Research Foundation in Northern Sweden, Sweden
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Eklöf, V., Wikberg, M., Edin, S. et al. The prognostic role of KRAS, BRAF, PIK3CA and PTEN in colorectal cancer. Br J Cancer 108, 2153–2163 (2013). https://doi.org/10.1038/bjc.2013.212
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2013.212
Keywords
This article is cited by
-
The role of the PTEN/mTOR axis in clinical response of rectal cancer patients
Molecular Biology Reports (2022)
-
Mutation Status and Prognostic Value of KRAS and BRAF in Southeast Iranian Colorectal Cancer Patients: First Report from Southeast of Iran
Journal of Gastrointestinal Cancer (2021)
-
NRAS mutant E132K identified in young-onset sporadic colorectal cancer and the canonical mutants G12D and Q61K affect distinct oncogenic phenotypes
Scientific Reports (2020)
-
A longitudinal study of prediagnostic metabolic biomarkers and the risk of molecular subtypes of colorectal cancer
Scientific Reports (2020)
-
Biomarker-guided therapy for colorectal cancer: strength in complexity
Nature Reviews Clinical Oncology (2020)